Modules:QDECModule-Documentation-3.4
Return to Slicer 3.4 Documentation
QDEC
QDEC
General Information
Module Type & Category
Type: Interactive
Category: Informatics
Authors, Collaborators & Contact
- Nicole Aucoin: Brigham and Women's Hospital
- Nick Schmansky: MGH
- Doug Greve: MGH
- Contact: Nicole Aucoin, nicole@bwh.harvard.edu
Module Description
The Qdec Module provides support for the QDEC program from MGH: Query, Design, Estimate, Contrast.
Slicer3 provides integrated file format support for the FreeSurfer geometry files, scalar overlays, and volume files. Through the QdecModule GUI, it also provides an interface to launch queries on subject populations that have been processed by FreeSurfer morphometry autosegmentation pipelines. Users can also load precomputed data sets and inspect the statistical processing results.
Usage
Examples, Use Cases & Tutorials
- This module is useful for visualising the output of FreeSurfer's GLM processing
- 2007 Project Week page
- Presentation from 2007
Quick Tour of Features and Use
Subject scans
- Obtained from your local MRI scanner, processed using FreeSurfer
- Place them in a subjects directory on disk
- Add a qdec directory at the same level as the subjects
- Create a qdec.table.dat file to describe the subject
Once you've set up a subjects directory, set your SUBJECTS_DIR environment variable (you'll need to restart Slicer3). In that directory, make sure you've put the fsaverage subject created from the subjects you wish to analyse. When the FreeSurfer analysis is done over a set of subjects, an average subject is computed. The fsaverage directory holds an averaged brain on which the group statistics will be displayed
Then make a subdirectory qdec, and in it place your qdec.table.dat file along with any .level files describing the discrete factors (ie gender.levels will have 'male' and 'female' on two separate lines).
Sample qdec.table.dat:
ID Gender Age CSF OAS1_0001_MR1 F 74 1229.0 OAS1_0002_MR1 F 55 773.0 OAS1_0003_MR1 F 73 1448.0 OAS1_0004_MR1 M 28 1286.0 OAS1_0005_MR1 M 18 1304.0 OAS1_0006_MR1 F 24 909.0
Sample gender.levels:
male female
You can also load precomputed .qdec archives using the 'Load Results Data File' button. This requires unzip and rm, and you can point Slicer to these applications via the Application Settings interface (View->Application Settings). Slicer3 uses it's QDEC library to unpack the .qdec file and load the contents:
- Average brain surface file
- Brain curvature overlay
- Statistical overlays corresponding to the contrast questions
- Volume holding data for each subject
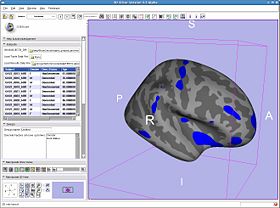
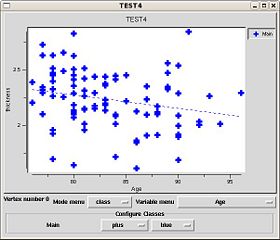
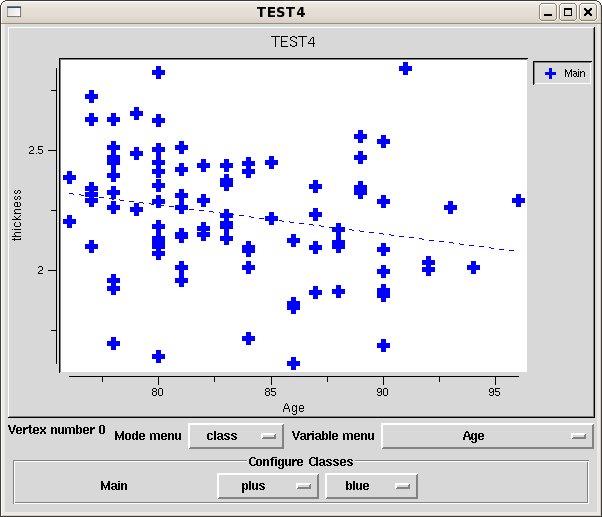
Once the results are loaded, the first point in the average brain is used to pop up a plot of all the subject values at that vertex. You can plot results at various vertices by setting the setting the 3DViewer mouse mode to 'pick' using the toolbar button and clicking on the brain model. The plot window shows the RAS and index of the chosen vertex.
Panels
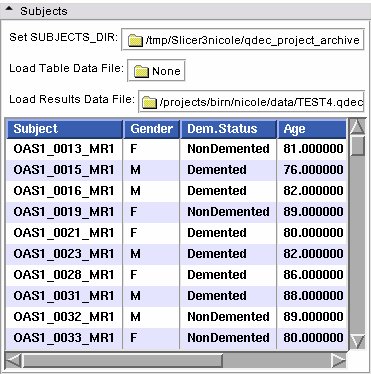
- Subjects panel:
| |
| Set SUBJECTS_DIR | Set the SUBJECTS_DIR environment variable |
| Load Table Data File | Load a .dat file |
| Load Results Data File | Load a .qdec file |
| Table of Subjects | Filled in from the subjects on disk |
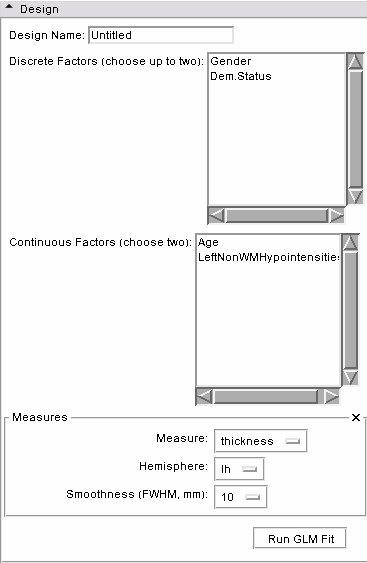
- Design panel:
- Display panel:
You can inspect all the statistical results of the analysis by switching to different overlays using the “Questions” menu.
Once the average brain surface and statistical overlays are loaded into Slicer3, you can use the QueryAtlas module to load the brain region labels and browse the annotations.
Development
Dependencies
The QDEC library (included with Slicer3) is required for this module. FreeSurfer is required for this module's use in calculating new GLM results, but is not necessary for loading pre computed ones.
Known bugs
Follow this link to the Slicer3 bug tracker.
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Module Source Code:
- vtkQdecModuleGUI.cxx, vtkQdecModuleGUI.h
- vtkQdecModuleLogic.cxx, vtkQdecModuleLogic.h
- vtkGDFReader.cxx, vtkGDFReader.h
Library Source Code:
- QdecContrast.cpp, QdecContrast.h
- QdecDataTable.cpp, QdecDataTable.h
- QdecFactor.cpp, QdecFactor.h
- QdecGlmDesign.cpp, QdecGlmDesign.h
- QdecGlmFit.cpp, QdecGlmFit.h
- QdecGlmFitResults.cpp, QdecGlmFitResults.h
- QdecProject.cpp, QdecProject.h
- QdecSubject.cpp, QdecSubject.h
- QdecUtilities.cpp, QdecUtilities.h
- vtkFreeSurferReaders.tcl
Module Doxygen documentation:
Library Doxygen documentation:
More Information
Acknowledgment
This work is part of the National Alliance for Medical Image Computing (NAMIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 EB005149.