Main Page/SlicerCommunity/2019
Go to 2022 :: 2021 :: 2020 :: 2019 :: 2018 :: 2017 :: 2016 :: 2015 :: 2014-2011 :: 2010-2000
The community that relies on 3D Slicer is large and active: (numbers below updated on December 1st, 2023)
- 1,467,466+ downloads in the last 11 years (269,677 in 2023, 206,541 in 2022)
- over 17.900+ literature search results on Google Scholar
- 2,147+ papers on PubMed citing the Slicer platform paper
- Fedorov A., Beichel R., Kalpathy-Cramer J., Finet J., Fillion-Robin J-C., Pujol S., Bauer C., Jennings D., Fennessy F.M., Sonka M., Buatti J., Aylward S.R., Miller J.V., Pieper S., Kikinis R. 3D Slicer as an Image Computing Platform for the Quantitative Imaging Network. Magnetic Resonance Imaging. 2012 Nov;30(9):1323-41. PMID: 22770690. PMCID: PMC3466397.
- 39 events in open source hackathon series continuously running since 2005 with 3260 total participants
- Slicer Forum with +8,138 subscribers has approximately 275 posts every week
The following is a sample of the research performed using 3D Slicer outside of the group that develops it. in 2019
We monitor PubMed and related databases to update these lists, but if you know of other research related to the Slicer community that should be included here please email: marianna (at) bwh.harvard.edu.
Contents
- 1 2019
- 1.1 Influence of Hyrax Screw Position on Dental Movement and Cortical Bone: A Study of Finite Elements
- 1.2 Clinical Relevance of the Left Brachiocephalic Vein Anatomy for Vascular Access in Dialysis Patients
- 1.3 the Effect of Intraoperative Imaging on Surgical Navigation for Laparoscopic Liver Resection Surgery
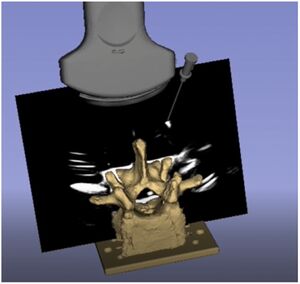
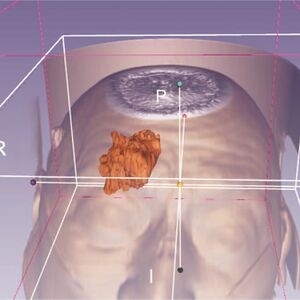
- 1.4 A New Application of Ultrasound-Magnetic Resonance Multimodal Fusion Virtual Navigation in Glioma Surgery
- 1.5 Correlation of Spontaneous and Traumatic Anterior Skull Base CSF Leak Flow Rates with Fluid Pattern on Early, Delayed, and Subtraction Volumetric Extended Echo Train T2-Weighted MRI
- 1.6 DCE-MRI Assessment of Response to Neoadjuvant SABR in Early Stage Breast Cancer: Comparisons of Single Versus Three Fraction Schemes and Two Different Imaging Time Delays Post-SABR
- 1.7 Deep Learning Approach For Automatic Out-Of-Plane Needle Localisation For Semi-Automatic Ultrasound Probe Calibration
- 1.8 Prediction of Immunohistochemistry of Suspected Thyroid Nodules by Use of Machine Learning-Based Radiomics
- 1.9 A Review on Multiplatform Evaluations of Semi-automatic Open-source Based Image Segmentation for Cranio-maxillofacial Surgery
- 1.10 Patellar Calcar: Morphometric Analysis by Knee Magnetic ResonanceImaging and Three-dimensional Reconstruction Software-assisted
- 1.11 Deep Convolutional Neural Networks for Automatic Detection of Orbital Blowout Fractures
- 1.12 Decision-making Based on 3D Printed Models in Laparoscopic Liver Resections with Intraoperative Ultrasound: A Prospective Observational Study
- 1.13 In-House Surgeon-Led Virtual Surgical Planning for Maxillofacial Reconstruction
- 1.14 Patient-specific Access Planning in Minimally Invasive Mitral Valve Surgery
- 1.15 Semi-Automatic Signature-Based Segmentation Method for Quantification of Neuromelanin in Substantia Nigra
- 1.16 Spatial Accuracy of a Clinically Established Noninvasive Electrocardiographic Imaging System for the Detection of Focal Activation in an Intact Porcine Model
- 1.17 Life Without a Brain: Neuroradiological and Behavioral Evidence of Neuroplasticity Necessary to Sustain Brain Function in the Face of Severe Hydrocephalus
- 1.18 a Case Report of Total Skin Photon Radiation Therapy for Cutaneous Epitheliotropic Lymphoma in a Dog
- 1.19 Considerations for Ultrasound Exposure During Transcranial MR Acoustic Radiation Force Imaging
- 1.20 Direct Evidence for Eudicot Pollen-feeding in a Cretaceous Stinging Wasp (Angiospermae; Hymenoptera, Aculeata) Preserved in Burmese Amber
- 1.21 Myoarchitectural Disarray of Hypertrophic Cardiomyopathy Begins Pre-Birth
- 1.22 Whole Lesion Histogram Analysis of Apparent Diffusion Coefficients on MRI Predicts Disease-free Survival in Locally Advanced Squamous Cell Cervical Cancer after Radical Chemo-radiotherapy
- 1.23 Variations in the Size and Shape of Human Cochlear Malformation Types
- 1.24 Pretreatment Prediction of Adaptive Radiation Therapy Eligibility Using MRI-Based Radiomics for Advanced Nasopharyngeal Carcinoma Patients
- 1.25 Volumetric Assessment of the Dental Crown for Sex Estimation by Means of Cone-beam Computed Tomography
- 1.26 Association of Serum Cystatin C with White Matter Abnormalities in Patients with Amnestic Mild Cognitive Impairment
- 1.27 Three-dimensional Clavicle Displacement Analysis and its Effect on ScapularPosition in Acute Clavicle Midshaft Fracture
- 1.28 The Evolutionary Radiation of Hominids: a Phylogenetic Comparative Study
- 1.29 Isolating Phyllotactic Patterns Embedded in the Secondary Growth of Sweet Cherry (Prunus Avium L.) Using Magnetic Resonance Imaging
- 1.30 Decoding Tumor Mutation Burden And Driver Mutations In Early Stage Lung Adenocarcinoma Using Ct-Based Radiomics Signature
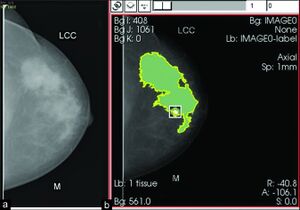
- 1.31 Mammographic Breast Density Assessed with Fully Automated Method and its Risk for Breast Cancer
- 1.32 Advantage of Proton-Radiotherapy for Pediatric Patients and Adolescents With Hodgkin's Disease
- 1.33 Potential Role of Convolutional Neural Network Based Algorithm in Patient Selection for DCIS Observation Trials Using a Mammogram Dataset
- 1.34 Texture Analysis of Pretreatment 18FFDG PET/CT for the Prognostic Prediction of Locally Advanced Salivary Gland Carcinoma Treated with Interstitial Brachytherapy
- 1.35 Levator Bowl Volume during Straining and its Relationship to other Levator Measures
- 1.36 Mixed Reality-Based Preoperative Planning for Training of Percutaneous Transforaminal Endoscopic Discectomy: A Feasibility Study.
- 1.37 Evaluation of Accuracy of a Three-Dimensional Printed Model in Open-Wedge High Tibial Osteotomy
- 1.38 Neuroradiological Changes Following Single or Repetitive Mild TBI
- 1.39 Technique Development and Measurement of Cross-Sectional Area of the Pubovisceral Muscle on MRI Scans of Living Women
- 1.40 A Comparative Evaluation of SteamVR Tracking and the OptiTrack System for Medical Device Tracking
- 1.41 The Middle Fossa Approach with Self-drilling Screws: a Novel Technique for BONEBRIDGE Implantation
- 1.42 Evaluation of Flow Changes After Telescopic Stenting of a Giant Fusiform Aneurysm of the Vertebrobasilar Junction
- 1.43 a New Acoustic Coupling Fluid With Ability to Reduce Ultrasound Imaging Artefacts in Brain Tumour Surgery-A Phase I Study
- 1.44 CT-Based Radiomic Signatures for Prediction of Pathologic Complete Response in Esophageal Squamous Cell Carcinoma After Neoadjuvant Chemoradiotherapy
- 1.45 Diffusion Abnormalities in the Corpus Callosum in First Episode Schizophrenia: Associated With Enlarged Lateral Ventricles and Symptomatology
- 1.46 Computed Tomographic Portography with Esophageal Variceal Measurements in the Evaluation of Esophageal Variceal Severity and Assessment of Esophageal Variceal Volume Efficacy
- 1.47 Intracranial Mirror Aneurysm: Epidemiology, Rupture Risk, New Imaging, Controversies, and Treatment Strategies
- 1.48 Development and Evaluation of a "Trackerless" Surgical Planning and Guidance System Based on 3D Slicer
- 1.49 Multi-objective Parameter Auto-tuning for Tissue Image Segmentation Workflows
- 1.50 In vivo Localization of Cortical Areas using a 3D Computerized Atlas of the Marmoset Brain
- 1.51 Quantification of Global Lung Inflammation Using Volumetric 18F-FDG PET/CT Parameters in Locally Advanced Non-Small-Cell Lung Cancer Patients Treated With Concurrent Chemoradiotherapy: A Comparison of Photon and Proton Radiation Therapy
- 1.52 Comprehensive, Multiscale Framework for Evaluation of Arrhythmias Arising From Cell Therapy in the Whole Post-Myocardial Infarcted Heart
- 1.53 Diagnostic Performance of Clinical Properties and Conventional Magnetic Resonance Imaging for Determining the IDH1 Mutation Status in Glioblastoma: A Retrospective Study
- 1.54 Processing Pipeline for Atlas-Based Imaging Data Analysis of Structural and Functional Mouse Brain MRI (AIDAmri).
- 1.55 Ancient Machine Tools for the Construction of the Antikythera Mechanism Parts
- 1.56 Comprehensive Review of 3D Segmentation Software Tools for MRI Usable for Pelvic Surgery Planning
- 1.57 New, Simple and Reliable Volumetric Calculation Technique in Incisional Hernias with Loss of Domain
- 1.58 Predicting Malignant Potential of Subsolid Nodules: Can Radiomics Preempt Longitudinal Follow up CT?
- 1.59 Imaging Phenotype Using Radiomics to Predict Dry Pleural Dissemination in Non-Small Cell Lung Cancer
- 1.60 Three-dimensional Neuronavigation in SEEG-guided Epilepsy Surgery
- 1.61 The Future of Biomechanical Spine Research: Conception and Design of a Dynamic 3D Printed Cervical Myelography Phantom
- 1.62 Treatment of Intracranial Hemorrhage With Neuroendoscopy Guided by Body Surface Projection
- 1.63 Alterations in Brain Neurocircuitry Following Treatment With the Chemotherapeutic Agent Paclitaxel in Rats
- 1.64 Comprehensive Analysis of Animal Models of Cardiovascular Disease using Multiscale X-Ray Phase Contrast Tomography
- 1.65 3D Superimposition of Craniofacial Imaging-The Utility of Multicentre Collaborations
- 1.66 Complete Thoracolumbar Fracture-dislocation with Intact Neurologic Function: Explanation of a Novel Cord Saving Mechanism
- 1.67 Whole-Body FDG PET/MR Atlas for Multiparametric Voxel-Based Analysis
- 1.68 Complexity of Tumor Shape, Spiculatedness, Correlates With Tumor Radiomic Shape Features
- 1.69 Virtual Reconstruction of Paranasal Sinuses from CT Data: A Feasibility Study for Forensic Application
- 1.70 Integrated in Utero MR Method for Assessing Structural Brain Abnormalities and Measuring Intracranial Volumes in Fetuses With Congenital Heart Disease: Results of a Prospective Case-Control Feasibility Study
- 1.71 Novel Application and Validation of in Vivo Micro-CT to Study Bone Modelling in 3D
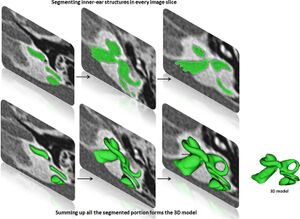
- 1.72 Human Inner-ear Malformation Types Captured in 3D
- 1.73 Bridging the Translational Gap: Implementation of Multimodal Small Animal Imaging Strategies for Tumor Burden Assessment in a Co-Clinical Trial
- 1.74 Validation of a Freehand Technique for Cortical Bone Trajectory Screws in the Lumbar Spine
- 1.75 Three-Dimensional Evaluation of the Root Resorption of Maxillary Incisors After the Orthodontic Traction of Bicortically Impacted Canines: Case Report
- 1.76 Automated 3-Dimensional Magnetic Resonance Imaging Allows for Accurate Evaluation of Glenoid Bone Loss Compared With 3-Dimensional Computed Tomography
- 1.77 Investigation of F-BAR Domain PACSIN Proteins Uncovers Membrane Tubulation Function in Cilia Assembly and Transport
- 1.78 Reproducibility and Non-Redundancy of Radiomic Features Extracted From Arterial Phase CT Scans in Hepatocellular Carcinoma Patients: Impact of Tumor Segmentation Variability
- 1.79 Real-Time Adaptive Planning Method for Radiotherapy Treatment Delivery for Prostate Cancer Patients, Based on a Library of Plans Accounting for Possible Anatomy Configuration Changes
- 1.80 Towards an Advanced Virtual Ultrasound-guided Renal Biopsy Trainer
- 1.81 Methods for Quantitative Characterization of Bone Injury From Computed-Tomography Images
- 1.82 Predicting Breast Cancer Molecular Subtype with MRI Dataset Utilizing Convolutional Neural Network Algorithm
- 1.83 Unique Metasomal Musculature in Sweat Bees (Hymenoptera, Apoidea, Halictidae) Revealed by Micro-CT Scanning
- 1.84 Quantitative Features Can Predict Further Growth of Persistent Pure Ground-Glass Nodule
- 1.85 3D Reconstruction of MR-Visible Fe3O4-Mesh Implants: Pelvic Mesh Measurement Techniques and Preliminary Findings
- 1.86 A Complete Workflow for Utilizing Monte Carlo Toolkits in Clinical Cases for a Double-Scattering Proton Therapy System
- 1.87 Accuracy Assessment of 3D-Printed Tooth Replicas
- 1.88 Glottic Configuration Changes and Outcomes of Endoscopic Arytenoid Abduction Lateropexy
- 1.89 Pyramidal Neuron Growth and Increased Hippocampal Volume During Labor and Birth in Autism
- 1.90 Opposing CSF Hydrodynamic Trends Found in the Cerebral Aqueduct and Prepontine Cistern Following Shunt Treatment in Patients With Normal Pressure Hydrocephalus
- 1.91 Computer Simulations Suggest That Prostate Enlargement Due to Benign Prostatic Hyperplasia Mechanically Impedes Prostate Cancer Growth
- 1.92 Machine Learning Derived Segmentation of Phase Velocity Encoded Cardiovascular Magnetic Resonance for Fully Automated Aortic Flow Quantification
- 1.93 Morphological Analysis of Sigmoid Sinus Anatomy: Clinical Applications to Neurotological Surgery
- 1.94 Biomarker Localization, Analysis, Visualization, Extraction, and Registration (BLAzER) Methodology for Research and Clinical Brain PET Applications
- 1.95 Imaging as Part of a Quality Assurance Program: Predictors of Interobserver Variability for Pretreatment Image Registration for Lung SBRT
2019
Influence of Hyrax Screw Position on Dental Movement and Cortical Bone: A Study of Finite Elements
|
Publication: J Clin Exp Dent. 2019 Dec 1;11(12):e1099-e1108. PMID: 31824589 | PDF Authors: Gómez-Gómez SL, Villarraga-Ossa JA, Arcila-Monsalve JC, Moreno-Garzón DM, Ardila CM. Institution: Orthodontics; Master in Epidemiology; Assistant Professor, School of Dentistry, Universidad de Antioquia, Colombia. Abstract: BACKGROUND: Rapid maxillary expansion (RME) has effects on the dental and periodontal structures of the parts involved, which vary according to the design and position of the expansion screw. The purpose of the study was to determine the optimal three-dimensional position of the Hyrax screw to obtain precise control of the dental movement and effect on the bone cortex using the finite element method (FEM). MATERIAL AND METHODS: RME was performed from the patient whom two Cone-Beam computerized tomography scans (CBCT) were obtained: T1 before expansion, and T2 three months after stabilization of RME. The FEM model was designed with T1 and of Hyrax photographs. FEM was obtained by comparing the simulation, T2, and clinical results. Three sagittal screw positions (anterior-middle-posterior) and vertical (upper-medium-low) were evaluated. RESULTS: A coronal- buccal displacement of premolars and first molars was found which decreased in the occlusal-apical direction, presenting different types of dental movement in the screw positions; besides, a tendency of translational movement in the posterior-high location was observed. In the posterior-high position a higher concentration of efforts and homogeneous deformations in the periodontal ligament and vestibular cortex of the cervical area of first molars, first and second premolars were observed, with variations according to the screw position and the distribution of stresses. CONCLUSIONS: The ideal location of the expansion screw for controlling dental movement and periodontal side effects was the high-posterior position. "...the three-dimensional CAD model of the maxilla was constructed and processed in 3D Slicer software (open source) to obtain a cloud of points of the upper maxillary system, alveolar bone, periodontal ligament, and maxillary teeth." |
Clinical Relevance of the Left Brachiocephalic Vein Anatomy for Vascular Access in Dialysis Patients
|
Publication: Clin Anat. 2019 Dec 31. PMID: 31891199 Authors: Vertemati M, Rizzetto F, Cassin S, Zerbi P, Giordano A, Cariati M, Gallieni M. Institution: Institute of Human Anatomy, Department of Biomedical and Clinical Sciences "Luigi Sacco", Università degli Studi di Milano, Milan, Italy. Abstract: INTRODUCTION: Most hemodialysis patients start renal replacement therapy with a central venous catheter (CVC). The left internal jugular vein (LIJV) is the second-choice vein for CVC positioning, after the right IJV. However, to reach the right atrium, the CVC must pass through the left brachiocephalic vein (LBV), which also drains blood from the left arm through the subclavian vein. The purpose of this study is to describe how the anatomy of the central venous system and in particular that of the LBV affects vascular access in hemodialysis patients. MATERIALS AND METHODS: Three-dimensional (3D) virtual model reconstructions of the central thoracic veins of three hemodialysis patients were obtained from contrast-enhanced computed tomography scans acquired in the venous phase. The images were exported as DICOM files and loaded on open-source software for visualizing and analyzing the medical imaging, 3D Slicer, Windows v.4.8.1. RESULTS: As expected, the 3D reconstructions showed that the LBV has a tortuous path with three main angulations that could be associated with external compression and stenosis. These could determine the difficulties and increased risks of venous injury during CVC placement, and an increased risk of medium to long-term catheter-associated vein thrombosis and stenosis. CONCLUSIONS: The anatomical features of the LBV indicate that the path of a CVC from the LIJV to the right atrium is tortuous and can easily be complicated by vein injury, negatively affecting the creation of future arterio-venous vascular accesses in the left arm. |
|
Publication: Sci Rep. 2019 Dec 10;9(1):18687. PMID: 31822701 | PDF Authors: Teatini A, Pelanis E, Aghayan DL, Kumar RP, Palomar R, Fretland ÅA, Edwin B, Elle OJ Institution: The Intervention Centre, Oslo University Hospital, Oslo, Norway. Abstract: Conventional surgical navigation systems rely on preoperative imaging to provide guidance. In laparoscopic liver surgery, insufflation of the abdomen (pneumoperitoneum) can cause deformations on the liver, introducing inaccuracies in the correspondence between the preoperative images and the intraoperative reality. This study evaluates the improvements provided by intraoperative imaging for laparoscopic liver surgical navigation, when displayed as augmented reality (AR). Significant differences were found in terms of accuracy of the AR, in favor of intraoperative imaging. In addition, results showed an effect of user-induced error: image-to-patient registration based on annotations performed by clinicians caused 33% more inaccuracy as compared to image-to-patient registration algorithms that do not depend on user annotations. Hence, to achieve accurate surgical navigation for laparoscopic liver surgery, intraoperative imaging is recommendable to compensate for deformation. Moreover, user annotation errors may lead to inaccuracies in registration processes. "...segmentations were later exported into 3D Slicer and reconstructed into a 3D model." |
|
Publication: Ann Transl Med. 2019 Dec; 7(23): 736. PMC6989994 Authors: Chaofeng Liang,Manting Li, Jin Gong, Baoyu Zhang, Cong Lin, Haiyong He, Ke Zhang, Ying Guo Institution: Department of Neurosurgery, 3rd Affiliated Hospital of Sun Yat-sen University, Sun Yat-sen University, Guangzhou, China. Abstract: Background Long-term survival and high-quality life of patients with gliomas depends on the extent of resection (EOR) and the protection of functional white matter fibers. The navigation system provides precise positioning for surgery based on preoperative magnetic resonance imaging (MRI) but the precision decreases when intraoperative brain drift occurs. Ultrasound (US) can support real-time imaging and correct brain shift. The real-time US-MRI multimodal fusion virtual navigation system (UMNS) is a new technique for glioma surgery. In order to obtain a maximum EOR and functional protection, this study aimed to explore the feasibility, efficiency, and safety of real-time UMNS for glioma surgery, and to evaluate the benefit of the new application by UMNS presetting markers between the tumor and functional white matter fiber surgery. Methods A retrospective analysis included 45 patients who underwent glioma surgery, 19 patients with only intraoperative US, and 26 patients with UMNS. A preoperative plan was made by 3D Slicer software based on preoperative MRI. This was combined with a reconstruction of diffusion tensor imaging (DTI) that designed the important locations as “warning points” between functional white matter fibers and tumor. Following patient registration, markers were injected into preset “warning points” under image-guided UMNS in order to give us a warning during surgery in case of postoperative function deficits. The operating time, volumetric assessment in glioma resection, and postoperative complications were evaluated and used to compared those surgeries using intraoperative US (iUS) with those surgeries using intraoperate MRI (iMRI) navigation. Results A total of 45 patients underwent glioma surgery. Gross total removal (GTR) of iUS alone was achieved in 6 of 19 cases, while this was achieved in 22 of 26 cases with UMNS alone, demonstrating an improvement in rate of GTR from 31.58% to 84.62%, respectively. This may be attributable to the superior US image quality provided by UMNS. In 13 of 26 cases, there was improved image quality (from poor/moderate to moderate/good) with the aid of UMNS. In addition, the consistency of EOR of postoperative MRI evaluated by UMNS (92.31%) was higher than when using iUS alone (42.11%). The whole process of intraoperative scanning time and marker injection did not lead to a significant delay of the operating time compared to using iUS alone, and has been reported to be shorter than with iMRI as well. Furthermore, the percentage of postoperative morbidity in the UMNS group was lower than that in the iUS group (motor deficit: 11.54% vs. 42.11%; aphasia: P =3.85% vs. 31.58%, respectively). Conclusions Real-time UMNS is an effective, timesaving technology that offers high quality intraoperative imaging. Injection markers between functional white matter fibers and tumor by UMNS can help to obtain a maximum EOR of glioma and functional protection postoperatively. The integration of iUS into the neuronavigation system offered quick and helpful intra-operative images. |
Correlation of Spontaneous and Traumatic Anterior Skull Base CSF Leak Flow Rates with Fluid Pattern on Early, Delayed, and Subtraction Volumetric Extended Echo Train T2-Weighted MRI
|
