Documentation/3.5
From Slicer Wiki
Revision as of 01:17, 11 December 2009 by Sylvainjaume (talk | contribs) (→Segmentation: add link to documentation of new EMSegment module)
Home < Documentation < 3.5
Contents
Main GUI
Main GUI
- Main Application GUI
- "Hot-keys" and Keyboard Shortcuts
- Loading Scenes and Individual Datasets through the Data Module
- Data Loading Details
- Saving Scenes and Data
- Creating and Restoring Scene Snapshots
- Extensions Management Wizard (Terry G.)
Modules
- Please copy the template linked below, paste it into your page and customize it with your module's information.
Slicer3:Module_Documentation-3.5_Template
- See below for info to be put into the Help and Acknowledgment Tabs
- To put your lab's logo into a module, see here
Please adhere to the naming scheme for the module documentation:
- [ [Modules:MyModuleNameNoSpaces-Documentation-3.5|My Module Name With Spaces] ] (First Last Name)
Requirements for Modules
|
Examples for the Help and
Acknowledgment Panels |
List of Modules new to 3.5
- MRI Bias Field Correction (Nicolas Rannou, Sylvain Jaume)
- 4D Image (Viewer) (Junichi Tokuda)
- 4D Analysis (Time-intensity curve plotting and analysis) (Junichi Tokuda)
- Fast Marching segmentation (Andriy Fedorov)
- Gyri Contour Segmentation (Peter Karasev)
- Subvolume extraction with ROI widget (Andriy Fedorov)
- Registration Metrics (HD and DSC) (Haytham Elhawary)
- Cast Image (Nicole Aucoin)
List of pre-existing Modules
Core
- Camera Module (Sebastian Barre)
- Welcome Module (Wendy Plesniak, Steve Pieper, Sonia Pujol, Ron Kikinis)
- Volumes Module (Alex Yarmarkovich, Steve Pieper)
- Diffusion Editor (Kerstin Kessel)
- Models Module (Alex Yarmarkovich)
- Fiducials Module (Nicole Aucoin)
- Data Module (Alex Yarmarkovich)
- Slices Module (Jim Miller)
- Color Module (Nicole Aucoin)
- Interactive Editor (Steve Pieper)
- ROI Module (Alex Yarmarkovich)
- Volume Rendering Module (Yanling Liu, Alex Yarmarkovich)
Specialized Modules
Please adhere to the naming scheme for the module documentation:
- [ [Modules:MyModuleNameNoSpaces-Documentation-3.5|My Module Name With Spaces] ] (First Last Name)
Wizards
- ChangeTracker (Andriy Fedorov)
- IA FE Meshing Module (Vince Magnotta)
Informatics Modules
- Fetch Medical Informatics Module (Wendy Plesniak)
- QDEC Module (Nicole Aucoin)
- Query Atlas Module (Wendy Plesniak)
Registration
- Overview:
- The Register Images module is an integrated solution to all your registration needs, if you want to have a resampled volume as output. It provides access to rigid, affine and b-spline itk technologies.
- The Transforms Module allows to manually align two volumes. This can be used for initial alignment.
- Linear, affine and Deformable B-Spline modules can be used stand-alone or one after the other. They can accept transformation matrices as the start pose and produce either transforms or resampled volumes as output.
- Transformation matrices derived from these modules can be used as input for resampling other volumes (including DTI) using the Resample Volume 2 module.
- Register Images (Stephen Aylward)
- Transforms Module (Alex Yarmarkovich)
- Linear Registration (Daniel Blezek)
- Affine Registration (Daniel Blezek)
- Deformable B-Spline Registration (Bill Lorensen)
- ACPC Transform (Nicole Aucoin)
Segmentation
- EM Segment (Sylvain Jaume, Nicolas Rannou)
- EM Segment Command-Line (Brad Davis, Will Schroeder)
- EM Segment Simple (Brad Davis, Will Schroeder)
- EM Segment Template Builder (Brad Davis, Will Schroeder)
- Simple Region Growing (Jim Miller)
- Otsu Threshold (Bill Lorensen)
Statistics
- Label Statistics (Steve Pieper)
Diffusion
DWI
- Estimation
- Diffusion Tensor Estimation (Raul San Jose Estepar)
- Python Extract Baseline DWI Volume (Julien von Siebenthal)
- Filter
- Joint Rician LMMSE Image Filter (Antonio Tristán Vega, Santiago Aja Fernandez)
- Rician LMMSE Image Filter (Antonio Tristán Vega, Santiago Aja Fernandez, Marc Niethammer)
- Unbiased Non Local Means filter for DWI (Antonio Tristán Vega, Santiago Aja Fernandez)
- Python Shift DWI Values (Julien von Siebenthal)
- Python Recenter Scalar to DWI Volume (Julien von Siebenthal)
DTI
- Resample DTI Volume (Francois Budin)
- Diffusion Tensor Scalar Measurements (Raul San Jose Estepar)
- Analysis
Tractography
- Label Seeding (Raul San Jose Estepar)
- Fiducial Seeding (Alex Yarmakovich, Steve Pieper)
- FiberBundles (Alex Yarmakovich)
- Python Stochastic Tractography (Julien von Siebenthal)
- ROI Select (Lauren O'Donnell)
IGT
- OpenIGTLinkIF Module (Junichi Tokuda)
- NeuroNav Module (Haiying Liu)
