Modules:VMTKVesselEnhancement
Return to Slicer 3.6 Documentation
Module Name
VMTKVesselEnhancement, part of the  collection
collection
General Information
Module Type & Category
Type: Scripted Module
Category: Filtering, Extension
Authors, Collaborators & Contact
- Author: Daniel Haehn, University of Heidelberg
- Acknowledgments: Luca Antiga, Mario Negri Institute; Steve Pieper, Isomics Inc.
- Contact: Daniel Haehn, haehn@bwh.harvard.edu
Module Description
This module provides the vessel enhancement filters of the Vascular Modeling Toolkit (http://www.vmtk.org) in 3D Slicer. These filters target the highlighting of line-like, plate-like and blob-like structures.
This work is part of the NA-MIC VMTK Collaboration.
Official project page: http://www.vmtk.org/Main/VmtkIn3DSlicer
Usage
Installation
This module depends on the VmtkSlicerModule: see this page for installation notes.
The VMTKVesselEnhancement module can be installed using the 3D Slicer extension wizard. The extension is called VMTKVesselEnhancement.
When the module was successfully installed, it is available within 3D Slicer's module selector inside the category Vascular Modeling Toolkit.
Examples, Use Cases
The following screencasts show different Use Cases of the VMTKVesselEnhancement module. It is always used as a pre-processing step and the actual segmentation is performed using the VMTKLevelSetSegmentation module.
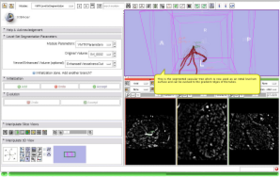
- Use Case #1: Highlighting the Coronary Tree inside a blood-pool MRI using the Sato Vesselness filter
 Watch the video.. |
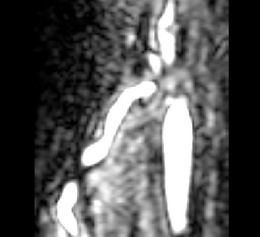
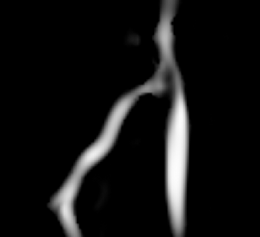
This screencast shows the extraction of a coronary tree inside a cardiac blood-pool MRI by using a combination of the VMTKVesselEnhancement and the VMTKLevelSetSegmentation modules. Before the actual segmentation, VMTKVesselEnhancement is used to highlight the coronary tree which results in a much better segmentation. Watch the video here. |
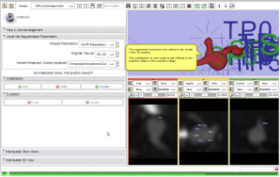
- Use Case #2: Enhancement of vessels connected to a cerebral aneurysm to support the segmentation
 Watch the video.. |
The segmentation of a cerebral aneurysm and its connecting vessels is shown in this screencast. Sato Vesselness of the VMTKVesselEnhancement module is used as a pre-processing step and the segmentation of the connecting vessels is performed on the vessel enhanced feature image using the VMTKLevelSetSegmentation module. The segmentation of the actual aneurysm is performed on the original image. How to establish the pipeline between VMTKVesselEnhancement and the VMTKLevelSetSegmentation module easily is explained as well. Watch the video here. |
Tutorials
The following tutorials demonstrate the use of VMTKVesselEnhancement as a processing step in a pipeline of VMTK modules to extract the centerlines of coronary arteries.
 Step-by-step online tutorial: Segmentation of the Right Coronary Artery using VMTK in 3D Slicer. View online here. |
 Step-by-step tutorial including example dataset: The first slide of the Centerline Extraction tutorial. Download here. |
Quick Tour of Features and Use
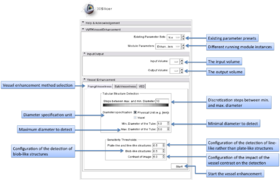
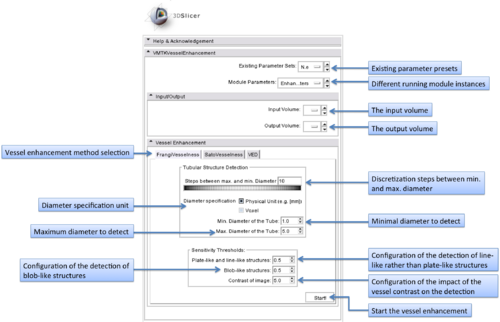
The graphical user interface of the VMTKVesselEnhancement module consists of two panels.
