Modules:LabelDiameterEstimation-Documentation-3.5
Return to Slicer 3.5 Documentation
Label Diameter Estimation
LabelDiameterEstimation
General Information
Module Type & Category
Type: CLI
Category: Statistics
Authors, Collaborators & Contact
- Andriy Fedorov, BWH
- Ron Kikinis, BWH
- Contact: Andriy Fedorov, fedorov at bwh
Module Description
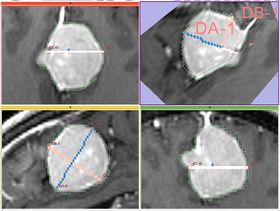
Given a binary label, this module estimates the largest diameter of the label, its largest in-plane perpendicular diameter, and the third diameter that is perpendicular to the plane of the first two.
Usage
Examples, Use Cases & Tutorials
This module was originally designed to calculate the dimensions of tumor, following the RECIST [1] and WHO guidelines.
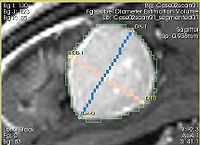
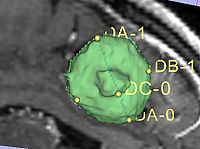
- Implementation details The module implements the following algorithm for calculating the dimensions of the label. First, the area of the label slice is computed in all the slices of the image that are passing through the label. This is done for the three orthogonal directions of the image. The largest area slice is found. The largest diameter (DA) is estimated by finding the two most distant points on the contour of the label cross-section. The second diameter (DB) is found by finding the two most distant points on the contour that lie on the line perpendicular to the first diameter. The third diameter (DC) is estimated by calculating the two points of intersection between the line perpendicular to the plane formed by DA and DBpassing through the point of their intersection, and the contour of the analyzed label.
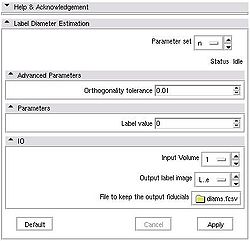
Quick Tour of Features and Use
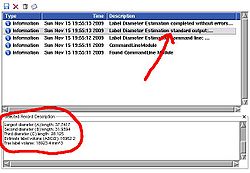
The module also outputs numeric values for the lengths of the computed diameters. To see these numbers, open Log Viewer, and select the first from top log record with the name Label Diameter Estimation: standard output. The reported numbers are the lenghts of the three diameters, the volume estimate calculated as A*B*C/2 (a formula suggested in the literature for estimating volume of hematomas from planimetric measurements [2]), and the true label volume in mm^3. |
|
Development
Standing issues under development:
- would be nice to have fiducials read automatically; currently not possible, since fiducials are not fully integrated with CLI
- use measurements widgets instead of label images for diameter visualization; measurements are also not integrated with CLI, and are not yet feature-complete
- update Acks tab to point to documentation and give credit to BSF
Dependencies
Known bugs
Follow this link to the Slicer3 bug tracker.
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Source code can accessed here
Links to documentation generated by doxygen.
More Information
Acknowledgment
Supported by Brain Science Foundation.
References
[1] E. Eisenhauer, P. Therasse, J. Bogaerts, L. Schwartz, D. Sargent, R. Ford, J. Dancey, S. Arbuck, S. Gwyther, and M. Mooney, "New response evaluation criteria in solid tumours: Revised recist guideline (version 1.1)," European Journal of Cancer, vol. 45, no. 2, pp. 228-247, January 2009. DOI
[2] H. B. Huttner, T. Steiner, M. Hartmann, M. Kohrmann, E. Juettler, S. Mueller, J. Wikner, U. Meyding-Lamade, P. Schramm, S. Schwab, and P. D. Schellinger, "Comparison of abc/2 estimation technique to computer-assisted planimetric analysis in warfarin-related intracerebral parenchymal hemorrhage," Stroke, vol. 37, no. 2, pp. 404-408, February 2006. DOI