Modules:ROISelect-Documentation-3.5
Return to Slicer 3.5 Documentation
Module Name
ROI Select
General Information
Module Type & Category
Type: CLI
Category: Tractography -> Editor
Authors, Collaborators & Contact
- Author1: Lauren O'Donnell, BWH
- Contributor1: Alex Yarmakovich, Isomics
- Contact: http://lmi.bwh.harvard.edu/~odonnell]
Module Description
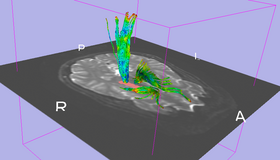
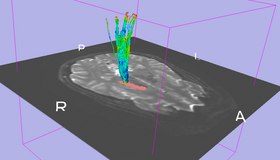
ROI Select allows a user to select tracts passing through a region of interest (ROI). The ROI is defined as a labelmap and has to be provided by the user. One approach to generate the ROI is by means of the Editor module.
Usage
Examples, Use Cases & Tutorials
We want to select only those tracts passing through a given region, where that region is defined using a labelmap. This can be used to get a more thorough seeding by seeding in a large region and selecting a subset of tracts that are of interest (as opposed to seeding directly in the region of interest). Additionally, ROIs can be used to "clean up" seeding from fiducials by retaining only those fibers of interest.
- First, a region of interest in the tract that is under study has to be defined for each subject, for example by manually delineating the ROI using the Editor module. The Fractional Anisotropy (FA) Volume can be used as reference volume to trace the ROI. The FA volume can be computed using Diffusion Tensor Scalar Measurements.
- Second, if the DTI volume is not direcly available, the DWI has to be loaded and the tensor has to be created using the | Diffusion Tensor Estimation.
- Third, seed tractography either using ROI Seeding (normally with a larger or different region than selection ROI) or Fiducial Seeding using the DTI volume.
- Fourth, select fibers of interest using ROI Select.
Quick Tour of Features and Use
The module is very straighforward to use:
- Input ROI A:set the labelmap volume that is going to be used for selection
- ROI definition panel:
- ROI pass label:set the label value that is going to be used for fiber selection (AND)
- ROI not pass label: (optional) set the label value that is going to be used for fiber rejection (NOT)
- Output panel:
- Viewing panel:
Caveats and other limitations
ROISelect module may not work when there are transforms applied to the ROI labelmap or fiber tracts in the scene.
Development
Dependencies
None
Known bugs
Follow this link to the Slicer3 bug tracker.
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Information about the DTMRI infrastructure in Slicer 3 and related classes can be found here
Links to documentation generated by doxygen.
More Information
Acknowledgment
Include funding and other support here.
References
Publications related to this module go here. Links to pdfs would be useful.