- I/O panel:
The MRML volume nodes to use as input and output can be specified at this panel.
- Vessel Enhancement panel:
Different filtering methods are available to highlight or enhance tubular structures. These can be chosen by using the tabs.
The range of tube diameters to be highlighted can be specified in voxels or millimeters. The steps between the minimal and maximal diameters regulate the accuracy of the vessel detection process.
Sensitivity thresholds can be specified to control the filtering for different types of structures. The lower a threshold is, the more sensitive is the detection of this structure type. A common approach is to set this value to 0.25*Max_Intensity.
For example to enhance line-like structures rather than plate-like or blob-like structures, set the sensitivity thresholds for Frangi's method to the following:
- Plate-like and line-like structures: 0.3
- Blob-like structures: 500
The third threshold specifies how important the contrast between the foreground and the background is to highlight the local structure. A higher value implies a higher contrast is required.
Development
Notes from the Developer(s)
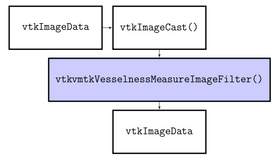
The algorithms provided by VMTK and used by this module are C++ classes implemented using VTK and ITK. The module using the following algorithms: FrangiVesselness based on Multiscale Vessel Enhancement Filtering by Frangi A. et al. (http://www.tecn.upf.es/~afrangi/articles/miccai1998.pdf), SatoVesselness based on Three-dimensional multi-scale line filter for segmentation and visualization of curvilinear structures in medical images by Sato Y. et al. (http://www.spl.harvard.edu/publications/bitstream/download/2732) and VesselEnhancingDiffusion based on Vessel Enhancing Diffusion Filter by Enquobahrie A. et al. (http://hdl.handle.net/1926/558).
Dependencies
This module depends on the VMTK libraries which are provided in the VmtkSlicerModule. Therefore the VmtkSlicerModule has to be installed before the VMTKVesselEnhancement module can be used.
Known bugs & Usability issues
Follow this link to the VMTK in 3D Slicer bug tracker.
Source code & documentation
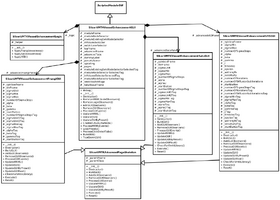
VMTKVesselEnhancement is a Python Scripted Module. It follows the conventions of the Model View Controller pattern of slicer modules. This implies the separation of logic and GUI.
The class SlicerVMTKVesselEnhancementGUI derives from ScriptedModuleGUI and saves the current user interface state to its own MRML node. The MRML nodes of the used volume and model data are additionally attached. This allows different instances of the vessel enhancement module to be run at the same time. The actual calls to the VMTK libraries are only performed in the class SlicerVMTKVesselEnhancementLogic.
Separate classes derived from the interface SlicerVMTKAdvancedPageSkeleton were created for each filtering method in order to ensure the possibility of maintenance and extension. The calls to the required methods like UpdateGUI() or UpdateMRML() get forwarded to each individual sub-class to support the modular design and enable code encapsulation.
The complete source code is available at a NITRC SVN repository.
More Information
Acknowledgment
This work was funded by a grant of the Thomas-Gessmann Foundation part of the Founder Federation for German Science.
References
- Antiga L, Piccinelli M, Botti L, Ene-Iordache B, Remuzzi A and Steinman DA. An image-based modeling framework for patient-specific computational hemodynamics. Medical and Biological Engineering and Computing, 46: 1097-1112, Nov 2008.
- D. Hähn. Integration of the vascular modeling toolkit in 3d slicer. SPL, 04 2009. Available online at http://www.spl.harvard.edu/publications/item/view/1728.
- D. Hähn. Centerline Extraction of Coronary Arteries in 3D Slicer using VMTK based Tools. Master's Thesis. Department of Medical Informatics, University of Heidelberg, Germany. Feb 2010.
- Piccinelli M, Veneziani A, Steinman DA, Remuzzi A, Antiga L (2009) A framework for geometric analysis of vascular structures: applications to cerebral aneurysms. IEEE Trans Med Imaging. In press.