Publication: J Neurosurg. 2019 Dec 27:1-9. PMID: 31881543 Authors: Rutland JW, Govindaraj S, Gill CM, Shohet M, Iloreta AMC, Bederson JB, Shrivastava RK, Delman BN. Institution: Departments of Neurosurgery, Icahn School of Medicine at Mount Sinai, New York, NY, USA. Abstract: OBJECTIVE: CSF leakage is a potentially fatal condition that may result when a skull base dural defect permits CSF communication between the cranial vault and sinonasal cavities. Flow rate is an important property of CSF leaks that can contribute to surgical decision-making and predispose patients to complications and inferior outcomes. Noninvasive preoperative prediction of the leak rate is challenging with traditional diagnostic tools. The present study compares fluid configurations on early and late volumetric extended echo train T2-weighted MRI by using image tracings and sequence subtraction as a novel method of quantifying CSF flow rate, and it correlates radiological results with intraoperative findings and clinical outcomes. METHODS: A total of 45 patients met inclusion criteria for this study and underwent 3-T MRI. Imaging sequences included two identical CUBE T2 (vendor trade name for volumetric extended echo train T2) acquisitions at the beginning and end of the scanning session, approximately 45 minutes apart. Twenty-five patients were confirmed to have definitive spontaneous or traumatic anterior skull base CSF leaks. Semiautomated volumetric segmentation of CSF intensity was performed on both CUBE data sets by using 3D Slicer software, and volumes were subtracted to obtain accumulated CSF volume. These imaging-derived fluid accumulations were correlated with high- or low-flow states, as well as ultimate treatment outcomes including recurrences. RESULTS: Of the 45 patients, 25 (55.6%) had definitive evidence of CSF leakage, and 22 (88%) of these underwent surgical repair. Patients with high-flow CSF leaks had higher early (4.058 cm3 vs 0.982 cm3, p = 0.04), late (4.58 cm3 vs 1.096 cm3, p = 0.04), and accumulated (0.53 cm3 vs 0.11 cm3, p = 0.01) fluid volume measurements than patients with low-flow leaks. The 5 (22.7%) patients who exhibited postoperative CSF leak recurrence had significantly greater early (6.30 cm3 vs 1.23 cm3, p = 0.008) and late (6.87 cm3 vs 1.45 cm3, p = 0.008) volumes. Accumulated volume was not significantly greater in patients with leak recurrence (0.58 cm3 vs 0.22 cm3, p = 0.07). Early, late, and accumulated volumes were significantly correlated with postoperative hospital stay as well as duration of postoperative lumbar drain placement (p < 0.05 for all measures). CONCLUSIONS: High-resolution CUBE T2 MRI, coupled with precise volumetric segmentation and subtraction of sinonasal hyperintensity, not only demonstrated predictive value in differentiating low- and high-flow CSF leaks, but also correlated with postoperative complications such as leak recurrence. These findings may be useful in the clinical workup and neurosurgical management of patients with skull base CSF leaks. |
DCE-MRI Assessment of Response to Neoadjuvant SABR in Early Stage Breast Cancer: Comparisons of Single Versus Three Fraction Schemes and Two Different Imaging Time Delays Post-SABR
|
Publication: Clin Transl Radiat Oncol. 2019 Dec 25;21:25-31. PMID: 32021911 | PDF Authors: Mouawad M, Biernaski H, Brackstone M, Lock M, Yaremko B, Shmuilovich O, Kornecki A, Ben Nachum I, Muscedere G, Lynn K, Prato FS, Thompson RT, Gaede S, Gelman N. Institution: Medical Biophysics, Western University, London, Ontario, Canada. Abstract: PURPOSE: To determine the effect of dose fractionation and time delay post-neoadjuvant stereotactic ablative radiotherapy (SABR) on dynamic contrast-enhanced (DCE)-MRI parameters in early stage breast cancer patients. MATERIALS AND METHODS: DCE-MRI was acquired in 17 patients pre- and post-SABR. Five patients were imaged 6-7 days post-21 Gy/1fraction (group 1), six 16-19 days post-21 Gy/1fraction (group 2), and six 16-18 days post-30 Gy/3 fractions every other day (group 3). DCE-MRI scans were performed using half the clinical dose of contrast agent. Changes in the surrounding tissue were quantified using a signal-enhancement threshold metric that characterizes changes in signal-enhancement volume (SEV). Tumour response was quantified using Ktrans and ve (Tofts model) pre- and post-SABR. Significance was assessed using a Wilcoxin signed-rank test. RESULTS: All group 1 and 4/6 group 2 patients' SEV increased post-SABR. All group 3 patients' SEV decreased. The mean Ktrans increased for group 1 by 76% (p = 0.043) while group 2 and 3 decreased 15% (p = 0.028) and 34% (p = 0.028), respectively. For ve, there was no significant change in Group 1 (p = 0.35). Groups 2 showed an increase of 24% (p = 0.043), and Group 3 trended toward an increase (23%, p = 0.08). CONCLUSION: Kinetic parameters measured 2.5 weeks post-SABR in both single fraction and three fraction groups were indicative of response but only the single fraction protocol led to enhancement in the surrounding tissue. Our results also suggest that DCE-MRI one-week post-SABR may be too early for response assessment, at least for single fraction SABR, whereas 2.5 weeks appears sufficiently long to minimize confounding acute effects. "To correct for intra-session motion, DCE images were deformably registered to the mid-time point post-contrast image using 3D Slicer v4.8.0." |
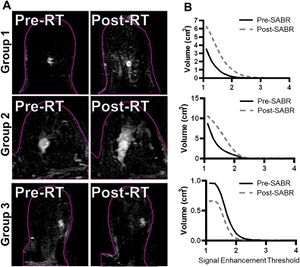 Subtraction images (A) and associated signal enhancement threshold vs volume curves (B) from representative patients for each of the three groups. The slices are approximately through the center of the tumour. The pink outline represents the skin/air interface. The single fraction images show clear evidence of an increase in the signal enhancement volume local to the tumour following SABR, and this is not seen in the 30 Gy/3 fraction patient. |
Deep Learning Approach For Automatic Out-Of-Plane Needle Localisation For Semi-Automatic Ultrasound Probe Calibration
|
Publication: Healthc Technol Lett. 2019 Dec 2;6(6):204-9. PMID: 32038858 | PDF Authors: Groves LA, VanBerlo B, Peters TM, Chen ECS. Institution: School of Biomedical Engineering, University of Western Ontario, London, Ontario, Canada. Abstract: The authors present a deep learning algorithm for the automatic centroid localisation of out-of-plane US needle reflections to produce a semi-automatic ultrasound (US) probe calibration algorithm. A convolutional neural network was trained on a dataset of 3825 images at a 6 cm imaging depth to predict the position of the centroid of a needle reflection. Applying the automatic centroid localisation algorithm to a test set of 614 annotated images produced a root mean squared error of 0.62 and 0.74 mm (6.08 and 7.62 pixels) in the axial and lateral directions, respectively. The mean absolute errors associated with the test set were 0.50 ± 0.40 mm and 0.51 ± 0.54 mm (4.9 ± 3.96 pixels and 5.24 ± 5.52 pixels) for the axial and lateral directions, respectively. The trained model was able to produce visually validated US probe calibrations at imaging depths on the range of 4-8 cm, despite being solely trained at 6 cm. This work has automated the pixel localisation required for the guided-US calibration algorithm producing a semi-automatic implementation available open-source through 3D Slicer. The automatic needle centroid localisation improves the usability of the algorithm and has the potential to decrease the fiducial localisation and target registration errors associated with the guided-US calibration method. |
Prediction of Immunohistochemistry of Suspected Thyroid Nodules by Use of Machine Learning-Based Radiomics
|
Publication: AJR Am J Roentgenol. 2019 Dec;213(6):1348-57. PMID: 31461321 Authors: Gu J, Zhu J, Qiu Q, Wang Y, Bai T, Yin Y. Institution: School of Medicine and Life Sciences, University of Jinan Shandong Academy of Medical Sciences, Jinan, Shandong, China. Abstract: OBJECTIVE. The purpose of this study was to develop and validate a radiomics model for evaluating immunohistochemical characteristics in patients with suspected thyroid nodules. MATERIALS AND METHODS. A total of 103 patients (training cohort-to-validation cohort ratio, ≈ 3:1) with suspected thyroid nodules who had undergone thyroidectomy and immunohistochemical analysis were enrolled. The immunohistochemical markers were cytokeratin 19, galectin 3, thyroperoxidase, and high-molecular-weight cytokeratin. All patients underwent CT before surgery, and a 3D Slicer was used to analyze images of the surgical specimen. Test-retest and Spearman correlation coefficient (ρ) were used to select reproducible and nonredundant features. The Kruskal-Wallis test (p < 0.05) was used for feature selection, and a feature-based model was built by support vector machine methods. The performance of the radiomic models was assessed with respect to accuracy, sensitivity, specificity, corresponding AUC, and independent validation. RESULTS. Eighty-six reproducible and nonredundant features selected from the 828 features were used to build the model. The best performance of the cytokeratin 19 model yielded accuracy of 84.4% in the training cohort and 80.0% in the validation cohort. The thyroperoxidase and galectin 3 predictive models yielded accuracies of 81.4% and 82.5% in the training cohort and 84.2% and 85.0% in the validation cohort. The performance of the high-molecular-weight cytokeratin predictive model was not good (accuracy, 65.7%) and could not be validated. CONCLUSION. A radiomics model with excellent performance was developed for individualized noninvasive prediction of the presence of cytokeratin 19, galectin 3, and thyroperoxidase based on CT images. This model may be used to identify benign and malignant thyroid nodules. |
A Review on Multiplatform Evaluations of Semi-automatic Open-source Based Image Segmentation for Cranio-maxillofacial Surgery
|
Publication: Comput Methods Programs Biomed. 2019 Dec;182:105102. PMID: 31610359 Authors: Wallner J, Schwaiger M, Hochegger K, Gsaxner C, Zemann W, Egger J. Institution: Medical University of Graz, Department of Oral and Maxillofacial Surgery, Graz, Austria. Abstract: BACKGROUND AND OBJECTIVES: Computer-assisted technologies, such as image-based segmentation, play an important role in the diagnosis and treatment support in cranio-maxillofacial surgery. However, although many segmentation software packages exist, their clinical in-house use is often challenging due to constrained technical, human or financial resources. Especially technological solutions or systematic evaluations of open-source based segmentation approaches are lacking. The aim of this contribution is to assess and review the segmentation quality and the potential clinical use of multiple commonly available and license-free segmentation methods on different medical platforms. METHODS: In this contribution, the quality and accuracy of open-source segmentation methods was assessed on different platforms using patient-specific clinical CT-data and reviewed with the literature. The image-based segmentation algorithms GrowCut, Robust Statistics Segmenter, Region Growing 3D, Otsu & Picking, Canny Segmentation and Geodesic Segmenter were investigated in the mandible on the platforms 3D Slicer, MITK and MeVisLab. Comparisons were made between the segmentation algorithms and the ground truth segmentations of the same anatomy performed by two clinical experts (n = 20). Assessment parameters were the Dice Score Coefficient (DSC), the Hausdorff Distance (HD), and Pearsons correlation coefficient (r). RESULTS: The segmentation accuracy was highest with the GrowCut (DSC 85.6%, HD 33.5 voxel) and the Canny (DSC 82.1%, HD 8.5 voxel) algorithm. Statistical differences between the assessment parameters were not significant (p < 0.05) and correlation coefficients were close to the value one (r > 0.94) for any of the comparison made between the segmentation methods and the ground truth schemes. Functionally stable and time-saving segmentations were observed. CONCLUSION: High quality image-based semi-automatic segmentation was provided by the GrowCut and the Canny segmentation method. In the cranio-maxillofacial complex, these segmentation methods provide algorithmic alternatives for image-based segmentation in the clinical practice for e.g. surgical planning or visualization of treatment results and offer advantages through their open-source availability. This is the first systematic multi-platform comparison that evaluates multiple license-free, open-source segmentation methods based on clinical data for the improvement of algorithms and a potential clinical use in patient-individualized medicine. The results presented are reproducible by others and can be used for clinical and research purposes. |
Patellar Calcar: Morphometric Analysis by Knee Magnetic ResonanceImaging and Three-dimensional Reconstruction Software-assisted
|
Publication: Surg Radiol Anat. 2019 Dec;41(12):1483-8. PMID: 31529166 Authors: Benavente S, Villagra J. Institution: Universidad Católica de la Santísima Concepción, Concepción, Chile. Abstract: PURPOSE: Patellar calcar corresponds to a greater trabecular bone density area in the patella lateral facet, whose morphometry is uncertain. This study aimed to describe patellar calcar morphometry by knee MRI and develop a 3D reconstruction software-assisted. MATERIALS AND METHODS: Consecutive adult patients, submitted to knee MRI, between 2014 and 2017, were entered in IMPAX software. Exclusion criteria are history of patellar surgical intervention, trauma, chondromalacia, bone edema or bipartite patella. All MRI images were retrospectively reviewed by three readers. MRI patellar calcar measurements are height, width, thickness and posterior distance. 3D model protocol reconstruction: 3D Slicer software was used to design a preliminary model for each patient, and then all were automatically merged into one, which was finalized using the software segmentation tools. For 3D patellar calcar location, the transpolar axis was designed. RESULTS: 250 MRI were analyzed, patellar calcar was present in 208 (83.2%); 101 men and 107 women. Mean age was 44.3 ± 15.6 years. MEASUREMENTS: height 13.84 ± 2.42 mm (male: 14.50 ± 2.42; female: 13.21 ± 2.26) (p < 0.0001), width 12.21 ± 2.26 mm (male 13.14 ± 2.22; female 11.33 ± 1.93) (p < 0.0001). No statistically significant difference of thickness 0.56 ± 0.22 mm (male: 0.56 ± 0.25; female: 0.56 ± 0.20) and posterior distance 2.37 ± 0.80 mm (male: 2.46 ± 0.89; female: 2.29 ± 0.69) between genders was found. 3D model results: transpolar axis went through the patellar calcar in all the cases. CONCLUSIONS: This study shows in a 3D model reconstruction, what was previously described in the literature, determining for the first time the patellar calcar morphometry in the knee MRI and identifying it as a regular finding in this imaging test. |
Deep Convolutional Neural Networks for Automatic Detection of Orbital Blowout Fractures
|
Publication: J Craniofac Surg. 2019 Dec 13. PMID: 31842071 Authors: Li L, Song X, Guo Y, Liu Y, Sun R, Zou H, Zhou H, Fan X1. Institution: Department of Ophthalmology, Ninth People's Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai. Abstract: Orbital blow out fracture is a common disease in emergency department and a delay or failure in diagnosis can lead to permanent visual changes. This study aims to evaluate the ability of an automatic orbital blowout fractures detection system based on computed tomography (CT) data.Orbital CT scans of adult orbital blowout fractures patients and normal cases were obtained from Shanghai Ninth People's Hospital between January and March 2017. The region of fractures was annotated using 3D Slicer. The Inception V3 convolutional neural networks were constructed utilizing the Python programming language with PyTorch as the framework to extract high dimension features from each slice in a CT scan. These extracted features are processed through a XGBoost model to make the final differentiation of fracture cases and nonfracture ones. Accuracy, receiver operating characteristics, and area under the curve were evaluated.This study used 94 CT scans diagnosed with orbital blowout fractures and 94 healthy control cases. The automatic detection system showed accuracy of 92% in single-image classification and 87% in patient level detection. The area under the receiver operating characteristic curve was 0.9574.Using a deep learning-based automatic detection system of orbital blowout fracture can accurately detect and classify orbital blowout fractures from CT scans. The convolutional neural networks model combined with an accurate annotation system could achieve good performance in a small dataset. Further studies with large and multicenter data are required to refine this technology for possible clinical applications. |
Decision-making Based on 3D Printed Models in Laparoscopic Liver Resections with Intraoperative Ultrasound: A Prospective Observational Study
|
Publication: Eur Radiol. 2019 Nov 26. PMID: 31773294 Authors: Witowski J, Budzyński A, Grochowska A, Ballard DH, Major P, Rubinkiewicz M, Złahoda-Huzior A, Popiela TJ, Wierdak M, Pędziwiatr M. Institution: 2nd Department of General Surgery, Jagiellonian University Medical College, Krakow, Poland. Abstract: OBJECTIVES: The aim of this study was to evaluate impact of 3D printed models on decision-making in context of laparoscopic liver resections (LLR) performed with intraoperative ultrasound (IOUS) guidance. METHODS: Nineteen patients with liver malignances (74% were colorectal cancer metastases) were prospectively qualified for LLR or radiofrequency ablation in a single center from April 2017 to December 2018. Models were 3DP in all cases based on CT and facilitated optical visualization of tumors' relationships with portal and hepatic veins. Planned surgical extent and its changes were tracked after CT analysis and 3D model inspection, as well as intraoperatively using IOUS. RESULTS: Nineteen patients were included in the analysis. Information from either 3DP or IOUS led to changes in the planned surgical approach in 13/19 (68%) patients. In 5/19 (26%) patients, the 3DP model altered the plan of the surgery preoperatively. In 4/19 (21%) patients, 3DP independently changed the approach. In one patient, IOUS modified the plan post-3DP. In 8/19 (42%) patients, 3DP model did not change the approach, whereas IOUS did. In total, IOUS altered surgical plans in 9 (47%) cases. Most of those changes (6/9; 67%) were caused by detection of additional lesions not visible on CT and 3DP. CONCLUSIONS: 3DP can be helpful in planning complex and major LLRs and led to changes in surgical approach in 26.3% (5/19 patients) in our series. 3DP may serve as a useful adjunct to IOUS. KEY POINTS: • 3D printing can help in decision-making before major and complex resections in patients with liver cancer. • In 5/19 patients, 3D printed model altered surgical plan preoperatively. • Most surgical plan changes based on intraoperative ultrasonography were caused by detection of additional lesions not visible on CT and 3D model. Segmentation was performed in 3D Slicer. |
In-House Surgeon-Led Virtual Surgical Planning for Maxillofacial Reconstruction
|
Publication: J Oral Maxillofac Surg. 2019 Nov 21. PMID: 31843280 Authors: Abo Sharkh H, Makhoul N. Institution: Maxillofacial Oncology and Microvascular Reconstruction, Department of Oral and Maxillofacial Surgery, McGill University Health Centre, Montreal, QC, Canada. Abstract: PURPOSE: Virtual surgical planning (VSP) and custom fabricated cutting guides for maxillofacial reconstruction have been shown to improve the accuracy of bony reconstruction and overall surgical efficiency and decrease the ischemia time. Our aim was to describe an in-house VSP technique for maxillofacial reconstructive procedures. MATERIALS AND METHODS: We used 2 free software applications. 3D Slicer was used to extract the bones of interest for the recipient and the donor sites from the computed tomography scan's DICOM (digital imaging and communications in medicine) data. The Autodesk Meshmixer (Autodesk Inc, San Rafael, CA) was used to perform VSP and fabrication of the cutting guides. A reconstructed jaw model was printed in-house using a commercially available fused deposition modeling-based desktop 3-dimensional (3D) printer (Qidi Technology, Zhejiang, China) and used to prebend the reconstruction plate. The cutting guides were printed using a commercially available resin-based stereolithography apparatus desktop 3D printer (Form 2, Dental SG Resin; Formlabs, Somerville, MA) to allow for sterilization of the guides. We performed this technique for 19 consecutive patients with maxillofacial benign or malignant tumors requiring microvascular bony reconstruction. We calculated the average time and associated costs using this in-house VSP technique. RESULTS: The technique was found to be simple and repeatable. The average time required for VSP was 158 minutes (2 hours, 38 minutes). The average cost for printing the reconstructed model per case was $5.21 Canadian dollars (CAD), and the average cost for printing the cutting guides per case was $12.80 CAD. CONCLUSIONS: Using this technique, in-house VSP and 3D printing can be performed by the treating surgeon, without an engineering background, within a reasonable period. |
Patient-specific Access Planning in Minimally Invasive Mitral Valve Surgery
|
Publication: Med Hypotheses. 2019 Nov 14;136:109475. PMID: 31812012 Authors: Di Perna D, Castro M, Gasc Y, Haigron P, Verhoye JP, Anselmi A. Institution: University of Rennes, France. Abstract: BACKGROUND: Minimally invasive mitral valve repair or replacement (MIMVR) approaches have been increasingly adopted for the treatment of mitral regurgitation, allowing a shorter recovery time and improving postoperative quality of life. However, inadequate positioning of the right mini thoracotomy access (working port) translates into suboptimal exposure, prolonged operative times and, potentially, reduction in the quality of mitral repair. At present, we are missing tools to further improve the positioning of the working port in order to ameliorate surgical exposure in a patient- specific fashion. METHODS AND EVALUATION OF THE HYPOTHESIS: We hypothesized that computation of relevant anatomical measurements from preoperative CT scans in patients undergoing MIMVR may provide patient-specific information in order to propose the surgical access that best fits to the patient's morphology. We hypothesized that this may systematize optimal mitral valve exposure, facilitating the procedure and potentially ameliorating the outcomes. We also hypothesized that preoperative simulation of the working port site and surgical instruments' insertion using a three-dimensional virtual model of the patient is feasible and may help in the customization of ports positioning. The hypothesis was evaluated by a multidisciplinary team including cardiac surgeons, experts in medical image processing and biomedical engineers. CT scans of 14 patients undergoing MIMVR were segmented to visualize 3D chest bones and heart structures meshes. The mitral valve annulus is pointed manually by the expert or extracted automatically when contrast-enhanced CT scan was available. The valve plane was then calculated and the optimal incision location analyzed according to a) the perpendicularity and b) the distance between the intercostal spaces and the valve plane. An angle-chart representation for the 4th, 5th and 6th intercostal spaces and a color map illustrating the distance between the skin and the mitral valve were created. We started the development of a simulation tool for preoperative planning using 3D Slicer software. CONCLUSIONS: Several patient-specific factors (including the orientation of the mitral valve plane and the morphology of the chest cage) may influence the performance of a MIMVR procedure, but they are not quantitatively considered in the current planning strategy. We suggest that the clinical results of MIMVR can be improved through preoperative virtual simulation and computer-assisted surgery (through determination of working port and surgical instruments insertion positioning). Further research is justified and the development of a software tool for clinical evaluation is warranted to verify the current hypothesis. |
Semi-Automatic Signature-Based Segmentation Method for Quantification of Neuromelanin in Substantia Nigra
|
