Modules:Model Maker-Documentation-3.4
Return to Slicer 3.4 Documentation
Model Maker
ModelMaker
General Information
Module Type & Category
Type: CLI
Category: Model Generation
Authors, Collaborators & Contact
- Nicole Aucoin: Brigham and Women's Hospital
- Bill Lorensen, GE
Module Description
The Modelmaker is used to create 3D surface models from segmented image data, called label maps. Label maps can be the result of automated segmentation or interactive editing.
Usage
Select an input volume from the label map volumes that are present in the scene. Create a new model hierarchy node. Pick a label, range of labels, or all, from which to generate a surface model. Press Apply.
Examples, Use Cases & Tutorials
- After using the Editor module to segment a volume, use this module to generate a 3D surface of one segment.
Quick Tour of Features and Use
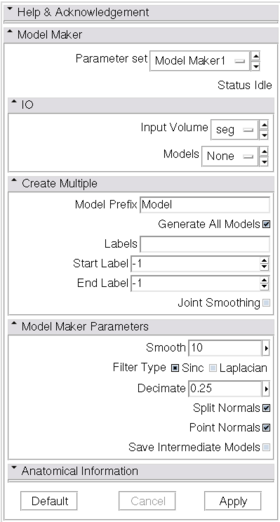
- Model Maker panel:
- Parameters Set: The standard CLI parameter node, created automatically when the values of the GUI widgets are changed from the defaults.
- I/O panel:
- The Input Volume drop down menu is populated with the label map volumes that are present in the scene
- Models are imported into Slicer under a model hierarchy node, and their colors are set by the color table associated with the input label map volume. The model hierarchy node must be created before running the model maker, by selecting Create New ModelHierarchy from the Models drop down menu. If you're running from the command line, a model hierarchy node in a new mrml scene will be created for you (Jan/09).
- Create Multiple panel:
- Any text entered in the Model Prefix box will be the starting string for the created model's file names. The label number and the color name will also be part of the file name.
- If you want to create all models that correspond to all values in a labelmap volume, click the Generate All Models and Joint Smoothing boxes
- Joint Smoothing will ensure that all resulting models fit together smoothly, like jigsaw puzzle pieces. Otherwise the models will be smoothed independently and may overlap.
- If you specify a list of Labels, it will override any start/end label settings. If you click Generate All Models it will override the list of labels and any start/end label settings.
- Model Maker Parameters panel:
- You can set the number of smoothing iterations, target reduction in number of polygons (decimal percentage). Use 0 and 1 if you wish no smoothing nor decimation.
- You can control the type of smoothing done on the models by selecting a Filter Type.
- You can set the flags to split normals or generate point normals in this pane as well.
- Split Normals is useful for visualizing sharp features. However it creates holes in surfaces which affects measurements.
- Point Normals is turned on if you wish to calculate the normal vectors for the points.
- You can save a copy of the models after intermediate steps (marching cubes, smoothing, and decimation if not joint smoothing, otherwise just after decimation); these models are not saved in the mrml file, turn off deleting temporary files first in the tcl window, and then load them manually:
[$::slicer3::CommandLineModuleGUI_Model_Maker GetLogic] DeleteTemporaryFilesOff
- Anatomical Information panel:
- Here you can specify an external file linking label values with anatomical names. This was the original method of linking label map values to resultant model colors and names, but it has been superseded by an internal linkage between the input label map volume and it's color node. If you wish to specify a file for a volume node that does not have a color node associated with it. It must be a comma separated file, extension .csv, in the format:
anatomyName1,label1 anatomyName2,label2
where anatomyName is a string, and label is an integer. If the volume node has a color node associated with it, that will override the anatomy label file.
- You can choose to not generate models from labels that do not have names by selecting the Skip Un-Named labels flag.
Development
The model maker is a pipeline of algorithms that start from the input label map, creates a binary label map with just the label(s) of interest set to 1, everything else to 0, generates a marching cubes model, runs triangle reduction and triangle smoothing algorithms. The pipeline was optimized for 1mm brain MRI data. For other geometries, adjustments of the parameters might be necessary.
The model maker is compiled as both a command line executable passed arguments via the command line, and as a binary plug in that communicates through shared memory with Slicer3.
Dependencies
The Volumes and Models modules are required for this module's use.
Known bugs
Follow this link to the Slicer3 bug tracker.
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Source Code:
Doxygen Documentation:
More Information
Acknowledgment
This work is part of the National Alliance for Medical Image Computing (NAMIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 EB005149.