- ProstateNav Module (Junichi Tokuda)
Filtering
- Checkerboard Filter (Bill Lorensen)
- Histogram Matching (Bill Lorensen)
- Image Label Combine (Alex Yarmarkovich)
- Resample Volume (Bill Lorensen)
- Resample Volume2 (Francois Budin)
- Threshold Image (Nicole Aucoin)
- Arithmetic
- Add Images (Bill Lorensen)
- Subtract Images (Bill Lorensen)
- Denoising
- Gradient Anisotropic Filter (Bill Lorensen checked this in)
- Curvature Anisotropic Diffusion (Bill Lorensen)
- Gaussian Blur (Julien Jomier, Stephen Aylward)
- Median Filter (Bill Lorensen)
- Morphology
- Voting Binary Hole Filling (Bill Lorensen)
- Grayscale Fill Hole (Bill Lorensen)
- Grayscale Grind Peak (Bill Lorensen)
Surface Models
- Modelmaker (Nicole Aucoin)
- Grayscale Model Maker (Bill Lorensen)
- Freesurfer Surface Section Extraction (Katharina Quintus)
- Python Surface Connectivity (Luca Antiga, Daniel Blezek)
- Python Surface ICP Registration (Luca Antiga, Daniel Blezek)
- Python Surface Toolbox (Luca Antiga, Daniel Blezek)
- Clip Model (Alex Yarmarkovich)
- Model into Label Volume (Nicole Aucoin)
Batch processing
- EM Segmenter batch (Julien Jomier, Brad Davis)
- Gaussian Blur batch (Julien Jomier, Stephen Aylward)
- Register Images batch (Julien Finet, Stephen Aylward)
- Resample Volume batch (Julien Finet)
Converters
- Create a Dicom Series (Bill Lorensen)
- Dicom to NRRD (Xiaodong Tao)
- Orient Images (Bill Lorensen)
- Python Explode Volume Transform (Luca Antiga, Daniel Blezek)
- Extract Subvolume (Steve Pieper)
Slicer Extensions
Extensions for Downloading
Introduction
- Slicer Extensions are a mechanism for third parties to provide modules which extend the functionality of 3d Slicer.
- Some of the extensions do not use the Slicer license. Please review carefully.
- For a subset of extensions, you can use the extension wizard in Slicer to find their webpages and to install/uninstall individual extensions. In case of problems with those modules, please talk directly to the developers of the extensions.
- The version that is available through the extension manager is chosen by the developer of that extension
We are using NITRC as the primary repository for contributed extensions. As a general rule, we do not test the extensions ourselves. Use them at your own risk. Click here to see a listing of Slicer 3 extensions on NITRC.
To add extension modules to an installed binary of slicer:
- Use the View->Extension Manager menu option
- The dialog will be initialized with the URL to the extensions that have been compiled to match your binary of slicer.
- Note installing extensions from a different repository URL is likely to be unstable due to platform and software version differences.
- You can select a local install directory for your downloaded extensions (be sure to choose a directory with enough free space).
- Select the extensions you wish to install and click to download them. Installed extensions will be available when you restart slicer.
- To turn modules on or off, you can use the Module Settings page of the View->Application Settings dialog.
- Extensions are compiled as part of the nightly build. In order to have your extension compiled nightly and made available to end users, please contact the Slicer team. For explanations for developers see here
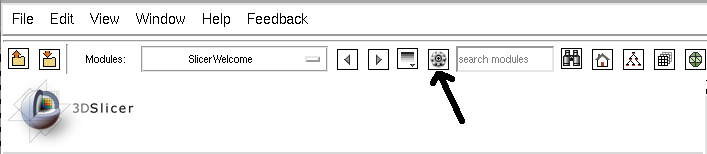
Installation
- Click on the icon to start the extensions wizard
Listing of available plug-ins
- VMTK (Daniel Haehn)
- You need to install the VmtkSlicerModule to run any of the other three.
- VMTKEasyLevelSetSegmentation - providing level-set segmentation of vessels, aneurysms and tubular structures using an easy interface
- VMTKLevelSetSegmentation - providing level-set segmentation of vessels, aneurysms and tubular structures using different algorithms
- VMTKVesselEnhancement - providing vessel enhancement filters to highlight vasculature or tubular structures
- EM DTI Clustering (Mahnaz Maddah)
- Label Diameter Estimation (Andriy Fedorov)
Extensions Contact List
This is a first pass list of extension authors; more to come!
- EMFiberClusteringModule Mahnaz Maddah maddah @nospam@ ge.com
- BRAINSROIAuto Greg Harris gregory-harris @nospam@ uiowa.edu
- Bubble Maker Carlos Mendoza carlos.sanchez.medoza @nospam@ gmail.com
- Hammer Registration Dinggang Shen dgshen @nospam@ med.unc.edu
- Lupus Lesion Mark Scully mscully @nospam@ mrn.org
- VmtkSlicerModule Daniel Haehn haehn @nospam@ bwh.harvard.edu
- Vmtkin3DSlicer Daniel Haehn
- BRAINSFit Eun Young Kim eunyoung-kim @nospam@ uiowa.edu
- ARCTIC Cedric Mathieu ced.mathieu @nospam@ gmail.com