Publication: Brain Sci. 2019 Nov 22;9(12). PMID: 31766668 | PDF Authors: Gašper Zupan, Dušan Šuput, Zvezdan Pirtošek, Andrej Vovk. Institution: Faculty of Medicine, University of Ljubljana, Ljubljana, Slovenia. Abstract: In Parkinson's disease (PD), there is a reduction of neuromelanin (NM) in the substantia nigra (SN). Manual quantification of the NM volume in the SN is unpractical and time-consuming; therefore, we aimed to quantify NM in the SN with a novel semi-automatic segmentation method. Twenty patients with PD and twelve healthy subjects (HC) were included in this study. T1-weighted spectral pre-saturation with inversion recovery (SPIR) images were acquired on a 3T scanner. Manual and semi-automatic atlas-free local statistics signature-based segmentations measured the surface and volume of SN, respectively. Midbrain volume (MV) was calculated to normalize the data. Receiver operating characteristic (ROC) analysis was performed to determine the sensitivity and specificity of both methods. PD patients had significantly lower SN mean surface (37.7 ± 8.0 vs. 56.9 ± 6.6 mm2) and volume (235.1 ± 45.4 vs. 382.9 ± 100.5 mm3) than HC. After normalization with MV, the difference remained significant. For surface, sensitivity and specificity were 91.7 and 95 percent, respectively. For volume, sensitivity and specificity were 91.7 and 90 percent, respectively. Manual and semi-automatic segmentation methods of the SN reliably distinguished between PD patients and HC. ROC analysis shows the high sensitivity and specificity of both methods. "...Manual delineation of the SN in mesencephalon was performed with 3D Slicer, v4.9.0." |
Spatial Accuracy of a Clinically Established Noninvasive Electrocardiographic Imaging System for the Detection of Focal Activation in an Intact Porcine Model
|
Publication: Circ Arrhythm Electrophysiol. 2019 Nov;12(11):e007570. PMID: 31707808 Authors: Hohmann S, Rettmann ME, Konishi H, Borenstein A, Wang S, Suzuki A, Michalak GJ, Monahan KH, Parker KD, Newman LK, Packer DL. Institution: Translational Interventional Electrophysiology Laboratory, Mayo Clinic, Rochester, MN, USA. Abstract: BACKGROUND: Noninvasive electrocardiographic imaging (ECGi) is used clinically to map arrhythmias before ablation. Despite its clinical use, validation data regarding the accuracy of the system for the identification of arrhythmia foci is limited. METHODS: Nine pigs underwent closed-chest placement of endocardial fiducial markers, computed tomography, and pacing in all cardiac chambers with ECGi acquisition. Pacing location was reconstructed from biplane fluoroscopy and registered to the computed tomography using the fiducials. A blinded investigator predicted the pacing location from the ECGi data, and the distance to the true pacing catheter tip location was calculated. RESULTS: A total of 109 endocardial and 9 epicardial locations were paced in 9 pigs. ECGi predicted the correct chamber of origin in 85% of atrial and 92% of ventricular sites. Lateral locations were predicted in the correct chamber more often than septal locations (97% versus 79%, P=0.01). Absolute distances in space between the true and predicted pacing locations were 20.7 (13.8-25.6) mm (median and [first-third] quartile). Distances were not significantly different across cardiac chambers. CONCLUSIONS: The ECGi system is able to correctly identify the chamber of origin for focal activation in the vast majority of cases. Determination of the true site of origin is possible with sufficient accuracy with consideration of these error estimates. "3D Slicer v.4.10.0 was used to integrate the CT imaging and 3D model data, and for visualization purposes." |
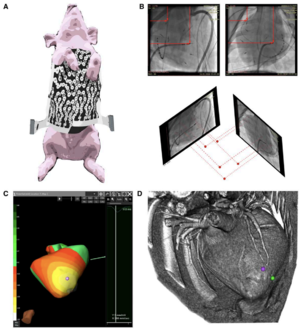 Study and analysis workflow. A, Animal with ECG acquisition vest. B, Geometric reconstruction of fiducial marker and catheter tip positions in 3D from corresponding biplane fluoroscopy images. Top, Measurements of clip and catheter positions in a set of biplane fluoroscopy images. This is shown for one clip and the catheter tip only but was performed for all clips and the catheter tip. Bottom, Reconstruction of a 3D point cloud from biplane fluoroscopy images. Each point (red sphere) is located at the intersection of its two projection vectors shown as red dashed lines. C, Electrocardiographic imaging (ECGi) map. ECGi information represented as isopotential lines on the segmented ventricular surface, from −2.8 mV (white) to +3.66 mV (dark green). The location of the earliest negativity has been marked by the purple marker. D, Corresponding volume rendered computed tomography (CT), oriented in the same caudal right posterior oblique view as (C). The inferior aspect of the heart is shown. The purple marker corresponds to the purple marker in (C). The green marker identifies the true pacing location as reconstructed from fluoroscopy (near the RV apex). Euclidean distance between the 2 points in space was 17.9 mm in this case. |
Life Without a Brain: Neuroradiological and Behavioral Evidence of Neuroplasticity Necessary to Sustain Brain Function in the Face of Severe Hydrocephalus
|
Publication: J Clin Exp Dent. 2019 Dec 1;11(12):e1099-e1108. PMID: 31712649 | PDF Authors: Ferris CF, Cai X, Qiao J, Switzer B, Baun J, Morrison T, Iriah S, Madularu D, Sinkevicius KW, Kulkarni P. Institution: Center for Translational NeuroImaging, Northeastern University, Boston, MA, USA. Abstract: A two-year old rat, R222, survived a life-time of extreme hydrocephaly affecting the size and organization of its brain. Much of the cortex was severely thinned and replaced by cerebrospinal fluid, yet R222 had normal motor function, could hear, see, smell, and respond to tactile stimulation. The hippocampus was malformed and compressed into the lower hindbrain together with the hypothalamus midbrain and pons, yet R222 showed normal spatial memory as compared to age-matched controls. BOLD MRI was used to study the reorganization of R222's brain function showing global activation to visual, olfactory and tactile stimulation, particularly in the brainstem/cerebellum. The results are discussed in the context of neuroadaptation in the face of severe hydrocephaly and subsequent tissue loss, with an emphasis on what is the "bare minimum" for survival. "...Brain tissue masks were manually drawn using 3D Slicer and applied for skull-stripping" |
a Case Report of Total Skin Photon Radiation Therapy for Cutaneous Epitheliotropic Lymphoma in a Dog
|
Publication: BMC Vet Res. 2019 Nov 9;15(1):407.. PMID: 31706321 | PDF Authors: Deveau MA, Sutton M, Baetge C, Diesel AB. Institution: Department of Small Animal Clinical Sciences, College of Veterinary Medicine & Biomedical Sciences, Texas A&M University, TX, USA. Abstract: BACKGROUND: Total skin electron beam radiation therapy (TSEBT) is an effective treatment for primary diffuse cutaneous lymphomas in humans. While several techniques exist, they all require significant commitment of staff time and resources. In veterinary medicine, canine-specific techniques and strategies have been adapted and delivered but deemed not "realistically" clinically implementable given the time commitment of over 2.5 h plus per fraction or have been relegated to palliative intent. Leveraging these technologies of helical tomotherapy and 3D printing, we developed and clinically implemented a radiotherapeutic treatment strategy for the management of medically refractory diffuse cutaneous lymphoma in the dog. CASE PRESENTATION: A 13.5-year-old female spayed Bichon Frise presented to the Oncology service at Texas A&M University, College of Veterinary Medicine due to the progression of diffuse cutaneous epitheliotropic lymphoma (CEL) that had failed medical management. Twenty-seven gray were delivered to the patient with a treatment time requirement under 40 min including real time monitoring of anesthesia during setup and treatment. A partial response was noticeable after four fractions and the tumor completely regressed progressively over the entire treated area by the end of therapy. A grade 1 lethargy, fatigue, weight loss, and oral mucositis and grade 2 alopecia, nail/claw changes, pruritus, scaling, anorexia, and diarrhea were noted during treatment. Additionally, a grade 3 thrombocytopenia developed after fraction eight requiring a treatment interruption of 6 weeks and prescription modification prior to treatment continuation and completion. From the beginning of total skin photon radiation therapy (TSPT) treatment until the time of the patient was euthanized unrelated to cutaneous epitheliotropic lymphoma (123 days), only one new lesion on the head was identified and confirmed by histopathology within the treated fields. CONCLUSIONS: The proposed technique is an acceptable alternative to TSEBT that is actually clinically implementable within a palliative or definitive setting and clinical constraints, however further testing and refinement is needed to reduce hematological complications and to confirm and expand on preliminary findings. "...To prepare the 3D mold scaffold for final printing, the 3D mold contour was then transformed into a 3D mesh using 3D Slicer" |
Considerations for Ultrasound Exposure During Transcranial MR Acoustic Radiation Force Imaging
|
Publication: Sci Rep. 2019 Nov 7;9(1):16235. PMID: 31700021 | PDF Authors: Phipps MA, Jonathan SV, Yang PF, Chaplin V, Chen LM, Grissom WA, Caskey CF. Institution: Vanderbilt University Medical Center Department of Radiology and Radiological Sciences, Nashville, TN, USA. Abstract: The aim of this study was to improve the sensitivity of magnetic resonance-acoustic radiation force imaging (MR-ARFI) to minimize pressures required to localize focused ultrasound (FUS) beams, and to establish safe FUS localization parameters for ongoing ultrasound neuromodulation experiments in living non-human primates. We developed an optical tracking method to ensure that the MR-ARFI motion-encoding gradients (MEGs) were aligned with a single-element FUS transducer and that the imaged slice was prescribed at the optically tracked location of the acoustic focus. This method was validated in phantoms, which showed that MR-ARFI-derived displacement sensitivity is maximized when the MR-ARFI MEGs were maximally aligned with the FUS propagation direction. The method was then applied in vivo to acquire displacement images in two healthy macaque monkeys (M fascicularis) which showed the FUS beam within the brain. Temperature images were acquired using MR thermometry to provide an estimate of in vivo brain temperature changes during MR-ARFI, and pressure and thermal simulations of the acoustic pulses were performed using the k-Wave package which showed no significant heating at the focus of the FUS beam. The methods presented here will benefit the multitude of transcranial FUS applications as well as future human applications. "...fiducials are manually identified in the T1-weighted image stack using 3D Slicer." Funding:
|
Direct Evidence for Eudicot Pollen-feeding in a Cretaceous Stinging Wasp (Angiospermae; Hymenoptera, Aculeata) Preserved in Burmese Amber
|
Publication: Commun Biol. 2019 Nov 7;2:408. PMID: 31728419 | PDF Authors: Grimaldi DA, Peñalver E, Barrón E, Herhold HW, Engel MS. Institution: American Museum of Natural History, New York, NY, USA. Abstract: Angiosperms and their insect pollinators form a foundational symbiosis, evidence for which from the Cretaceous is mostly indirect, based on fossils of insect taxa that today are anthophilous, and of fossil insects and flowers that have apparent anthophilous and ento- mophilous specializations, respectively. We present exceptional direct evidence preserved in mid-Cretaceous Burmese amber, 100 mya, for feeding on pollen in the eudicot genus Tri- colporoidites by a basal new aculeate wasp, Prosphex anthophilos, gen. et sp. nov., in the lineage that contains the ants, bees, and other stinging wasps. Plume of hundreds of pollen grains wafts from its mouth and an apparent pollen mass was detected by micro-CT in the buccal cavity: clear evidence that the wasp was foraging on the pollen. Eudicots today comprise nearly three-quarters of all angiosperm species. Prosphex feeding on Tricolporoidites supports the hypothesis that relatively small, generalized insect anthophiles were important pollinators of early angiosperms. Post-processing and analysis performed in 3D Slicer. |
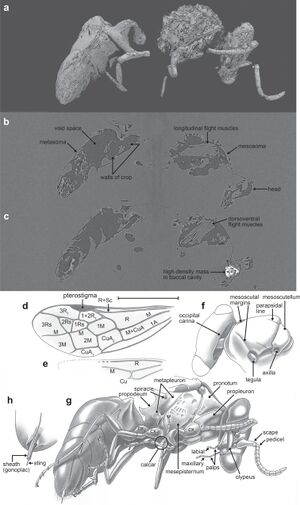 CT images and illustrations of the holotype of P. anthophilos, new genus, new species, AMNH Bu-KL18-31. a CT image, external surface. Portions of thin and/or distorted cuticle are missing, particularly in the mid-section. b, c Two CT slices through (a), showing the longitudinal (b) and dorsoventral flight muscles (c), walls of the crop (b), and a high-density mass in the buccal cavity (c), likely a pollen mass that is partially pyritized. d–h Illustrated rendering of Prosphex, showing the forewing (d, with conventional abbreviations for wing veins and cells) and the hind wing (e) (slightly reconstructed), dorsal view of head and thorax (f), left ventrolateral habitus (g: cx, coxa) and detail of sting (h). Body of the wasp is rendered as preserved, without reconstruction. All images to the same scale (scale line 1.0 mm); h is slightly magnified |
Myoarchitectural Disarray of Hypertrophic Cardiomyopathy Begins Pre-Birth
|
Publication: J Anat. 2019 Nov;235(5):962-76. PMID: 31347708 | PDF Authors: Garcia-Canadilla P, Cook AC, Mohun TJ, Oji O, Schlossarek S, Carrier L, McKenna WJ, Moon JC, Captur G. Institution: Institute of Cardiovascular Science, University College London, London, UK. Abstract: Myoarchitectural disarray - the multiscalar disorganisation of myocytes, is a recognised histopathological hallmark of adult human hypertrophic cardiomyopathy (HCM). It occurs before the establishment of left ventricular hypertrophy (LVH) but its early origins and evolution around the time of birth are unknown. Our aim is to investigate whether myoarchitectural abnormalities in HCM are present in the fetal heart. We used wild-type, heterozygous and homozygous hearts (n = 56) from a Mybpc3-targeted knock-out HCM mouse model and imaged the 3D micro-structure by high-resolution episcopic microscopy. We developed a novel structure tensor approach to extract, display and quantify myocyte orientation and its local angular uniformity by helical angle, angle of intrusion and myoarchitectural disarray index, respectively, immediately before and after birth. In wild-type, we demonstrate uniformity of orientation of cardiomyocytes with smooth transitions of helical angle transmurally both before and after birth but with traces of disarray at the septal insertion points of the right ventricle. In comparison, heterozygous mice free of LVH, and homozygous mice showed not only loss of the normal linear helical angulation transmural profiles observed in wild-type but also fewer circumferentially arranged myocytes at birth. Heterozygous and homozygous showed more disarray with a wider distribution than in wild-type before birth. In heterozygous mice, disarray was seen in the anterior, septal and inferior walls irrespective of stage, whereas in homozygous mice it extended to the whole LV circumference including the lateral wall. In conclusion, myoarchitectural disarray is detectable in the fetal heart of an HCM mouse model before the development of LVH "...3D volume‐rendered reconstructions were performed in 3D Slicer." |
Whole Lesion Histogram Analysis of Apparent Diffusion Coefficients on MRI Predicts Disease-free Survival in Locally Advanced Squamous Cell Cervical Cancer after Radical Chemo-radiotherapy
|
Publication: BMC Cancer. 2019 Nov 15;19(1):1115. PMID: 31729974 | PDF Authors: Zhao B, Cao K, Li XT, Zhu HT, Sun YS. Institution: Department of Radiology, Key Laboratory of Carcinogenesis and Translational Research, Ministry of Education, Peking University Cancer Hospital and Institute, Beijing, China. Abstract: BACKGROUND: The aim was to investigate the prognostic value of MR apparent diffusion coefficients (ADC) using histogram analysis (HA) in predicting disease-free survival (DFS) of cervical cancer after chemo-radiation therapy. METHODS: We retrospectively analyzed 103 women with pathologically proven squamous cell uterine cancer who received chemo-radiation therapy between 2009 and 2013. All patients were followed up for more than 2 years. Pre-treatment MR images were retrieved and imported for HA using an in-house developed software program based on 3D Slicer. Regions of interest of whole tumors were drawn manually on DWI with reference to T2WI. HA features (mean, max, min, 50, 10, 90%, kurtosis, and skewness) were extracted from apparent diffusion coefficient (ADC) maps and compared between the recurrence and non-recurrence groups after the 2-year follow-up. Univariate and multivariate Cox regression analysis was used to correlate ADC HA features and relevant clinical variables (age, grade, maximal diameter of tumor, FIGO stage, SCC-Ag) with DFS. RESULTS: One hundred three patients with stage IB-IV cervical cancers were followed up for 2.0-94.6 months (median 48.9 months). Twenty patients developed recurrence within 2 years. In the recurrence group, the min (P = 0.001) and 10% (P = 0.048) ADC values were significantly lower than those of the non-recurrence group. Univariate and multivariate Cox regression analysis revealed that ADCmin (P = 0.006, HR = 0.110) was significantly correlated with DFS. CONCLUSION: Pre-treatment volumetric ADCmin in histogram analysis is an independent factor that is correlated with DFS in cervical cancer patients treated with chemo-radiation therapy. |
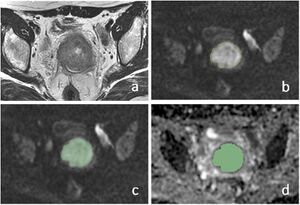 Manual segmentation of ROIs in cervical lesion and schematic diagram of parameters. a–c Referring to T2WI and DWI, ROIs were drawn manually slice by slice on DWI images along the edge of the lesions in order to cover as much tumor area as possible without excluding cystic or necrotic areas. d The same ROIs were registered to ADC maps. |
Variations in the Size and Shape of Human Cochlear Malformation Types
|
Publication: Anat Rec (Hoboken). 2019 Oct;302(10):1792-9. PMID: 30980504 | PDF Authors: Dhanasingh A. Institution: MED-EL GmbH, Innsbruck, Austria. Abstract: The objective of this study is to determine the variations in size and shape of the most widely recognized cochlear malformation types using three-dimensional (3D) visualization. Using 3D Slicer freeware, the complete inner-ear structures were segmented from 46 anonymized high-resolution computed tomography (HRCT) image datasets. Cochlear height, internal auditory canal height, and width were measured from the axial plane. Cochlear basal turn diameter was measured from the oblique coronal plane. Number of cochlear turns was measured from the 3D images and the corresponding cochlear duct length (CDL) was estimated using the CDL equations given in Alexiades et al. [Otol Neurotol 36 (2015) 904-907]. Out of 46 preoperative HRCT image datasets of human temporal bone, cochlear anatomy types including normal anatomy (4), enlarged vestibular aqueduct syndrome (3), cochlear aplasia (2), incomplete partition Types I (8), II (Mondini's deformity) (3), and III (X-linked) (4), cochlear hypoplasia (CH) (17), and common cavity (CC) (5) were identified. Majority of CH cases had cochlear height shorter than 4 mm whereas the CC cases measured cochlear height above 6 mm. For all the other malformation types, cochlear height was between 4 and 6 mm. In terms of "A" value, majority of CH cases showed shorter "A" value of <7.5 mm, which is in the lower end in comparison to the rest of the malformation types reported in this study. 3D-visualization shows the size and shape variations of all the structures of inner ear and also improves the clinicians' ability to visualize cochlear anatomy and nearby structures much easier than from the 2D image slices. |
Pretreatment Prediction of Adaptive Radiation Therapy Eligibility Using MRI-Based Radiomics for Advanced Nasopharyngeal Carcinoma Patients
|
Publication: Sci Rep. 2019 Oct 24;9(1):15267. PMID: 31649259 | PDF Authors: Yu TT, Lam SK, To LH, Tse KY, Cheng NY, Fan YN, Lo CL, Or KW, Chan ML, Hui KC, Chan FC, Hui WM, Ngai LK, Lee FK, Au KH, Yip CW, Zhang Y, Cai J. Institution: Department of Health Technology and Informatics, Hong Kong Polytechnic University, Hung Hom, Hong Kong. Abstract: Background and purpose: Adaptive radiotherapy (ART) can compensate for the dosimetric impacts induced by anatomic and geometric variations in patients with nasopharyngeal carcinoma (NPC); Yet, the need for ART can only be assessed during the radiation treatment and the implementation of ART is resource intensive. Therefore, we aimed to determine tumoral biomarkers using pre-treatment MR images for predicting ART eligibility in NPC patients prior to the start of treatment. Methods: Seventy patients with biopsy-proven NPC (Stage II-IVB) in 2015 were enrolled into this retrospective study. Pre-treatment contrast-enhanced T1-w (CET1-w), T2-w MR images were processed and filtered using Laplacian of Gaussian (LoG) filter before radiomic features extraction. A total of 479 radiomics features, including the first-order (n = 90), shape (n = 14), and texture features (n = 375), were initially extracted from Gross-Tumor-Volume of primary tumor (GTVnp) using CET1-w, T2-w MR images. Patients were randomly divided into a training set (n = 51) and testing set (n = 19). The least absolute shrinkage and selection operator (LASSO) logistic regression model was applied for radiomic model construction in training set to select the most predictive features to predict patients who were replanned and assessed in the testing set. A double cross-validation approach of 100 resampled iterations with 3-fold nested cross-validation was employed in LASSO during model construction. The predictive performance of each model was evaluated using the area under the receiver operator characteristic (ROC) curve (AUC). Results: In the present cohort, 13 of 70 patients (18.6%) underwent ART. Average AUCs in training and testing sets were 0.962 (95%CI: 0.961-0.963) and 0.852 (95%CI: 0.847-0.857) with 8 selected features for CET1-w model; 0.895 (95%CI: 0.893-0.896) and 0.750 (95%CI: 0.745-0.755) with 6 selected features for T2-w model; and 0.984 (95%CI: 0.983-0.984) and 0.930 (95%CI: 0.928-0.933) with 6 selected features for joint T1-T2 model, respectively. In general, the joint T1-T2 model outperformed either CET1-w or T2-w model alone. Conclusions: Our study successfully showed promising capability of MRI-based radiomics features for pre-treatment identification of ART eligibility in NPC patients. "...all MR images were processed using 3D Slicer v4.11.0." |
Volumetric Assessment of the Dental Crown for Sex Estimation by Means of Cone-beam Computed Tomography
|
Publication: Forensic Sci Int. 2019 Oct;303:109920. PMID: 31442711 Authors: Manhaes-Caldas D, Oliveira ML, Groppo FC, Haiter-Neto F. Institution: Department of Oral Diagnosis, Division of Oral Radiology, Piracicaba Dental School, University of Campinas, São Paulo, Brazil. Abstract: Sex estimation has a vital role in the solution of forensic cases when the identification of a large number of victims is needed. Considering the sexual dimorphism of the human teeth, the objective of this study was to estimate human sex by means of cone beam computed tomography (CBCT)-based volumetric assessment of the dental crown. A total of 78 CBCT images of the upper central incisors, upper and lower canines, and lower lateral incisors were equally selected from a Brazilian population aged between 8 and 36 years old. The dental crowns were subjected to image-based volumetric assessment by manual segmentation using the 3D Slicer software, and the outcomes were compared by the Mann-Whitney test, unpaired t-test, Pearson correlation test, conditional backward stepwise logistic regression and intraclass correlation coefficient (α=0.05). The volumetric accuracy of the upper central incisor, upper canine and lower canine for sex estimation were 64.1%, 74.4% and 79.5%, respectively. The combined analysis of the upper and lower canines allowed an average accuracy of 83.7%. In conclusion, the combined volumetric analysis of the crown of the upper and lower canines can be applied for sex estimation in the studied population. |
Association of Serum Cystatin C with White Matter Abnormalities in Patients with Amnestic Mild Cognitive Impairment
|
Publication: Geriatr Gerontol Int. 2019 Oct;19(10):1036-40. PMID: 31489777 | PDF Authors: Hirao K, Yamashita F, Tsugawa A, Haime R, Fukasawa R, Sato T, Okita M, Shimizu S, Kanetaka H, Umahara T, Sakurai H, Hanyu H. Institution: Department of Geriatric Medicine, Tokyo Medical University, Tokyo, Japan. Abstract: AIM: White matter hyperintensities (WMH) on MRI have been reported to be a risk factor for the conversion from mild cognitive impairment (MCI) to Alzheimer's disease, although the reason remains unclear. In the present study, we hence investigated the associations between WMH volumes and cognitive function, blood levels of various molecules, and the presence of lifestyle-associated diseases in patients with amnestic MCI. METHODS: The initial data of 38 patients with amnestic MCI and 10 normal control individuals were analyzed. The volumes of periventricular hyperintensities (PVH) and deep WMH (DWMH) were measured on T2 fluid-attenuated inversion recovery using the imaging software, 3D Slicer; and the association between PVH/DWMH volumes and cognitive function, blood levels of molecules (such as cystatin C [CysC], 25-hydroxyvitamin D and homocysteine) and the presence of lifestyle-associated diseases (such as hypertension, hyperlipidemia and diabetes mellitus) were analyzed. RESULTS: In the MCI group, the PVH volume : intracranial volume ratio significantly correlated with Trail Making Test-A/B scores and CysC level by Pearson's analysis, and the PVH volume : intracranial volume ratio significantly correlated with only CysC levels, whereas the DWMH volume : intracranial volume ratio did not correlate with any items at all by linear multiple regression analysis. CONCLUSIONS: PVH volume was closely associated with frontal lobe dysfunction, particularly with attention and executive dysfunction. Serum CysC level was associated with PVH volume, which suggests that CysC might be a useful marker for determining treatment strategies for white matter abnormalities in amnestic MCI. |
Three-dimensional Clavicle Displacement Analysis and its Effect on ScapularPosition in Acute Clavicle Midshaft Fracture
|
Publication: J Shoulder Elbow Surg. 2019 Oct;28(10):1877-85. PMID: 31272891 Authors: Kim JH, Gwak HC, Kim CW, Lee CR, Kim YJ, Seo HW. Institution: Department of Orthopedic Surgery, Inje University Busan Paik Hospital, Inje University College of Medicine, Busan, Republic of Korea. Abstract:BACKGROUND: The purpose of this study was to measure the distance of the clavicle in 3 dimensions (3D) and each direction (anterior to posterior, medial to lateral, and superior to inferior) and to analyze the correlation of the angular orientation of the scapula according to each directional distance of the clavicle. METHODS: Sixty-seven patients with Robinson 2B1 and 2B2 clavicle midshaft fracture (46.0 ± 17.4 years, men = 50, women = 17) were selected as final subjects. Patients' computed tomography was reconstructed using an image processing program (3D Slicer v.4.3 software). Anteroposterior (AP) distance, medial-to-lateral distance, superior-to-inferior distance, and 3D distance of both clavicles were measured. The plane connecting the 3 points (superior pole, inferior pole, and center of glenoid) of the scapula was used to calculate differences in the angular orientation between both scapulae. RESULTS: Among each directional distance of the clavicle, only the AP distance showed negative correlation with scapular angular orientation with anterior tilting, internal rotation, and upward rotation of the scapula (Pearson's correlation coefficient: -0.68, -0.24, and -0.28; P < .001, P = .048, and P = .021). CONCLUSION: The shortening of the AP distance of the clavicle was related to the angular orientation of the scapula in acute clavicle fracture. AP shortening should be considered when determining the treatment of clavicle fracture. |
The Evolutionary Radiation of Hominids: a Phylogenetic Comparative Study
|
Publication: Sci Rep. 2019 Oct 24;9(1):15267. PMID: 31649259 | PDF Authors: Garcia-Canadilla P, Cook AC, Mohun TJ, Oji O, Schlossarek S, Carrier L, McKenna WJ, Moon JC, Captur G. Institution: Institute of Cardiovascular Science, University College London, London, UK. Abstract: Over the last 150 years the diversity and phylogenetic relationships of the hominoids have been one of the main focuses in biological and anthropological research. Despite this, the study of factors involved in their evolutionary radiation and the origin of the hominin clade, a key subject for the further understanding of human evolution, remained mostly unexplored. Here we quantitatively approach these events using phylogenetic comparative methods and craniofacial morphometric data from extant and fossil hominoid species. Specifically, we explore alternative evolutionary models that allow us to gain new insights into this clade diversification process. Our results show a complex and variable scenario involving different evolutionary regimes through the hominid evolutionary radiation -modeled by Ornstein-Uhlenbeck multi-selective regime and Brownian motion multi-rate scenarios-. These different evolutionary regimes might relate to distinct ecological and cultural factors previously suggested to explain hominid evolution at different evolutionary scales along the last 10 million years. "...CT scans were processed by means of 3D Slicer and MeshLab software." |
Isolating Phyllotactic Patterns Embedded in the Secondary Growth of Sweet Cherry (Prunus Avium L.) Using Magnetic Resonance Imaging
|
Publication: Plant Methods. 2019 Oct 4;15:111. PMID: 31592133 | PDF Authors: Eithun M, Larson J, Lang G, Chitwood DH, Munch E. Institution: Department of Computational Mathematics, Science and Engineering, Michigan State University, East Lansing, MI, USA. Abstract: BACKGROUND: Epicormic branches arise from dormant buds patterned during the growth of previous years. Dormant epicormic buds remain just below the surface of trees, pushed outward from the pith during secondary growth, but maintain vascular connections. Epicormic buds can be activated to elongate into a new shoot, either through natural processes or horticultural intervention, to potentially rejuvenate orchards and restructure tree architecture. Because epicormic structures are embedded within secondary growth, tomographic approaches are a useful method to study them and understand their development. RESULTS: We apply techniques from image processing to determine the locations of epicormic vascular traces embedded within secondary growth of sweet cherry (Prunus avium L.), revealing the juvenile phyllotactic pattern in the trunk of an adult tree. Techniques include the flood fill algorithm to find the pith of the tree, edge detection to approximate the radius, and a conversion to polar coordinates to threshold and segment phyllotactic features. Intensity values from magnetic resonance imaging (MRI) of the trunk are projected onto the surface of a perfect cylinder to find the locations of traces in the "boundary image". Mathematical phyllotaxy provides a means to capture the patterns in the boundary image by modeling phyllotactic parameters. Our cherry tree specimen has the conspicuous parastichy pair (2,3), phyllotactic fraction 2/5, and divergence angle of approximately 143°. CONCLUSIONS: The methods described provide a framework not only for studying phyllotaxy, but also for processing of volumetric image data in plants. Our results have practical implications for orchard rejuvenation and directed approaches to influence tree architecture. The study of epicormic structures, which are hidden within secondary growth, using tomographic methods also opens the possibility of studying genetic and environmental influences such structures. |
 Cherry tree trunk and 3D reconstruction. MRI produces a voxel-based image of the specimen. Using 3D Slicer, an open-source software tool, we created a 3D reconstruction. Two types of traces (a branching trace and a single trace) are shown as physical cross-sections made by cutting the trunk and as slices from the scan. |
Decoding Tumor Mutation Burden And Driver Mutations In Early Stage Lung Adenocarcinoma Using Ct-Based Radiomics Signature
|
Publication: Thorac Cancer. 2019 Oct;10(10):1904-1912. PMID: 31414580 | PDF Authors: Wang X, Kong C, Xu W, Yang S, Shi D, Zhang J, Du M, Wang S, Bai Y, Zhang T, Chen Z, Ma Z, Wang J, Dong G, Sun M, Yin R, Chen F. Institution: Department of Biostatistics, School of Public Health, Nanjing Medical University, Nanjing, China. Abstract: BACKGROUND: Tumor mutation burden (TMB) is an important determinant and biomarker for response of targeted therapy and prognosis in patients with lung cancer. The present study aimed to determine whether radiomics signature could non-invasively predict the TMB status and driver mutations in patients with resectable early stage lung adenocarcinoma (LUAD). METHODS: A total of 61pulmonary nodules (PNs) from 51 patients post-operatively diagnosed LUAD were enrolled for analysis. Two datasets were divided according to two-thirds of patients from different commercial Comprehensive Genomic Profiling (CGP) panels: a training cohort including 41 PNs and a testing cohort including rest 20PNs. We sequenced all tumor specimens and paired blood cells using next generation sequencing (NGS), so as to detect TMB status and somatic mutations. We collected 718 quantitative 3D radiomics features extracted from segmented volumes of each PNs and 78 clinical and pathological features retrieved from medical records as well. Support vector machine methods were performed to establish the predictive model. RESULTS: We established an efficient fusion-positive tumor prediction model that predicts TMB status and EGFR/TP53 mutations of early stage LUAD. The radiomics signature yielded a median AUC value of 0.606, 0.604, and 0.586 respectively. Combining radiomics with the clinical information can further improve the prediction performance, which the median AUC values are 0.671 for TMB, 0.697 and 0.656 for EGFR/TP53 respectively. CONCLUSION: It is feasible and effective to facilitate TMB and somatic driver mutations prediction by using the radiomics signature and NGS data in early stage LUAD. "...CT images were imported into the 3D Slicer v4.7.0 software and the tumors were then contoured manually by three independent observers using the built‐in paint tool. " |
Mammographic Breast Density Assessed with Fully Automated Method and its Risk for Breast Cancer
|
Publication: J Clin Imaging Sci. 2019 Oct 11;9:43. PMID: 31662951 | PDF Authors: Saikiran P, Ramzan R, Nadish S, Kamineni PD, Priyanka, John AM. Institution: Department of Medical Imaging Technology, Manipal College of Health Professions, Manipal, Karnataka, India. Abstract: OBJECTIVES: We evaluated the association between breast cancer and breast density (BD) measured using fully automated software. We also evaluated the performance of cancer risk models such as only clinical risk factors, density related measures, and both clinical risk factors and density-related measures for determining cancer risk. MATERIALS AND METHODS: This is a retrospective case-control study. The data were collected from August 2015 to December 2018. Two hundred fifty women with breast cancer and 400 control subjects were included in this study. We evaluated the BD qualitatively using breast imaging-reporting and data system density and quantitatively using 3D Slicer. We also collected clinical factors such as age, familial history of breast cancer, menopausal status, number of births, body mass index, and hormonal replacement therapy use. We calculated the odds ratio (OR) for BD to determine the risk of breast cancer. We performed receiver operating characteristic (ROC) curve to assess the performance of cancer risk models. RESULTS: The OR for the percentage BD for second, third, and fourth quartiles was 1.632 (95% confidence intervals [CI]: 1.102-2.416), 2.756 (95% CI: 1.704-4.458), and 3.163 (95% CI: 1.356-5.61). The area under ROC curve for clinical risk factors only, mammographic density measures, combined mammographic, and clinical risk factors was 0.578 (95% CI: 0.45, 0.64), 0.684 (95% CI: 0.58, 0.75), and 0.724 (95% CI: 0.64, 0.80), respectively. CONCLUSION: Mammographic BD was found to be positively associated with breast cancer. The density related measures combined clinical risk factors, and density model had good discriminatory power in identifying the cancer risk. |
Advantage of Proton-Radiotherapy for Pediatric Patients and Adolescents With Hodgkin's Disease
|
Publication: Radiat Oncol. 2019 Sep 2;14(1):157. PMID: 31477141 | PDF Authors: Lautenschlaeger S, Iancu G, Flatten V, Baumann K, Thiemer M, Dumke C, Zink K, Hauswald H, Vordermark D, Mauz-Körholz C, Engenhart-Cabillic R, Eberle F. Institution: Klinik für Strahlentherapie und Radioonkologie, Klinikum der Philipps Universität Marburg, Marburg, Germany. Abstract: Radiotherapy is frequently used in the therapy of lymphoma. Since lymphoma, for example Hodgkin's disease, frequently affect rather young patients, the induction of secondary cancer or other long-term adverse effects after irradiation are important issues to deal with. Especially for mediastinal manifestations numerous organs and substructures at risk play a role. The heart, its coronary vessels and cardiac valves, the lungs, the thyroid and, for female patients, the breast tissue are only the most important organs at risk. In this study we investigated if proton-radiotherapy might reduce the dose delivered to the organs at risk and thus minimize the therapy-associated toxicity. METHODS: In this work we compared the dose delivered to the heart, its coronary vessels and valves, the lungs, the thyroid gland and the breast tissue by different volumetric photon plans and a proton plan, all calculated for a dose of 28.8 Gy (EURO-NET-PHL-C2). Target Volumes have been defined by F18-FDG PET-positive areas, following a modified involved node approach. Data from ten young female patients with mediastinal lymphoma have been evaluated. Three different modern volumetric IMRT (VMAT) photon plans have been benchmarked against each other and against proton-irradiation concepts. For plan-evaluation conformity- and homogeneity-indices have been calculated as suggested in ICRU 83. The target volume coverage as well as the dose to important organs at risk as the heart with its substructures, the lungs, the breast tissue, the thyroid and the spinal cord were calculated and compared. For statistical evaluation mean doses to organs at risk were evaluated by non- parametric Kruskal-Wallis calculations with pairwise comparisons. RESULTS: Proton-plans and three different volumetric photon-plans have been calculated. Proton irradiation results in significant lower doses delivered to organ at risk. The median doses and the mean doses could be decreased while PTV coverage is comparable. As well conformity as homogeneity are slightly better for proton plans. For several organs a risk reduction for secondary malignancies has been calculated using literature data as reference. According to the used data derived from literature especially the secondary breast cancer risk, the secondary lung cancer risk and the risk for ischemic cardiac insults can be reduced significantly by using protons for radiotherapy of mediastinal lymphomas. CONCLUSION: Irradiation with protons for mediastinal Hodgkin-lymphoma results in significant lower doses for almost all organs at risk and is suitable to reduce long term side effects for pediatric and adolescent patients. "...Mean DVHs involving all individual DVHs for each organ at risk were computed using Eclipse 13.7 (Varian) and 3D Slicer v4.8.1. " |
Potential Role of Convolutional Neural Network Based Algorithm in Patient Selection for DCIS Observation Trials Using a Mammogram Dataset
|
Publication: Acad Radiol. 2019 Sep 13. PMID: 31526687 Authors: Mutasa S, Chang P, Van Sant EP, Nemer J, Liu M, Karcich J, Patel G, Jambawalikar S, Ha R. Institution: Department of Medical Physics and Radiology, Columbia University Medical Center, NY, USA. Abstract: RATIONALE AND OBJECTIVES: We investigated the feasibility of utilizing convolutional neural network (CNN) for predicting patients with pure Ductal Carcinoma In Situ (DCIS) versus DCIS with invasion using mammographic images. MATERIALS AND METHODS: An IRB-approved retrospective study was performed. 246 unique images from 123 patients were used for our CNN algorithm. In total, 164 images in 82 patients diagnosed with DCIS by stereotactic-guided biopsy of calcifications without any upgrade at the time of surgical excision (pure DCIS group). A total of 82 images in 41 patients with mammographic calcifications yielding occult invasive carcinoma as the final upgraded diagnosis on surgery (occult invasive group). Two standard mammographic magnification views (CC and ML/LM) of the calcifications were used for analysis. Calcifications were segmented using an open source software platform 3D Slicer and resized to fit a 128 × 128 pixel bounding box. A 15 hidden layer topology was used to implement the neural network. The network architecture contained five residual layers and dropout of 0.25 after each convolution. Five-fold cross validation was performed using training set (80%) and validation set (20%). Code was implemented in open source software Keras with TensorFlow on a Linux workstation with NVIDIA GTX 1070 Pascal GPU. RESULTS: Our CNN algorithm for predicting patients with pure DCIS achieved an overall diagnostic accuracy of 74.6% (95% CI, ±5) with area under the ROC curve of 0.71 (95% CI, ±0.04), specificity of 91.6% (95% CI, ±5%) and sensitivity of 49.4% (95% CI, ±6%). CONCLUSION: It's feasible to apply CNN to distinguish pure DCIS from DCIS with invasion with high specificity using mammographic images. |
Texture Analysis of Pretreatment 18FFDG PET/CT for the Prognostic Prediction of Locally Advanced Salivary Gland Carcinoma Treated with Interstitial Brachytherapy
|
Publication: EJNMMI Res. 2019 Sep 11;9(1):89. PMID: 31511990 | PDF Authors: Wu WJ, Li ZY, Dong S, Liu SM, Zheng L, Huang MW, Zhang JG. Institution: Department of Oral and Maxillofacial Surgery, Peking University School and Hospital of Somatology, Beijing, China. Abstract: BACKGROUND: The aim of this study was to evaluate the prognostic value of positron emission tomography (PET) parameters and the PET texture features of fluorine 18-fluorodeoxyglucose ([18F]FDG) uptake on pretreatment PET/computed tomography (CT) in patients with locally advanced salivary gland carcinoma treated with interstitial brachytherapy. METHODS: Forty-three patients with locally advanced salivary gland carcinoma of the head and neck were treated with 125I interstitial brachytherapy as the sole modality and underwent 18FFDG PET/CT scanning before treatment. Tumor segmentation and texture analysis were performed using the 3D Slicer software. In total, 54 features were extracted and categorized as first-order statistics, morphology and shape, gray-level co-occurrence matrix, and gray-level run length matrix. Up to November 2018, the follow-up time ranged from 6 to 120 months (median 18 months). Cumulative survival was calculated by the Kaplan-Meier method. Factors between groups were compared by the log-rank test. Multivariate Cox regression analysis with a backward conditional method was used to predict progression-free survival (PFS). RESULTS: The 3- and 5-year locoregional control (LC) rates were 55.4% and 37.0%, respectively. The 3- and 5-year PFS rates were 51.2% and 34.1%, respectively. The 3- and 5-year overall survival (OS) rates were 77.0% and 77.0%, respectively. Univariate analysis revealed that minimum intensity, mean intensity, median intensity, root mean square, and long run emphasis (LRE) were significant predictors of PFS, whereas clinicopathological factors, conventional PET parameters, and PET texture features failed to show significance. Multivariate Cox regression analysis showed that minimum intensity and LRE were significant predictors of PFS. CONCLUSIONS: The texture analysis of pretreatment [18F]FDG PET/CT provided more information than conventional PET parameters for predicting patient prognosis of locally advanced salivary gland carcinoma treated with interstitial brachytherapy. The minimum intensity was a risk factor for PFS, and LRE was a favorable factor in prognostic prediction according to the primary results. |
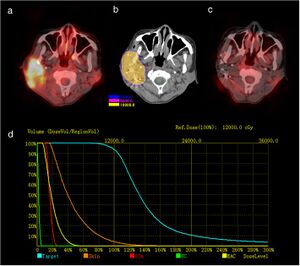 The procedure of brachytherapy based on PET/CT figure: Pretreatment PET/CT showed a T4 parotid gland carcinoma (a). Quality verification using postoperative CT images in the brachytherapy treatment planning system showed that the D90 curve covered the tumor area (b). PET/CT showed that the area of focal FDG uptake regressed 6 months after brachytherapy (c). Dose-volume histograms of the target area and organs at risk are shown (d). ICA internal carotid artery, SC spinal cord, EAC external auditory canal. |
Levator Bowl Volume during Straining and its Relationship to other Levator Measures
|
Publication: Int Urogynecol J. 2019 Sep;30(9):1457-63. PMID: 31222569 Authors: Nandikanti L, Sammarco AG, Chen L, Ashton-Miller JA, DeLancey JO. Institution: School of Public Health, University of Michigan, Ann Arbor, MI, USA. Abstract: INTRODUCTION AND HYPOTHESIS: This study was aimed at measuring levator ani bowl volume at rest and while straining, comparing women with and without prolapse (controls), and assessing the ability of measures of the mid-sagittal bowl area, levator hiatus (LH), and urogenital hiatus (UGH) to predict bowl volume. METHODS: Forty MRI scans previously acquired in case-control prolapse studies, including 20 women with prolapse and 20 women without prolapse, of similar age and parity, were selected. 3D models of rest and strain bowl volumes were made using sagittal scans and 3D Slicer. Mid-sagittal bowl area, UGH, and LH were measured using ImageJ. Data were analyzed using two sample t tests, effect sizes, and Pearson's correlation coefficients at the 0.05 significance level. RESULTS: Data were acquired in a total of 40 total women. Levator bowl volume at strain had a correlation coefficient of 0.5 with bowl volume at rest. During straining, prolapse subjects had a 53% larger bowl volume than control subjects (254 ± 86 cm3 vs 166 ± 44 cm3, p < 0.001), but at rest, the difference was 34% (138 ± 40 cm3 vs 103 ± 25 cm3, p = 0.002). Effect sizes for all parameters were large (d > 0.75). The strongest correlation with straining bowl volume was mid-sagittal straining bowl area (r = 0.86), followed by LH strain (r = 0.80), then UGH strain (r = 0.76). CONCLUSIONS: Straining levator bowl volume is substantially different than measures made at rest, with only a quarter of straining values explained by resting measurements. The bowl area at strain is the best 2D measurement estimating bowl volume and explains 74% of straining bowl volume.
|
Mixed Reality-Based Preoperative Planning for Training of Percutaneous Transforaminal Endoscopic Discectomy: A Feasibility Study.
|
Publication: World Neurosurg. 2019 Sep;129:e767-e775. PMID: 31203062 Authors: Yu H, Zhou Z, Lei X, Liu H, Fan G, He S. Institution: Orthopedic Department, Shanghai Tenth People's Hospital, Shanghai, China. Abstract: OBJECTIVE: To explore the effect of preoperative planning using mixed reality (MR) on training of percutaneous transforaminal endoscopic discectomy (PTED). METHODS: Before the training, we invited an experienced chief physician to plan the puncture path of PTED on the X-ray films of the lumbar spine model and the 3D Slicer platform, respectively, and used this as the standard to guide trainees. In the aggregate, 60 young residents were randomly divided into Group A (N = 30) and Group B (N = 30). Group A learned the 2-dimensional standard planning route, whereas Group B learned the standard route planning based on MR through the 3D Slicer platform. Then, trainees were asked to conduct PTED puncture on a lumbar spine model. Questionnaires were distributed to trainees before and after the training. During the training, puncture times, operating time (minutes), and fluoroscopy times were recorded. RESULTS: After the training, it was obvious that more trainees showed their recognition of MR, believing that MR could help preoperative planning and training of PTED. Their high satisfaction with the training indicated the success of our training. Moreover, puncture times, operating time (minutes), and fluoroscopy times of Group B were significantly lower than those of Group A. CONCLUSIONS: MR technology contributes to preoperative planning of PTED and is beneficial in the training of PTED. It significantly reduces puncture times and fluoroscopy times, providing a standardized method for the training of PTED. |
Evaluation of Accuracy of a Three-Dimensional Printed Model in Open-Wedge High Tibial Osteotomy
|
Publication: J Knee Surg. 2019 Sep;32(9):841-6. PMID: 30189435 | PDF Authors: Kim HJ, Park J, Park KH, Park IH, Jang JA, Shin JY, Kyung HS. Institution: Department of Orthopaedic Surgery, School of Medicine, Kyungpook National University, Kyungpook National University Hospital, Daegu, South Korea. Abstract: The purpose of this study was to evaluate the usefulness of a three-dimensional (3D) printed model for open-wedge high tibial osteotomy (HTO). This study retrospectively evaluated 20 patients with medial knee osteoarthritis and varus deformity. Between October 2015 and July 2016, the patients underwent open-wedge HTO using a 3D printed model. The mean age of patients was 55.2 years (range, 51-60 years). The mean preoperative mechanical femorotibial angle (mFTA) was varus 7.8 degrees (range, varus 4.7-11.6 degrees). After measuring the target angle using full-length lower limb weight-bearing radiography, the osteotomy was simulated using 3D images obtained from computed tomography (CT) with the 3D Slicer program. On the basis of the simulated osteotomy section and the target angle, the model was then designed and printed. Open-wedge HTO was then performed by applying the 3D printed model to the opening gap. The accuracy of osteotomy and the change in posterior tibial slope (PTS) angle were evaluated. The weight-bearing line on the tibial plateau was corrected from a preoperative mean of 19.5 ± 9.8% to a postoperative mean of 63.1 ± 6.1% (p < 0.001). The postoperative values were not statistically significantly different from the preoperative target points (p = 0.688). The mFTA was corrected to a postoperative mean of valgus 3.8 ± 1.4 degrees. The PTS angle showed no significant change (p = 0.256). A 3D printed model using CT may be useful for preoperative planning of open-wedge HTO. Satisfactory correction can be obtained without a change in the PTS. |
Neuroradiological Changes Following Single or Repetitive Mild TBI
|
Publication: Front Syst Neurosci. 2019 Aug 2;13:34. PMID: 31427931 | PDF Authors: Kulkarni P, Morrison TR, Cai X, Iriah S, Simon N, Sabrick J, Neuroth L, Ferris CF. Institution: Center for Translational NeuroImaging, Northeastern University, Boston, MA, USA. Abstract: OBJECTIVES: To test the hypothesis that there are differences in neuroradiological measures between single and repeated mild traumatic brain injury using multimodal MRI. METHODS: A closed-head momentum exchange model was used to produce one or three mild head injuries in young adult male rats compared to non-injured, age and weight-matched controls. Six-seven weeks post-injury, rats were studied for deficits in cognitive and motor function. Seven-eight weeks post-injury changes in brain anatomy and function were evaluated through analysis of high resolution T2 weighted images, resting-state BOLD functional connectivity, and diffusion weighted imaging with quantitative anisotropy. RESULTS: Head injuries occurred without skull fracture or signs of intracranial bleeding or contusion. There were no significant differences in cognitive or motors behaviors between experimental groups. With a single mild hit, the affected areas were limited to the caudate/putamen and central amygdala. Rats hit three times showed altered diffusivity in white matter tracts, basal ganglia, central amygdala, brainstem, and cerebellum. Comparing three hits to one hit showed a similar pattern of change underscoring a dose effect of repeated head injury on the brainstem and cerebellum. Disruption of functional connectivity was pronounced with three mild hits. The midbrain dopamine system, hippocampus, and brainstem/cerebellum showed hypoconnectivity. Interestingly, rats exposed to one hit showed enhanced functional connectivity (or hyperconnectivity) across brain sites, particularly between the olfactory system and the cerebellum. INTERPRETATION: Neuroradiological evidence of altered brain structure and function, particularly in striatal and midbrain dopaminergic areas, persists long after mild repetitive head injury. These changes may serve as biomarkers of neurodegeneration and risk for dementia later in life. "...Brain tissue masks for resting-state functional images were manually drawn using 3D Slicer. " Funding:
|
Technique Development and Measurement of Cross-Sectional Area of the Pubovisceral Muscle on MRI Scans of Living Women
|
Publication: Int Urogynecol J. 2019 Aug;30(8):1305-12. PMID: 29974138 Authors: Masteling M, Ashton-Miller JA, DeLancey JOL. Institution: Department of Mechanical Engineering, University of Michigan, Ann Arbor, MI, USA. Abstract: INTRODUCTION AND HYPOTHESIS: Measurements of the anatomic cross-sectional area (CSA) of the pubovisceral muscle (PVM) in women are confounded by the difficulty of separating the muscle from the adjacent puborectal (PRM) and iliococcygeal (ICM) muscles when visualized in a plane orthogonal to the fiber direction. We tested the hypothesis that it might be possible to measure the PVM CSA within a defined region of interest based on magnetic resonance images (MRI). METHODS: MRI scans of 11 women with unilateral PVM tears and seven primiparous women with intact muscles following elective C-section were used to identify the PVM injury zone defined by the mean location of its boundaries with the adjacent intact PRM and ICM from existing anatomic reference points using 3D Slicer and ImageJ software. Then, from the 15 or more 2-mm transverse slices available, the slice with the maximum anatomic CSA of the left and right PVM was found in 24 primiparous women with bilaterally intact muscles who had delivered via C-section. RESULTS: Mean [± standard deviation (SD)] of the maximum left or right PVM cross-section areas for the 24 women, measured by two different raters, was 1.25 ± 0.29 cm2 (range 0.75-1.86). The 5th, 50th, and 95th percentile values were 0.77, 1.23, and 1.80 cm2, respectively. Inter- and intrarater measurement repeatability intraclass correlation coefficients exceeded 0.89 and 0.90, respectively. CONCLUSIONS: It is possible to use MRI to identify the volume of interest with the maximum anatomic cross section of the PVM belly while minimizing the inadvertent inclusion of adjacent PRM or ICM in that measurement. |
A Comparative Evaluation of SteamVR Tracking and the OptiTrack System for Medical Device Tracking
|
Publication: Conf Proc IEEE Eng Med Biol Soc. 2019 Jul;2019:1465-70. PMID: 31946170 Authors: Ameler T, Warzecha M, Hes D, Fromke J, Schmitz-Stolbrink A, Friedrich CM, Blohme K, Brandt L, Brungel R, Hensel A, Huber L, Kuper F, Swoboda J, Warnecke M. Abstract: Tracking of medical devices can be used in diverse situations, e.g., training as well as image guidance for surgery and surgery planning. Therefore, position and orientation of a device, for instance, an ultrasound probe, need to be identified as precisely as possible. This enables correct representation of digital 3D models in medical image processing platforms such as 3D Slicer or MevisLab. In this manuscript, a comparative evaluation of the low-cost Swept Angle Laser Tracking (SALT) system SteamVR Tracking and the multi-camera-based Opti-Track System is presented. Their potential for medical device tracking is demonstrated in the use case of ultrasound probe tracking for simulation purposes. An evaluation of tracking errors is performed using a Universal Robotics UR5 industrial robot under non-laboratory conditions, involving common issues such as reflections and occlusions. A discussion on the tracking accuracy of both systems is given. The communication of tracking data is established for 3D Slicer and MeVisLab with the use of the PLUS Toolkit via the OpenIGTLink protocol. |
The Middle Fossa Approach with Self-drilling Screws: a Novel Technique for BONEBRIDGE Implantation
|
Publication: J Otolaryngol Head Neck Surg. 2019 Jul 29;48(1):35. PMID: 31358057 | PDF Authors: You P, Siegel LH, Kassam Z, Hebb M, Parnes L, Ladak HM, Agrawal SK. Institution: Department of Otolaryngology-Head and Neck Surgery, Schulich School of Medicine & Dentistry, Western University, London, Canada. Abstract: BACKGROUND: Bone conduction implants can be used in the treatment of conductive or mixed hearing loss. The BONEBRIDGE bone conduction implant (BB-BCI) is an active, transcutaneous device. BB-BCI implantation can be performed through either the transmastoid or retrosigmoid approach with their respective limitations. Here, we present a third, novel approach for BB-BCI implantation. OBJECTIVE: Describe the detailed surgical technique of BB-BCI implantation through a middle fossa approach with self-drilling screws and present preliminary audiometric outcome data following this approach. METHODS: A single institution, retrospective chart review was completed for patients implanted with the BB-BCI via the middle fossa approach. Preoperative planning and modelling were performed using 3D Slicer. Audiological testing was performed pre- and post-operatively following standard audiometric techniques. RESULTS: Forty patients underwent BB-BCI implantation using the middle fossa approach. Modelling techniques allowed for implantation through the use of external landmarks, obviating the need for intraoperative image guidance. The surgical technique was refined over time through experience and adaptation. Mean follow-up was 29 months (range 3-71 months) with no surgical complications, favourable cosmesis, and expected audiometric outcomes. An average functional gain of 39.6 dB (± 14.7 SD) was found. CONCLUSION: The middle fossa technique with self-drilling screws is a safe and effective option for BONEBRIDGE implantation. As a reference for other groups considering this approach, an annotated video has been included as a supplement to the study. |
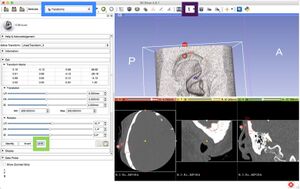 Screen capture of the 3D Slicer interface in Conventional view for preoperative planning of middle fossa approach to BONEBRIDGE implantation. Traditional axial, sagittal, and coronal views of a temporal bone CT are visible at the bottom of the screen. The BC-FMT is highlighted with a red outline on these slices, and a red fiducial marks the centre of the implant on the skin. Blue box = module drop down menu. Green box = option to select Transform selection to alternate between global versus regional transform. Purple box = option to add fiducials. F = digital fiducial |
Evaluation of Flow Changes After Telescopic Stenting of a Giant Fusiform Aneurysm of the Vertebrobasilar Junction
|
Publication: Biomed Eng Online. 2019 Jul 24;18(1):82. PMID: 31340820 | PDF Authors: Sindeev S, Kirschke JS, Prothmann S, Frolov S, Liepsch D, Berg P, Zimmer C, Friedrich B. Institution: Department of Biomedical Engineering, Tambov State Technical University, Tambov, Russia. Abstract: BACKGROUND: The use of flow-diverters for non-saccular cerebral posterior circulation aneurysms requires complex deployment techniques and is associated with high mortality and morbidity. Therefore, further studies are required to clarify the effect of stenting on post-treatment hemodynamics in such aneurysms. In this study, we evaluated flow alterations in a treated giant fusiform aneurysm of the vertebrobasilar junction and correlated them with the clinical outcome. METHODS: A patient-specific aneurysm model was acquired by rotational angiography, and three SILK flow-diverters (4.5 × 40, 5 × 40 and 5.5 × 40 mm) were virtually deployed in series along the basilar and right vertebral arteries. Image-based blood flow simulations before and after the treatment were performed under realistic pulsatile flow conditions. The flow reduction, velocity and wall shear stress (WSS) distribution, streamlines and WSS-derived parameters were evaluated before and after the treatment. RESULTS: The computed velocity streamlines showed substantial alterations of the flow pattern in the aneurysm and successful redirection of blood flow along the series of flow-diverters with no flow through the overlapping stents. The obtained flow reduction of 86% was sufficient to create thrombogenic flow conditions. Moreover, a 6.2-fold increase in relative residence time and a decrease by 87% of time-averaged WSS contributed to a successful treatment outcome observed during the follow-up. CONCLUSIONS: We found a correlation between the numerically predicted flow alterations and the available treatment outcome. This shows the potential of image-based simulations to be used in clinical practice for treatment planning and estimation of possible risk factors associated with a complex stent deployment in fusiform aneurysms of the posterior circulation. "..The data were segmented with open-source software 3D Slicer v4.8.1." |
a New Acoustic Coupling Fluid With Ability to Reduce Ultrasound Imaging Artefacts in Brain Tumour Surgery-A Phase I Study
|
Publication: Acta Neurochir (Wien). 2019 Jul;161(7):1475-86. PMID: 31104122 | PDF Authors: Unsgård G, Sagberg LM, Müller S, Selbekk T. Institution: Department of Neuromedicine and Movement Science, Norwegian University of Science and Technology, Trondheim, Norway. Abstract: BACKGROUND: A novel acoustic coupling fluid (ACF), with the potential to reduce surgically induced image artefacts during intraoperative ultrasound imaging in brain tumour surgery, has been evaluated with respect to image quality and safety in a clinical phase 1 study. METHODS: Fifteen patients with glioblastoma (WHO grade IV) were included. All adverse events were registered in a 6-month study period. During acquisition of 3D ultrasound image volumes, three different concentrations of the ACF and Ringer's solution were filled into the resection cavity. The effect of ACF on the ultrasound images was rated by the operating surgeon, and by five independent neurosurgeons evaluating a pair of blinded images from all patients. Images from all patients were analysed by comparing pixel brightness in a noise-affected region and a reference region. RESULTS: The operating surgeon deemed the ACF images to have less noise than images obtained with Ringers's solution. The blinded evaluations by the independent neurosurgeons were significantly in favour of ACF (p < 0.0001). The analyses of pixel intensities showed that the ACF images had lower amount of noise than images obtained with Ringer's solution. No radiological sign of inflammation nor circulatory changes was found in the early postoperative MR images. Of the nine complications registered as serious events in the study period, none was deemed to be caused by the ACF. CONCLUSION: The ultrasound (US) images obtained using ACF have significantly less noise than US images obtained with Ringer's solution. The rate of adverse events was comparable to what has been reported for similar groups of patients. "...quantitative image analyses were performed postoperatively by a single operator blinded for the results of the qualitative image analyses using the software 3D Slicer v4.4.0." |
CT-Based Radiomic Signatures for Prediction of Pathologic Complete Response in Esophageal Squamous Cell Carcinoma After Neoadjuvant Chemoradiotherapy
|
Publication: J Radiat Res. 2019 Jul 1;60(4):538-45. PMID: 31111948 | PDF Authors: Yang Z, He B, Zhuang X, Gao X, Wang D, Li M, Lin Z, Luo R. Institution: Department of Radiation Oncology, Cancer Hospital of Shantou University Medical College, Guangdong, China. Abstract: The objective of this study was to build models to predict complete pathologic response (pCR) after neoadjuvant chemoradiotherapy (nCRT) in esophageal squamous cell carcinoma (ESCC) patients using radiomic features. A total of 55 consecutive patients pathologically diagnosed as having ESCC were included in this study. Patients were divided into a training cohort (44 patients) and a testing cohort (11 patients). The logistic regression analysis using likelihood ratio forward selection was performed to select the predictive clinical parameters for pCR, and the least absolute shrinkage and selection operator (LASSO) with logistic regression to select radiomic predictors in the training cohort. Model performance in the training and testing groups was evaluated using the area under the receiver operating characteristic curves (AUC). The multivariate logistic regression analysis identified no clinical predictors for pCR. Thus, only radiomic features selected by LASSO were used to build prediction models. Three logistic regression models for pCR prediction were developed in the training cohort, and they were able to predict pCR well in both the training (AUC, 0.84-0.86) and the testing cohorts (AUC, 0.71-0.79). There were no differences between these AUCs. We developed three predictive models for pCR after nCRT using radiomic parameters and they demonstrated good model performance. "...3D Slicer (v4.8.1, Stable Release) with its extension (radiomics) was used for collecting the radiomic features from pre-treatment CT" |
Diffusion Abnormalities in the Corpus Callosum in First Episode Schizophrenia: Associated With Enlarged Lateral Ventricles and Symptomatology
|
Publication: Psychiatry Res. 2019 Jul;277:45-51. PMID: 30808608 | PDF Authors: Del Re EC, Bouix S, Fitzsimmons J, Blokland GAM, Mesholam-Gately R, Wojcik J, Kikinis Z, Kubicki M, Petryshen T, Pasternak O, Shenton ME, Niznikiewicz M. Institution: Laboratory of Neuroscience, Department of Psychiatry, VA Boston Healthcare System, Brockton Division, and Harvard Medical School, Boston, MA, USA. Abstract: INTRODUCTION: Abnormalities in the corpus callosum (CC) and the lateral ventricles (LV) are hallmark features of schizophrenia. These abnormalities have been reported in chronic and in first episode schizophrenia (FESZ). Here we explore further associations between CC and LV in FESZ using diffusion tensor imaging (DTI). METHODS: Sixteen FESZ patients and 16 healthy controls (HC), matched on age, gender, and handedness participated in the study. Diffusion and structural imaging scans were acquired on a 3T GE Signa magnet. Volumetric measures for LV and DTI measures for five CC subdivisions were completed in both groups. In addition, two-tensor tractography, the latter corrected for free-water (FAt), was completed for CC. Correlations between LV and DTI measures of the CC were examined in both groups, while correlations between DTI and clinical measures were examined in only FESZ. RESULTS: Results from two-tensor tractography demonstrated decreased FAt and increased trace and radial diffusivity (RDt) in the five CC subdivisions in FESZ compared to HC. Central CC diffusion measures in FESZ were significantly correlated with volume of the LV, i.e., decreased FAt values were associated with larger LV volume, while increased RDt and trace values were associated with larger LV volume. In controls, correlations were also significant, but they were in the opposite direction from FESZ. In addition, decreased FAt in FESZ was associated with more positive symptoms. DISCUSSION: Partial volume corrected FAt, RDt, and trace abnormalities in the CC in FESZ suggest possible de- or dys-myelination, or changes in axonal diameters, all compatible with neurodevelopmental theories of schizophrenia. Correlational findings between the volume of LV and diffusion measures in FESZ reinforce the concept of a link between abnormalities in the LV and CC in early stages of schizophrenia and are also compatible with neurodevelopmental abnormalities in this population. "...In reconstructing regions of interest (ROIs), tensors were estimated by the least squares method using 3D Slicer software v4.3.1." |
Computed Tomographic Portography with Esophageal Variceal Measurements in the Evaluation of Esophageal Variceal Severity and Assessment of Esophageal Variceal Volume Efficacy
|
Publication: Acad Radiol. 2019 Jul 11. PMID: 31303576 Authors: Wan S, Wei Y, Yu H, Li Y, Yao S, Song B. Institution: Department of Radiology, West China Hospital of Sichuan University, Chengdu, Sichuan, China. Abstract: RATIONALE AND OBJECTIVES: The aim of our study is to evaluate the severity of esophageal varices (EV), based on the computed tomographic portography (CTP) measurement of EV in the distal esophagus and to assess the prediction value of EV volume. PATIENTS AND METHODS: A total of 53 EV patients examined by CTP within 4 weeks of upper endoscopy were evaluated, the patients were divided into a nonconspicuous EV group (mild-to-moderate EV, n = 28) and a conspicuous EV group (severe EV, n = 25) according to endoscopy results. The diameter, cross-sectional surface area (CSA), and volume of EV were measured independently using 3D Slicer by two experienced abdominal radiologists blinded to endoscopy findings. The averaged values measured by the two observers were used in the final dataset, these indicators' predictive performances were studied by using receiver operating characteristic curve analysis, and the area under the curve (Az) and the cutoff values were calculated to distinguish mild-to-moderate from severe EV. RESULTS: The Az values of volume, diameter and CSA in differentiating severe EV were 0.817, 0.794, and 0.784 for observer-1, corresponding values for observer-2 were 0.796, 0.774, and 0.707, there was almost perfect interobserver agreement for all measurements. All indices were larger in the conspicuous group than the nonconspicuous group in both observers (p ≤ 0.01). In the final dataset, application of a 654.0-mm3-volume criterion yielded sensitivity, specificity of 96%, 50%, application of a 5.2-mm-diameter criterion yielded sensitivity, specificity of 80%, 75%, and application of a 68.6-mm2-CSA criterion yielded sensitivity, specificity of 52%, 93%. CONCLUSION: The volume of EV could be used as a new effective indictor for evaluating EV, and use of volume, diameter, and CSA of EV based on CTP allows discrimination between mild-to-moderate and severe EV in cirrhotic patients. |
Intracranial Mirror Aneurysm: Epidemiology, Rupture Risk, New Imaging, Controversies, and Treatment Strategies
|
Publication: World Neurosurg. 2019 Jul;127:165-175 PMID: 30954748 Authors: Liu HJ, Zhou H, Lu DL, Jiao YB, Chen SF, Cheng J, Yao XJ, Ren JY, Li SF, Liu W, Gao JC, Yue Y, Xu JX, Zhang PN, Feng YG. Institution: Qingdao University, Qingdao, China. Abstract: There are some controversies about the surgical treatment strategy of mirror aneurysms. Whether to choose 1-stage or 2-stage surgery, bilateral or unilateral craniotomy, or surgical or interventional treatment are the main points in dispute. In this review, the different surgery strategies faced by patients are discussed. Different surgical methods are adopted based on the patient's individual state and the location and size of the aneurysm. A new imaging method is introduced using 3D Slicer, which clearly recognizes the relationship among aneurysm, brain tissue, skull, and nerve. The 3D Slicer can help surgeons undertake adequate preoperative preparation. In addition, we also introduce some ruptured factors (e.g., age, gender, hypertension, morphologic, and hemodynamic) concerning mirror aneurysm. Systematic discussion of the controversies and methods in surgical treatment of mirror aneurysms may provide new perspectives in future research for the prevention and treatment of mirror aneurysms. |
Development and Evaluation of a "Trackerless" Surgical Planning and Guidance System Based on 3D Slicer
|
Publication: J Med Imaging (Bellingham). 2019 Jul;6(3):035002. PMID: 31528660 Authors: Yang X, Narasimhan S, Luo M, Thompson RC, Chambless LB, Morone PJ, He L, Dawant BM, Miga MI. Institution: Vanderbilt University, Department of Electrical Engineering and Computer Science, Nashville, TN, USA. Abstract: Conventional optical tracking systems use cameras sensitive to near-infrared (NIR) light and NIR illuminated/active-illuminating markers to localize instrumentation and the patient in the operating room (OR) physical space. This technology is widely used within the neurosurgical theater and is a staple in the standard of care for craniotomy planning. To accomplish, planning is largely conducted at the time of the procedure in the OR with the patient in a fixed head orientation. We propose a framework to achieve this in the OR without conventional tracking technology, i.e., a "trackerless" approach. Briefly, we investigate an extension of the 3D Slicer which combines surgical planning and craniotomy designation. While taking advantage of the well-developed 3D Slicer platform, we implement advanced features to aid the neurosurgeon in planning the location of the anticipated craniotomy relative to the preoperatively imaged tumor in a physical-to-virtual setup, and then subsequently aid the true physical procedure by correlating that physical-to-virtual plan with an intraoperative magnetic resonance imaging-to-physical registered field-of-view display. These steps are done such that the craniotomy can be designated without the use of a conventional optical tracking technology. To test this approach, four experienced neurosurgeons performed experiments on five different surgical cases using our 3D Slicer module as well as the conventional procedure for comparison. The results suggest that our planning system provides a simple, cost-efficient, and reliable solution for surgical planning and delivery without the use of conventional tracking technologies. We hypothesize that the combination of this craniotomy planning approach and our past developments in cortical surface registration and deformation tracking using stereo-pair data from the surgical microscope may provide a fundamental realization of an integrated trackerless surgical guidance platform. |
Multi-objective Parameter Auto-tuning for Tissue Image Segmentation Workflows
|
Publication: J Digit Imaging. 2019 Jun;32(3):521-33. PMID: 30402669 Authors: Taveira LFR, Kurc T, Melo ACMA, Kong J, Bremer E, Saltz JH, Teodoro G. Institution: Department of Computer Science, University of Brasília, Brasília, Brazil. Abstract: We propose a software platform that integrates methods and tools for multi-objective parameter auto-tuning in tissue image segmentation workflows. The goal of our work is to provide an approach for improving the accuracy of nucleus/cell segmentation pipelines by tuning their input parameters. The shape, size, and texture features of nuclei in tissue are important biomarkers for disease prognosis, and accurate computation of these features depends on accurate delineation of boundaries of nuclei. Input parameters in many nucleus segmentation workflows affect segmentation accuracy and have to be tuned for optimal performance. This is a time-consuming and computationally expensive process; automating this step facilitates more robust image segmentation workflows and enables more efficient application of image analysis in large image datasets. Our software platform adjusts the parameters of a nuclear segmentation algorithm to maximize the quality of image segmentation results while minimizing the execution time. It implements several optimization methods to search the parameter space efficiently. In addition, the methodology is developed to execute on high-performance computing systems to reduce the execution time of the parameter tuning phase. These capabilities are packaged in a Docker container for easy deployment and can be used through a friendly interface extension in 3D Slicer. Our results using three real-world image segmentation workflows demonstrate that the proposed solution is able to (1) search a small fraction (about 100 points) of the parameter space, which contains billions to trillions of points, and improve the quality of segmentation output by × 1.20, × 1.29, and × 1.29, on average; (2) decrease the execution time of a segmentation workflow by up to 11.79× while improving output quality; and (3) effectively use parallel systems to accelerate parameter tuning and segmentation phases. Funding:
|
In vivo Localization of Cortical Areas using a 3D Computerized Atlas of the Marmoset Brain
|
Publication: Brain Struct Funct. 2019 Jun;224(5):1957-69. PMID: 30963231 Authors: Risser L, Sadoun A, Mescam M, Strelnikov K, Lebreton S, Boucher S, Girard P, Vayssière N, Rosa MGP, Fonta C. Institution: Institut de Mathématiques de Toulouse, UMR5219, Université de Toulouse, CNRS, UPS IMT, 31062, Toulouse Cedex 9, France. Abstract: We created a volumetric template of the marmoset (Callithrix jacchus) brain, which enables localization of the cortical areas defined in the Paxinos et al. (The marmoset brain in stereotaxic coordinates. Elsevier Academic Press, Cambridge, 2012) marmoset brain atlas, as well as seven broader cortical regions (occipital, temporal, parietal, prefrontal, motor, limbic, insular), different brain compartments (white matter, gray matter, cerebro-spinal fluid including ventricular spaces), and various other structures (brain stem, cerebellum, olfactory bulb, hippocampus). The template was designed from T1-weighted MR images acquired using a 3 T MRI scanner. It was based on a single fully segmented marmoset brain image, which was transported onto the mean of 13 adult marmoset brain images using a diffeomorphic strategy that fully preserves the brain topology. In addition, we offer an automatic segmentation pipeline which fully exploits the proposed template. The segmentation pipeline was quantitatively assessed by comparing the results of manual and automated segmentations. An associated program, written in Python, can be used from a command-line interface, or used interactively as a module of the 3D Slicer software. This program can be applied to the analysis of multimodal images, to map specific cortical areas in lesions or to define the seeds for further tractography analyses. |
Quantification of Global Lung Inflammation Using Volumetric 18F-FDG PET/CT Parameters in Locally Advanced Non-Small-Cell Lung Cancer Patients Treated With Concurrent Chemoradiotherapy: A Comparison of Photon and Proton Radiation Therapy
|
Publication: Nucl Med Commun. 2019 Jun;40(6):618-25. PMID: 31095527 Authors: Rice SR, Saboury B, Houshmand S, Salavati A, Kalbasi A, Goodman CR, Werner TJ, Vujaskovic Z, Simone CB 2nd, Alavi A. Institution: Department of Radiation Oncology, University of Maryland School of Medicine, Baltimore, MD, USA. Abstract: INTRODUCTION: Radiation pneumonitis is a major dose-limiting complication in thoracic radiation therapy (RT) and presents clinically in the first few months after RT. We evaluated the feasibility of quantifying pulmonary parenchymal glycolysis (PG) as a surrogate of global lung inflammation and radiation-induced pulmonary toxicity using a novel semiautomatic lung segmentation technique in non-small-cell lung cancer (NSCLC) patients and compared PG in patients treated with photon or proton RT. PATIENTS AND METHODS: We evaluated 18 consecutive locally advanced NSCLC patients who underwent pretreatment and post-treatment F-FDG PET/CT treated with definitive (median: 66.6 Gy; 1.8 Gy fractions) photon or proton RT between 2010 and 2014. Lung volume segmentation was conducted using 3D Slicer by performing simple thresholding. Pulmonary PG was calculated by summing F-FDG uptake in the whole lung. RESULTS: In nine patients treated with photon RT, significant increases in PG in both ipsilateral (mean difference: 1400±510; P=0.02) and contralateral (mean difference: 1200±450; P=0.03) lungs were noted. In nine patients treated with proton therapy, no increase in pulmonary PG was observed in either the ipsilateral (P=0.30) or contralateral lung (P=0.98). CONCLUSION: We observed a significant increase in global lung inflammation bilaterally as measured by quantification of PG. However, no significant change in global lung inflammation was noted after proton therapy. Future larger studies are needed to determine whether this difference correlates with lower risks of radiation pneumonitis in NSCLC patients treated with proton therapy. |
Comprehensive, Multiscale Framework for Evaluation of Arrhythmias Arising From Cell Therapy in the Whole Post-Myocardial Infarcted Heart
|
Publication: Sci Rep. 2019 Jun 25;9(1):9238. PMID: 31239508 | PDF Authors: Yu JK, Franceschi W, Huang Q, Pashakhanloo F, Boyle PM, Trayanova NA. Institution: Institute for Computational Medicine, Johns Hopkins University, Baltimore, MD, USA. Abstract: Direct remuscularization approaches to cell-based heart repair seek to restore ventricular contractility following myocardial infarction (MI) by introducing new cardiomyocytes (CMs) to replace lost or injured ones. However, despite promising improvements in cardiac function, high incidences of ventricular arrhythmias have been observed in animal models of MI injected with pluripotent stem cell-derived cardiomyocytes (PSC-CMs). The mechanisms of arrhythmogenesis remain unclear. Here, we present a comprehensive framework for computational modeling of direct remuscularization approaches to cell therapy. Our multiscale 3D whole-heart modeling framework integrates realistic representations of cell delivery and transdifferentiation therapy modalities as well as representation of spatial distributions of engrafted cells, enabling simulation of clinical therapy and the prediction of emergent electrophysiological behavior and arrhythmogenensis. We employ this framework to explore how varying parameters of cell delivery and transdifferentiation could result in three mechanisms of arrhythmogenesis: focal ectopy, heart block, and reentry. "...patient MRIs are cropped in 3D Slicer." |
Diagnostic Performance of Clinical Properties and Conventional Magnetic Resonance Imaging for Determining the IDH1 Mutation Status in Glioblastoma: A Retrospective Study
|
Publication: PeerJ. 2019 Jun 21;7:e7154. PMID: 31275753 | PDF Authors: Wang Q, Zhang J, Li F, Xu X, Xu B. Institution: Department of Neurosurgery, Chinese PLA General Hospital, Beijing, Beijing, China. Abstract: Glioblastoma (GBM), the most malignant form of gliomas, is a relatively common primary brain tumor in adults. Preoperative identification of isocitrate dehydrogenase 1 (IDH1) mutations in GBM is of critical prognostic importance. The aim of the present study was to explore the feasibility and diagnostic performance of basic patient information combined with conventional magnetic resonance imaging (MRI) findings for determination of the IDH1 status (mutant vs wild type) in patients with GBM. METHODS: From January 1, 2016 to December 31, 2017, a consecutive series of 50 patients with GBM was retrospectively collected. The patients were divided into two group according to their IDH1 mutation status. Basic information and MRI features were analyzed for the establishment of a diagnostic prediction model using logistic regression. A receiver operating characteristic curve was used to evaluate the diagnostic performance. RESULTS: Patients with IDH1-mutant tumors were younger than those with IDH1-wild type tumors, and exhibited a larger tumor volume. The diagnostic predictive model established by combining age and the tumor size exhibited a sensitivity and specificity of 70% and 93%, respectively. The area under the curve was 0.88, which indicated high diagnostic performance. CONCLUSION: Patient age and tumor volume can be used as indicators of IDH1 mutation status in patients with GBM, with high diagnostic performance for simple evaluations in clinical practice. The combined use of these two indicators can further enhance the diagnostic specificity. "...The BRAINS module of the 3D Slicer software was used for linear registration of rigid bodies between sequences.." |
Processing Pipeline for Atlas-Based Imaging Data Analysis of Structural and Functional Mouse Brain MRI (AIDAmri).
|
Publication: Front Neuroinform. 2019 Jun 4;13:42. PMID: 31231202 | PDF Authors: Pallast N, Diedenhofen M, Blaschke S, Wieters F, Wiedermann D, Hoehn M, Fink GR, Aswendt M. Institution: Department of Neurology, Faculty of Medicine and University Hospital Cologne, University of Cologne, Cologne, Germany. Abstract: Magnetic resonance imaging (MRI) is a key technology in multimodal animal studies of brain connectivity and disease pathology. In vivo MRI provides non-invasive, whole brain macroscopic images containing structural and functional information, thereby complementing invasive in vivo high-resolution microscopy and ex vivo molecular techniques. Brain mapping, the correlation of corresponding regions between multiple brains in a standard brain atlas system, is widely used in human MRI. For small animal MRI, however, there is no scientific consensus on pre-processing strategies and atlas-based neuroinformatics. Thus, it remains difficult to compare and validate results from different pre-clinical studies which were processed using custom-made code or individual adjustments of clinical MRI software and without a standard brain reference atlas. Here, we describe AIDAmri, a novel Atlas-based Imaging Data Analysis pipeline to process structural and functional mouse brain data including anatomical MRI, fiber tracking using diffusion tensor imaging (DTI) and functional connectivity analysis using resting-state functional MRI (rs-fMRI). The AIDAmri pipeline includes automated pre-processing steps, such as raw data conversion, skull-stripping and bias-field correction as well as image registration with the Allen Mouse Brain Reference Atlas (ARA). Following a modular structure developed in Python scripting language, the pipeline integrates established and newly developed algorithms. Each processing step was optimized for efficient data processing requiring minimal user-input and user programming skills. The raw data is analyzed and results transferred to the ARA coordinate system in order to allow an efficient and highly-accurate region-based analysis. AIDAmri is intended to fill the gap of a missing open-access and cross-platform toolbox for the most relevant mouse brain MRI sequences thereby facilitating data processing in large cohorts and multi-center studies. "...ARA template that was semi-automatically registered by two independent observers O1 and O2 using a previously described landmark-based registration approach with the help of the software 3D Slicer." |
Ancient Machine Tools for the Construction of the Antikythera Mechanism Parts
|
Publication: Digital Applications in Archaeology and Cultural Heritage. 2019 Jun; 13:e00092. Authors: Aristeidis Voulgaris, Christophoros Mouratidis, Andreas Vossinakis. Institution: Thessaloniki Astronomy Club, Thessaloniki, Greece. Abstract: The present work deals with the study, design, original reconstruction and use of the bow drill of the late archaic period (ca 490 BC), as depicted in two different red figure vases and the vertical lathe depicted on an engraved wall painting of the Petosiris tomb of the Ptolemaic era (300 BC). After the reconstruction of the three ancient tools, during the implementation of the FRAMe Project, their use was thoroughly studied, from which useful conclusions were drawn about the material processing in antiquity, as well as the details of the construction of the Antikythera Mechanism components. Following the new findings detected from the authors’ study of the X-Ray Computed Tomographies from Antikythera Mechanism Research Project, these ancient machine tools can be considered as the progenitors of the Hellenistic period machine tools, which were used for the construction of the mechanical components of the Mechanism. |
Comprehensive Review of 3D Segmentation Software Tools for MRI Usable for Pelvic Surgery Planning
|
Publication: J Digit Imaging. 2019 Jun 24. PMID: 31236743 Authors: Virzì A, Muller CO, Marret JB, Mille E, Berteloot L, Grévent D, Boddaert N, Gori P, Sarnacki S, Bloch I. Institution: LTCI, Télécom Paris, Institut Polytechnique de Paris, Paris, France. Abstract: Patient-specific 3D modeling is the first step towards image-guided surgery, the actual revolution in surgical care. Pediatric and adolescent patients with rare tumors and malformations should highly benefit from these latest technological innovations, allowing personalized tailored surgery. This study focused on the pelvic region, located at the crossroads of the urinary, digestive, and genital channels with important vascular and nervous structures. The aim of this study was to evaluate the performances of different software tools to obtain patient-specific 3D models, through segmentation of magnetic resonance images (MRI), the reference for pediatric pelvis examination. Twelve software tools freely available on the Internet and two commercial software tools were evaluated using T2-w MRI and diffusion-weighted MRI images. The software tools were rated according to eight criteria, evaluated by three different users: automatization degree, segmentation time, usability, 3D visualization, presence of image registration tools, tractography tools, supported OS, and potential extension (i.e., plugins). A ranking of software tools for 3D modeling of MRI medical images, according to the set of predefined criteria, was given. This ranking allowed us to elaborate guidelines for the choice of software tools for pelvic surgical planning in pediatric patients. The best-ranked software tools were Myrian Studio, ITK-SNAP, and 3D Slicer, the latter being especially appropriate if nerve fibers should be included in the 3D patient model. To conclude, this study proposed a comprehensive review of software tools for 3D modeling of the pelvis according to a set of eight criteria and delivered specific conclusions for pediatric and adolescent patients that can be directly applied to clinical practice. |
New, Simple and Reliable Volumetric Calculation Technique in Incisional Hernias with Loss of Domain
|
Publication: Hernia. 2019 Jun 19. PMID: 31218439 Authors: Martre P, Sarsam M, Tuech JJ, Coget J, Schwarz L, Khalil H. Institution: Department of Digestive Surgery, Rouen University Hospital, Rouen, France. Abstract: INTRODUCTION: The management of hernias with loss of domain is a challenging problem. It has been shown that the volume of the incisional hernia/peritoneal volume ratio < 20% was a predictive factor for tension-free fascia closure, after pre-operative pneumoperitoneum preparation (Goni Moreno technique). In this study, we propose an easy, reliable and fast technique to perform volumetric calculation, by the surgeon alone. MATERIALS AND METHODS: 3D Slicer software (free open-source software) was used to calculate with precision the intra-peritoneal and intra-hernia volumes, and to create a 3D reconstruction of both volumes. The measurement technique is described step by step using detailed figures and videos. RESULTS: The method was used to calculate the volumes for five consecutive patients, managed between January 2018 and March 2019. All the five patients had a ratio greater than 20% and, therefore, received a PPP program. The effectiveness of the procedure is objectified by the increase of the intraabdominal volume and the reduction of the incisional hernia/peritoneal volume ratio. The feasibility of a tension-free fascia closure was confirmed for the five patients. CONCLUSION: In addition to a standardized definition of "loss of domain", a standardized volumetric technique, easy to reproduce, needs to be adopted. Our method can be done by any surgeon with basic computer skills and radiological knowledge in an autonomous and a fast manner, thus helping to select the right technique for the right patient. |
Predicting Malignant Potential of Subsolid Nodules: Can Radiomics Preempt Longitudinal Follow up CT?
|
Publication: Cancer Imaging. 2019 Jun 10;19(1):36. PMID: 31182167 | PDF Authors: Digumarthy SR, Padole AM, Rastogi S, Price M, Mooradian MJ, Sequist LV, Kalra MK. Institution: Department of Radiology, Massachusetts General Hospital, Boston, MA, USA. Abstract: BACKGROUND: To assess if radiomics can differentiate benign and malignant subsolid lung nodules (SSNs) on baseline or follow up chest CT examinations. If radiomics can differentiate between benign and malignant subsolid lung nodules, the clinical implications are shorter follow up CT imaging and early recognition of lung adenocarcinoma on imaging. MATERIALS AND METHODS: The IRB approved retrospective study included 36 patients (mean age 69 ± 8 years; 5 males, 31 females) with 108 SSNs (31benign, 77 malignant) who underwent follow up chest CT for evaluation of indeterminate SSN. All SSNs were identified on both baseline and follow up chest CT. DICOM CT images were deidentified and exported into the open access 3D Slicer v.4.7 software to obtain radiomic features. Logistic regression analyses and receiver operating characteristic (ROC) curves for various quantitative parameters were generated with SPSS statistical software. RESULTS: Only 2/92 radiomic features (cluster shade and surface volume ratio) enabled differentiation between malignant and benign SSN on baseline chest CT (P = 0.01 and 0.03) with moderate accuracy [AUC 0.624 (0.505-0.743)]. On follow-up CT, 52/92 radiomic features were significantly different between benign and malignant SSN (P: 0.04 - < 0.0001) with improved accuracy [AUC: 0.708 (0.605-0.811), P = 0.04 - < 0.0001]. Radiomics of benign SSN were stable over time, whereas 63/92 radiomic features of malignant SSNs changed significantly between the baseline and follow up chest CT (P: 0.04 - < 0.0001). CONCLUSIONS: Temporal changes in radiomic features of subsolid lung nodules favor malignant etiology over benign. The change in radiomics features of subsolid lung nodules can allow shorter follow up CT imaging and early recognition of lung adenocarcinoma on imaging. Radiomic features have limited application in differentiating benign and early malignant SSN on baseline chest CT. |
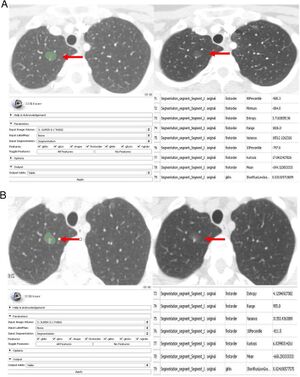 a Radiomic analysis of a malignant SSN on baseline chest CT of a 62-year-old woman. b Radiomic analysis of a malignant SSN of the same patient 3 years later. Radiomic features (e.g., entropy, kurtosis, mean, grey level variance) were substantially different on the follow-up CT compared to the baseline CT. |
Imaging Phenotype Using Radiomics to Predict Dry Pleural Dissemination in Non-Small Cell Lung Cancer
|
Publication: Ann Transl Med. 2019 Jun;7(12):259. PMID: 31355226 | PDF Authors: Yang M, Ren Y, She Y, Xie D, Sun X, Shi J, Zhao G, Chen C. Institution: Department of Cardiothoracic Surgery, Ningbo No. 2 Hospital, Ningbo, China. Abstract: BACKGROUND: Dry pleural dissemination (DPD) in non-small cell lung cancer (NSCLC) is defined as having solid pleural metastases without malignant pleural effusion. We aim to identify DPD by applying radiomics, a novel approach to decode the tumor phenotype. METHODS: Preoperative chest computed tomographic images and basic clinical feature were retrospectively evaluated in patients with surgically resected NSCLC between January 1, 2015 and December 31, 2016. Propensity score was applied to match the DPD and non-DPD groups. One thousand and eighty radiomics features were quantitatively extracted by the 3D Slicer software and "pyradiomics" package. Least absolute shrinkage and selection operator (LASSO) binary model was applied for feature selection and developing the radiomics signature. The discrimination was evaluated using area under the curve (AUC) and Youden index. RESULTS: Sixty-four DPD patients and paired 192 non-DPD patients were enrolled. Using the LASSO model, this study developed a radiomics signature including 10 radiomic features. The mean ± standard deviation values of the radiomics signature with DPD status (-2.129±1.444) was significantly higher compared to those with non-DPD disease (0.071±0.829, P<0.001). The ten-feature based signature showed good discrimination between DPD and non-DPD, with an AUC of 0.93 (95% confidence-interval, 0.891-0.958) respectively. The sensitivity and specificity of the radiomics signature was 85.94% and 85.94%, with the optimal cut-off value of -0.696 and Youden index of 0.71. CONCLUSIONS: The signature based on radiomics features can provide potential predictive value to identify DPD in patients with NSCLC. |
|
Publication: Acta Neurochir (Wien). 2019 May;161(5):917-23. PMID: 30937608 Authors: Minkin K, Gabrovski K, Sirakov S, Penkov M, Todorov Y, Karakostov V, Dimova P. Institution: Department of Neurosurgery, University Hospital "Saint Ivan Rilski", Sofia, Bulgaria. Abstract: OBJECTIVES: Epilepsy surgery is mainly cortical surgery and the precise definition of the epileptogenic zone on the complex cortical surface is of paramount importance. Stereoelectroencephalography (SEEG) may delineate the epileptogenic zone even in cases of non-lesional epilepsy. The aim of our study was to present a technique of 3D neuronavigation based on the brain surface and SEEG electrodes reconstructions using FSL and 3D Slicer software. PATIENTS AND METHODS: Our study included 26 consecutive patients operated on for drug-resistant epilepsy after SEEG exploration between January 2015 and December 2017. All patients underwent 1.5 T pre-SEEG MRI, post-SEEG CT, DICOM data post-processing using FSL and 3D Slicer, preoperative planning on 3D Slicer, and intraoperative 3D neuronavigation. Accuracy and precision of 3D SEEG reconstruction and 3D neuronavigation was assessed. RESULTS: We identified 125 entry points of SEEG electrodes during 26 operations. The accuracy of 3D reconstruction was 0.8 mm (range, 0-2 mm) with a precision of 1.5 mm. The accuracy of 3D SEEG neuronavigation was 2.68 mm (range, 0-6 mm). The precision of 3D neuronavigation was 1.48 mm. CONCLUSION: 3D neuronavigation for SEEG-guided epilepsy surgery using free software for post-processing of common MRI sequences is possible and a reliable method even with navigation systems without a brain extraction tool. |
The Future of Biomechanical Spine Research: Conception and Design of a Dynamic 3D Printed Cervical Myelography Phantom
|
Publication: Cureus. 2019 May 3;11(5):e4591. PMID: 31309016 Authors: Clifton W, Nottmeier E, Damon A, Dove C, Pichelmann M. Institution: Neurosurgery, Mayo Clinic, Jacksonville, USA. Abstract: Background Three-dimensional (3D) printing is a growing practice in the medical community for patient care and trainee education as well as production of equipment and devices. The development of functional models to replicate physiologic systems of human tissue has also been explored, although to a lesser degree. Specifically, the design of 3D printed phantoms that possess comparable biomechanical properties to human cervical vertebrae is an underdeveloped area of spine research. In order to investigate the functional uses of cervical 3D printed models for replicating the complex physiologic and biomechanical properties of the human subaxial cervical spine, our institution has created a prototype that accurately reflects these properties and provides a novel method of assessing spinal canal dimensions using simulated myelography. To our knowledge, this is the first 3D printed phantom created to study these parameters. Materials and methods A de-identified cervical spine computed tomography imaging file was segmented using threshold modulation in 3D Slicer software. The subaxial vertebrae (C3-C7) of the scan were individualized by separating the facet joint spaces and uncovertebral joints within the software in order to create individual stereolithography (STL) files. Each individual vertebra was printed on an Ultimaker S5 dual-extrusion printer using white "tough" polylactic acid filament. A human cadaveric subaxial cervical spine was harvested to provide a control for our experiment. Both models were assessed and compared in flexion and extension dynamic motion grossly and fluoroscopically. The maximum angles of deformation on X-ray imaging were recorded using DICOM (Digital Imaging and Communications in Medicine) viewing software. In order to compare the ability to assess canal dimensions of the models using fluoroscopic imaging, a myelography simulation was designed. Results The cervical phantom demonstrated excellent ability to resist deformation in flexion and extension positions, attributed to the high quality of initial segmentation. The gross and fluoroscopic dynamic movement of the phantom was analogous to the cadaver model. The myelography simulator adequately demonstrated the canal dimensions in static and dynamic positions for both models. Pertinent anatomic landmarks were able to be effectively visualized for assessment of canal measurements for sagittal and transverse dimensions. Conclusions By utilizing the latest technologies in DICOM segmentation and 3D printing, our institution has created the first cervical myelography phantom for biomechanical evaluation and trainee instruction. By combining new technologies with anatomical knowledge, quality 3D printing shows great promise in becoming a standard player in the future of spinal biomechanical research. |
Treatment of Intracranial Hemorrhage With Neuroendoscopy Guided by Body Surface Projection
|
Publication: Medicine (Baltimore). 2019 May;98(19):e15503. PMID: 31083190 | PDF Authors: Qiu S, Liu T, Cao G, Wu K, Zhao T. Institution: Department of Neurosurgery, The Second People's Hospital of Hefei, Hefei Hospital Affiliated to Medical University of Anhui, Hefei, China. Abstract: BACKGROUND: We aimed to study the feasibility of body surface projection in neuroendoscopic treatment of intracranial hemorrhage (ICH), and to evaluate the prognosis of muscle strength using diffusion tensor imaging (DTI) technique. METHODS: We utilized 3D Slicer software and adopted hematoma body surface projection orientation to eliminate ICH by using neuroendoscope for 69 cases of spontaneous intracerebral hemorrhage. The standard of correct location was determined by the direct view of hematoma at the first operation. Evacuation rate by comparing computed tomography (CT) before and after the surgery and Glasgow coma scale (GCS) was computed. DTI was used for pyramidal tract imaging 3 weeks after the operation, while the prognosis of muscle strength was assessed after 6 months. The control group included 69 patients with basal ganglia hemorrhage who received conservative treatment during the same period. RESULTS: The hematoma evacuation rate was 90.75% in average. The average GCS score rose by 4 points one week after the surgery. The shape of pyramidal tract affected the prognosis of body muscle strength, and the simple disruption type was the worst. There was no difference in mortality between the surgery group (10.1%) and the conservative group (4.3%). The muscle strength improvement value and modulate RANK score (MRS) in the surgery group were better than the control group. CONCLUSION: It is convenient and feasible to use the surface projection to determine the target of operation, and the clearance rate of hematoma is high. Pyramidal tract imaging can predict the prognosis of muscle strength. |
Alterations in Brain Neurocircuitry Following Treatment With the Chemotherapeutic Agent Paclitaxel in Rats
|
Publication: Neurobiol Pain. 2019 May 27;6:100034. PMID: 31223138 | PDF Authors: Ferris CF, Nodine S, Pottala T, Cai X, Knox TM, Fofana FH, Kim S, Kulkarni P, Crystal JD, Hohmann AG. Institution: Center for Translational NeuroImaging, Northeastern University, Boston, MA, USA. Abstract: Human and animal studies suggest that both traumatic nerve injury and toxic challenge with chemotherapeutic agents involves the reorganization of neural circuits in the brain. However, there have been no prospective studies, human or animal, using magnetic resonance imaging (MRI) to identify changes in brain neural circuitry that accompany the development of chemotherapy-induced neuropathic pain (i.e. within days following cessation of chemotherapy treatment and without the confound cancer). To this end, different MRI protocols were used to ascertain whether a reorganization of brain neural circuits is observed in otherwise normal rats exposed to the taxane chemotherapeutic agent paclitaxel. We conducted an imaging study to evaluate the impact of a well-established paclitaxel dosing regimen, validated to induce allodynia in control rats within eight days of treatment, on brain neural circuitry. Rats received either paclitaxel (2 mg/kg/day i.p; cumulative dose of 8 mg/kg) or its vehicle four times on alternate days (i.e. day 0, 2, 4, 6). Following the cessation of treatments (i.e. on day 8), all rats were tested for responsiveness to cold followed by diffusion weighted magnetic resonance imaging and assessment of resting state functional connectivity. Imaging data were analyzed using a 3D MRI rat with 173 segmented and annotated brain areas. Paclitaxel-treated rats were more sensitive to a cold stimulus compared to controls. Diffusion weighted imaging identified brain areas involved in the emotional and motivational response to chronic pain that were impacted by paclitaxel treatment. Affected brain regions included the prefrontal cortex, amygdala, hippocampus, hypothalamus and the striatum/nucleus accumbens. This putative reorganization of gray matter microarchitecture formed a continuum of brain areas stretching from the basal medial/lateral forebrain to the midbrain. Resting state functional connectivity showed reorganization between the periaqueductal gray, a key node in nociceptive neural circuitry, and connections to the brainstem. Our results, employing different imaging modalities to assess the central nervous system effects of chemotherapy, fit the theory that chronic pain is regulated by emotion and motivation and influences activity in the periaqueductal gray and brainstem to modulate pain perception. "...Brain tissue masks for resting-state functional images were manually drawn using 3D Slicer and applied for skull-stripping.." Funding:
|
Comprehensive Analysis of Animal Models of Cardiovascular Disease using Multiscale X-Ray Phase Contrast Tomography
|
Publication: Sci Rep. 2019 May 6;9(1):6996. PMID: 31061429 | PDF Authors: Dejea H, Garcia-Canadilla P, Cook AC, Guasch E, Zamora M, Crispi F8, Stampanoni M, Bijnens B, Bonnin A. Institution: Paul Scherrer Institut, Villigen PSI, Switzerland. Abstract: Cardiovascular diseases (CVDs) affect the myocardium and vasculature, inducing remodelling of the heart from cellular to whole organ level. To assess their impact at micro and macroscopic level, multi-resolution imaging techniques that provide high quality images without sample alteration and in 3D are necessary: requirements not fulfilled by most of current methods. In this paper, we take advantage of the non-destructive time-efficient 3D multiscale capabilities of synchrotron Propagation-based X-Ray Phase Contrast Imaging (PB-X-PCI) to study a wide range of cardiac tissue characteristics in one healthy and three different diseased rat models. With a dedicated image processing pipeline, PB-X-PCI images are analysed in order to show its capability to assess different cardiac tissue components at both macroscopic and microscopic levels. The presented technique evaluates in detail the overall cardiac morphology, myocyte aggregate orientation, vasculature changes, fibrosis formation and nearly single cell arrangement. Our results agree with conventional histology and literature. This study demonstrates that synchrotron PB-X-PCI, combined with image processing tools, is a powerful technique for multi-resolution structural investigation of the heart ex-vivo. Therefore, the proposed approach can improve the understanding of the multiscale remodelling processes occurring in CVDs, and the comprehensive and fast assessment of future interventional approaches. "...Cell aggregates as well as the lumen of vessel sections were segmented using 3D Slicer v4.5, by manually labelling individual 2D image slices and performing a 3D interpolation." |
3D Superimposition of Craniofacial Imaging-The Utility of Multicentre Collaborations
|
Publication: Orthod Craniofac Res. 2019 May;22 Suppl 1:213-220. PMID: 31074129 | PDF Authors: Yatabe M, Prieto JC, Styner M, Zhu H, Ruellas AC, Paniagua B, Budin F, Benavides E, Shoukri B, Michoud L, Ribera N, Cevidanes L. Institution: Department for Orthodontics and Pediatric Dentistry, University of Michigan, Ann Arbor, Michigan, USA. Abstract: Clinical applications of 3D image registration and superimposition have contributed to better understanding growth changes and clinical outcomes. The use of 3D dental and craniofacial imaging in dentistry requires validate image analysis methods for improved diagnosis, treatment planning, navigation and assessment of treatment response. Volumetric 3D images, such as cone-beam computed tomography, can now be superimposed by voxels, surfaces or landmarks. Regardless of the image modality or the software tools, the concepts of regions or points of reference affect all quantitative of qualitative assessments. This study reviews current state of the art in 3D image analysis including 3D superimpositions relative to the cranial base and different regional superimpositions, the development of open source and commercial tools for 3D analysis, how this technology has increased clinical research collaborations from centres all around the globe, some insight on how to incorporate artificial intelligence for big data analysis and progress towards personalized orthodontics. Funding:
|
Complete Thoracolumbar Fracture-dislocation with Intact Neurologic Function: Explanation of a Novel Cord Saving Mechanism
|
Publication: J Spinal Cord Med. 2018 May;41(3):367-76. PMID: 28648115 | PDF Authors: Rahimizadeh A, Asgari N, Rahimizadeh A. Institution: Department of Neurosurgery, Pars Advanced and Minimally Invasive Medical Manners Research Center, Pars Hospital, Iran University of Medical Science, Tehran, Iran. Abstract: BACKGROUND: The thoracolumbar junction from T11 to L2 is a common site of injury in which fracture and dislocations are the most prevalent ones occurring at this location. Fracture dislocation is defined as failure of all three columns of the spine with gross displacement. Considering the significant violence necessary to produce fracture dislocations, these injuries are often associated with major neural deficit, with the majority of casualties becoming paraplegic immediately. Preservation of neurological function following complete fracture dislocation is quite rare entity. OBJECTIVE: To represent the possibility of existence of a preservation mechanism for functional integrity of cord despite spinal gross fracture dislocation by reproducing the injury on a plastic model and simulating a corresponding model using 3D Slicer software, detailed description the pathomechanism of neurologic sparing. CASE REPORT: A 19-year-old female who sustained severe thoracolumbar fracture dislocation but with normal neurology is presented. Despite the severity of the condition, the diagnosis was initially missed due to associated vital injuries. RESULTS: Combined posterior and anterior surgery resulted in optimal coronal and sagittal alignment, as well as proper stabilization without any complication. At 9-year follow-up, the patient was found to be doing well. CONCLUSION: The prognosis for complete recovery with preplanned surgical intervention in thoracolumbar injuries affecting all three columns but with normal neurologic function is promising based on images, plastic models and 3D simulated model based on digital images. |
Whole-Body FDG PET/MR Atlas for Multiparametric Voxel-Based Analysis
|
Publication: Sci Rep. 2019 Apr 16;9(1):6158. PMID: 30992502 | PDF Authors: Sjöholm T, Ekström S, Strand R, Ahlström H, Lind L, Malmberg F, Kullberg J. Institution: Department of Surgical Sciences, Uppsala University, Uppsala, Sweden. Abstract: Quantitative multiparametric imaging is a potential key application for Positron Emission Tomography/Magnetic Resonance (PET/MR) hybrid imaging. To enable objective and automatic voxel-based multiparametric analysis in whole-body applications, the purpose of this study was to develop a multimodality whole-body atlas of functional 18F-fluorodeoxyglucose (FDG) PET and anatomical fat-water MR data of adults. Image registration was used to transform PET/MR images of healthy control subjects into male and female reference spaces, producing a fat-water MR, local tissue volume and FDG PET whole-body normal atlas consisting of 12 male (66.6 ± 6.3 years) and 15 female (69.5 ± 3.6 years) subjects. Manual segmentations of tissues and organs in the male and female reference spaces confirmed that the atlas contained adequate physiological and anatomical values. The atlas was applied in two anomaly detection tasks as proof of concept. The first task automatically detected anomalies in two subjects with suspected malignant disease using FDG data. The second task successfully detected abnormal liver fat infiltration in one subject using fat fraction data. "...All segmentations were performed manually in 3D Slicer using transaxial fat and water fraction MR atlas images in reference space. " |
Complexity of Tumor Shape, Spiculatedness, Correlates With Tumor Radiomic Shape Features
|
Publication: Sci Rep. 2019 Mar 13;9(1):4329. PMID: 30867443 | PDF Authors: Limkin EJ, Reuzé S, Carré A, Sun R, Schernberg A, Alexis A, Deutsch E, Ferté C, Robert C. Institution: Gustave Roussy, Université Paris-Saclay, Department of Radiotherapy, Villejuif, France. Abstract: Radiomics extracts high-throughput quantitative data from medical images to contribute to precision medicine. Radiomic shape features have been shown to correlate with patient outcomes. However, how radiomic shape features vary in function of tumor complexity and tumor volume, as well as with method used for meshing and voxel resampling, remains unknown. The aims of this study are to create tumor models with varying degrees of complexity, or spiculatedness, and evaluate their relationship with quantitatively extracted shape features. Twenty-eight tumor models were mathematically created using spherical harmonics with the spiculatedness degree d being increased by increments of 3 (d = 11 to d = 92). Models were 3D printed with identical bases of 5 cm, imaged with a CT scanner with two different slice thicknesses, and semi-automatically delineated. Resampling of the resulting masks on a 1 × 1 × 1 mm3 grid was performed, and the voxel size of each model was then calculated to eliminate volume differences. Four MATLAB-based algorithms (isosurface (M1), isosurface filter (M2), isosurface remeshing (M3), and boundary (M4)) were used to extract nine 3D features (Volume, Surface area, Surface-to-volume, Compactness1, Compactness2, Compactness3, Spherical Disproportion, Sphericity and Fractional Concavity). To quantify the impact of 3D printing, acquisition, segmentation and meshing, features were computed directly from the stereolithography (STL) file format that was used for 3D printing, and compared to those computed. Changes in feature values between 0.6 and 2 mm slice acquisitions were also compared. Spearman's rank-order correlation coefficients were computed to determine the relationship of each shape feature with spiculatedness for each of the four meshing algorithms. Percent changes were calculated between shape features extracted from the original and resampled contoured images to evaluate the influence of spatial resampling. Finally, the percent change in shape features when the volume was changed from 25% to 150% of their original volume was quantified for three distinct tumor models and compared to the percent change observed when modifying the spiculatedness of the model from d = 11 to d = 92. Values extracted using isosurface remeshing method are the closest to the STL reference ones, with mean differences less than 10.8% (Compactness2) for all features. Seven of the eight features had strong significant correlations with tumor model complexity irrespective of the meshing algorithm (r > 0.98, p < 10-4), with fractional concavity having the lowest correlation coefficient (r = 0.83, p < 10-4, M2). Comparisons of features extracted from the 0.6 and 2 mm slice thicknesses showed that mean differences were from 2.1% (Compactness3) to 12.7% (Compactness2) for the isosurface remeshing method. Resampling on a 1 × 1 × 1 mm3 grid resulted in between 1.3% (Compactness3) to 9.5% (Fractional Concavity) mean changes in feature values. Compactness2, Compactness3, Spherical Disproportion, Sphericity and Fractional Concavity were the features least affected by volume changes. Compactness1 had a 90.4% change with volume, which was greater than the change between the least and most spiculated models. This is the first methodological study that directly demonstrates the relationship of tumor spiculatedness with radiomic shape features, that also produced 3D tumor models, which may serve as reference phantoms for future radiomic studies. Surface Area, Surface-to-volume, and Spherical Disproportion had direct relationships with spiculatedness while the three formulas for Compactness, Sphericity and Fractional Concavity had inverse relationships. The features Compactness2, Compactness3, Spherical Disproportion, and Sphericity should be prioritized as these have minimal variations with volume changes, slice thickness and resampling. "...Scans were contoured using the thresholding function of 3D Slicer... " |
Virtual Reconstruction of Paranasal Sinuses from CT Data: A Feasibility Study for Forensic Application
|
Publication: Diagn Interv Imaging. 2019 Mar;100(3):163-8. PMID: 30553743 Authors: Gach P, Tuchtan-Torrents L, Delteil C, Adalian P, Piercecchi MD, Ebert LC, Gorincour G. Institution: LiiE, EA 4264, CERIMED, Aix-Marseille University, 13005 Marseille, France. Abstract: PURPOSE: The purpose of this study was to report the feasibility of computed modelization and reconstitution of the paranasal sinuses, before and after trauma, from CT data. MATERIALS AND METHODS: We modeled and reconstructed the paranasal sinuses of two patients (A and B), before and after trauma, using two different softwares (3D Slicer and Blender®). Both patients had different numbers and locations of fractures. The 3D Slicer software was used to create a 3D model from CT data. We then imported the 3D data into the Blender® software, to reconstruct and compare the dimensions of the paranasal sinuses before and after trauma. RESULTS: The 3 fragments of patient A and the 7 fragments of patient B could be repositioned in the pre-traumatic configuration. Distance measurements proved to be similar between pre- and post-traumatic 3D volumes. CONCLUSION: After simple trauma, bone facial anatomy reconstruction is manually feasible. The whole procedure could benefit from automatization through machine learning. However, this feasibility must be confirmed on more severely fractured paranasal sinuses, to consider an application in forensic identification. |
Integrated in Utero MR Method for Assessing Structural Brain Abnormalities and Measuring Intracranial Volumes in Fetuses With Congenital Heart Disease: Results of a Prospective Case-Control Feasibility Study
|
Publication: Neuroradiology. 2019 May;61(5):603-11. PMID: 30796469 | PDF Authors: Griffiths PD, Mousa HA, Finney C, Mooney C, Mandefield L, Chico TJA, Jarvis D. Institution: Academic Unit of Radiology, University of Sheffield, Floor C, Royal Hallamshire Hospital, Glossop Road, Sheffield, UK. Abstract: PURPOSE: To refine methods that assess structural brain abnormalities and calculate intracranial volumes in fetuses with congenital heart diseases (CHD) using in utero MR (iuMR) imaging. Our secondary objective was to assess the prevalence of brain abnormalities in this high-risk cohort and compare the brain volumes with normative values. METHODS: We performed iuMR on 16 pregnant women carrying a fetus with CHD and gestational age ≥ 28-week gestation and no brain abnormality on ultrasonography. All cases had fetal echocardiography by a pediatric cardiologist. Structural brain abnormalities on iuMR were recorded. Intracranial volumes were made from 3D FIESTA acquisitions following manual segmentation and the use of 3D Slicer software and were compared with normal fetuses. Z scores were calculated, and regression analyses were performed to look for differences between the normal and CHD fetuses. RESULTS: Successful 2D and 3D volume imaging was obtained in all 16 cases within a 30-min scan. Despite normal ultrasonography, 5/16 fetuses (31%) had structural brain abnormalities detected by iuMR (3 with ventriculomegaly, 2 with vermian hypoplasia). Brain volume, extra-axial volume, and total intracranial volume were statistically significantly reduced, while ventricular volumes were increased in the CHD cohort. CONCLUSION: We have shown that it is possible to perform detailed 2D and 3D studies using iuMR that allow thorough investigation of all intracranial compartments in fetuses with CHD in a clinically appropriate scan time. Those fetuses have a high risk of structural brain abnormalities and smaller brain volumes even when brain ultrasonography is normal. |
Novel Application and Validation of in Vivo Micro-CT to Study Bone Modelling in 3D
|
Publication: Orthod Craniofac Res. 2019 May;22 Suppl 1:90-5. PMID: 31074146 | PDF Authors: Oz U, Ruellas AC, Westgate PM, Cevidanes LH, Huja SS. Institution: Department of Orthodontics, Near East University School of Dentistry, Nicosia, Northern Cyprus. Abstract: OBJECTIVES: The aim is to highlight a novel three-dimensional (3D) imaging methodology using micro-CT scans to visualize and measure bone modelling in an animal model. In order to validate the new methodology, we compared the 3D imaging method to traditional two-dimensional (2D) histomorphometry to assess growth changes in the jaws of a rodent. SETTING AND SAMPLE POPULATION: Rodent animal models. MATERIAL AND METHODS: Eleven rats were obtained from a larger previously published study. Sixty undecalcified histological sections from the maxilla and corresponding high-resolution in vivo micro-CT reconstructions were obtained. Bone modelling changes on specific alveolar surfaces were measured using traditional histomorphometry. Measurements of bone growth were also obtained via 3D Slicer software from 3D micro-CT generated models from the same plane containing the histological images. Both qualitative and quantitative 3D methods were compared to traditional histological measurements. Quantitative agreement between methods was categorized as follows: poor (>150 μm), good (150-100 μm) and excellent (<100 μm). RESULTS: Both qualitative (88.3%) and quantitative (86.7%) 3D measurements showed excellent agreement, when compared to histomorphometric measurements. Only 1.7% and 5% of the comparisons exhibited poor agreement (>150 μm) for qualitative and quantitative methods, respectively. DISCUSSION: The new 3D superimposition method compares very favourably with traditional histology. It is likely that in the future, such methods will be used in studies of bone adaptation. CONCLUSION: The 3D micro-CT qualitative and quantitative methods are reliable for measuring bone modelling changes and compare favourably to histology for the specific application described. |
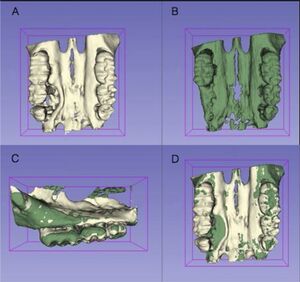 Pre (T1) and post (T2) sacrifice 3D models. A, Occlusal view of 3D pre-treatment model. B, Occlusal view of post-sacrifice 3D model. Right side maxillary 2nd and 3rd molars were extracted. C, Left sagittal registered and superimposed view. D, Occlusal view of pre-, and post-sacrifice 3D registered and superimposed models (Voxel-based registration by 3D Slicer software). |
Human Inner-ear Malformation Types Captured in 3D
|
Publication: J Int Adv Otol. 2019 Apr;15(1):77-82. PMID: 31058598 | PDF Authors: Dhanasingh A, Dietz A, Jolly C, Roland P. Institution: MED-EL GmbH, Implants, Innsbruck, Austria. Abstract: OBJECTIVES: Capture the human inner-ear malformation types in 3D by segmenting the inner-ear structures from clinical CT (computed tomography) and MR (magnetic resonance) image datasets. Volumetric analysis was done to find the variations in the volume of cochlear part alone from complete inner-ear followed by 3D printing from the 3D segmented models. MATERIALS AND METHODS: Using 3D Slicer freeware, the complete inner-ear structures were segmented from anonymized CT and MR image by setting a tight grey-scale threshold to avoid capturing unwanted structures followed by volumetric analysis of the cochlear part alone. 3D printing was done using Form labs desktop 3D printer. RESULTS: We identified 2x normal anatomy (NA) cochlea, 1x enlarged vestibular aqueduct syndrome (EVAS), 3x incomplete partition (IP) type-I, 4x IP type-II, 3x IP type-III, 5x common cavity (CC) and 5x cochlear hypoplasia (CH). 3D segmented models along with the 3D printed models showed huge variation in size, shape and the anatomy among the image data-sets analyzed. Volumetric analysis showed that on average, volume of CC was above 150mm3, volume of CH fell below 80mm3, Volume of NA, EVAS and IP-I were all around 85-105mm3 whereas the volume of IP-II was around 50mm3. CONCLUSION: Visualizing human inner-ear malformation types in 3D both as computer models and as 3D printed models is a whole-new experience as demonstrated in this study. The volumetric analysis showed a huge variation among the volume of cochlear part alone among the malformation types. |
Bridging the Translational Gap: Implementation of Multimodal Small Animal Imaging Strategies for Tumor Burden Assessment in a Co-Clinical Trial
|
Publication: PLoS One. 2019 Apr 8;14(4):e0207555. PMID: 30958825 | PDF Authors: Blocker SJ, Mowery YM, Holbrook MD, Qi Y, Kirsch DG, Johnson GA, Badea CT. Institution: Center for In Vivo Microscopy, Duke University Medical Center, Durham, NC, USA. Abstract: In designing co-clinical cancer studies, preclinical imaging brings unique challenges that emphasize the gap between man and mouse. Our group is developing quantitative imaging methods for the preclinical arm of a co-clinical trial studying immunotherapy and radiotherapy in a soft tissue sarcoma model. In line with treatment for patients enrolled in the clinical trial SU2C-SARC032, primary mouse sarcomas are imaged with multi-contrast micro-MRI (T1 weighted, T2 weighted, and T1 with contrast) before and after immune checkpoint inhibition and pre-operative radiation therapy. Similar to the patients, after surgery the mice will be screened for lung metastases with micro-CT using respiratory gating. A systems evaluation was undertaken to establish a quantitative baseline for both the MR and micro-CT systems against which others systems might be compared. We have constructed imaging protocols which provide clinically-relevant resolution and contrast in a genetically engineered mouse model of sarcoma. We have employed tools in 3D Slicer for semi-automated segmentation of both MR and micro-CT images to measure tumor volumes efficiently and reliably in a large number of animals. Assessment of tumor burden in the resulting images was precise, repeatable, and reproducible. Furthermore, we have implemented a publicly accessible platform for sharing imaging data collected during the study, as well as protocols, supporting information, and data analyses. In doing so, we aim to improve the clinical relevance of small animal imaging and begin establishing standards for preclinical imaging of tumors from the perspective of a co-clinical trial. |
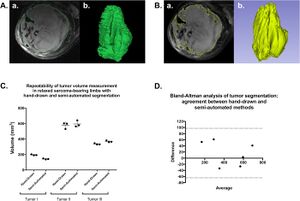 Semi-automated tumor segmentation with 3D Slicer is an acceptable alternative to hand-drawn ROIs for sarcoma volume measurement. T2-weighted images of three tumors of varying size and morphology were analyzed with hand-drawn ROIs and semi-automated segmentation to calculate tumor volume. Examples of hand-drawn (A) and semi-automated (B) segmentation of the same tumor are shown, including an axial slice of the segmented tumor (a.) and the rendered 3D representation of the tumor according to each segmentation (b.). Resulting volume calculations were compared between triplicate measures with each method for each tumor (C), demonstrating similar measurements and comparable precision. Bland-Altman analysis of both methods, when employed in 6 separate tumors, indicated that semi-automated segmentation is a viable alternative to hand-drawn segmentation (D). |
Validation of a Freehand Technique for Cortical Bone Trajectory Screws in the Lumbar Spine
|
Publication: J Neurosurg Spine. 2019 Apr 19:1-8. PMID: 31003218 Authors: Tan Z, McLachlin S, Whyne C, Finkelstein J. Institution: Division of Orthopaedic Surgery, University of Toronto, Toronto, Canada. Abstract: OBJECTIVE : The cortical bone trajectory (CBT) technique for pedicle screw placement has gained popularity among spinal surgeons. It has been shown biomechanically to provide better fixation and improved pullout strength compared to a traditional pedicle screw trajectory. The CBT technique also allows for a less invasive approach for fusion and may have lower incidence of adjacent-level disease. A limitation of the current CBT technique is a lack of readily identifiable and reproducible visual landmarks to guide freehand CBT screw placement in comparison to the well-defined identifiable landmarks for traditional pedicle screw insertion. The goal of this study was to validate a safe and intuitive freehand technique for placement of CBT screws based on optimization of virtual CBT screw placement using anatomical landmarks in the lumbar spine. The authors hypothesized that virtual identification of anatomical landmarks on 3D models of the lumbar spine generated from CT scans would translate to a safe intraoperative freehand technique. METHODS: Customized, open-source medical imaging and visualization software 3D Slicer was used in this study to develop a workflow for virtual simulation of lumbar CBT screw insertion. First, in an ex vivo study, 20 anonymous CT image series of normal and degenerative lumbar spines and virtual screw insertion were conducted to place CBT screws bilaterally in the L1-5 vertebrae for each image volume. The optimal safe CBT trajectory was created by maximizing both the screw length and the cortical bone contact with the screw. Easily identifiable anatomical surface landmarks for the start point and trajectory that best allowed the reproducible idealized screw position were determined. An in vivo validation of the determined landmarks from the ex vivo study was then performed in 10 patients. Placement of virtual "test" cortical bone trajectory screws was simulated with the surgeon blinded to the real-time image-guided navigation, and the placement was evaluated. The surgeon then placed the definitive screw using image guidance. RESULTS: From the ex vivo study, the optimized technique and landmarks were similar in the L1-4 vertebrae, whereas the L5 optimized technique was distinct. The in vivo validation yielded ideal, safe, and unsafe screws in 62%, 16%, and 22% of cases, respectively. A common reason for the nonidealized trajectories was the obscuration of patient anatomy secondary to severe degenerative changes. CONCLUSIONS: CBT screws were placed ideally or safely 78% of the time in a virtual simulation model. A 22% rate of unsafe freehand trajectories suggests that the CBT technique requires use of image-guided navigation or x-ray guidance and that reliable freehand CBT screw insertion based on anatomical landmarks is not reliably feasible in the lumbar spine. |
Three-Dimensional Evaluation of the Root Resorption of Maxillary Incisors After the Orthodontic Traction of Bicortically Impacted Canines: Case Report
|
Publication: Prog Orthod. 2019 Apr 1;20(1):13. PMID: 30931492 | PDF Authors: Arriola-Guillén LE, Rodríguez-Cárdenas YA, Ruíz-Mora GA, Aliaga-Del Castillo A, Schilling J, Dias-Da Silveira HL. Institution:Division of Orthodontics, School of Dentistry, Universidad Científica del Sur, Miraflores, Lima, Perú. Abstract: BACKGROUND: The root resorption of the maxillary incisors after the orthodontic traction of impacted canines is a concern for clinicians. The aim of this case series report was to evaluate the root resorption of the maxillary incisors after traction until the occlusal plane of the bicortically impacted canines (placed between the two cortical bones in the middle of the alveolar process) located in a complex position using three-dimensional superimposition. This case series report describes the root resorption of the maxillary incisors after orthodontic traction with NiTi closed coil springs and a heavy anchorage appliance in three cases of bilateral impacted canines located in a complex position (bicortically) near to midline. Cone-beam computed tomographies (CBCTs) were obtained before and after traction. Root resorption in all root surfaces of the maxillary incisors was evaluated with color-coded maps using the ITK-SNAP and the 3D Slicer software to indicate loss of the root surface (in red) or gain of the surface (in blue) and was quantified in millimeters by the superimposition method. RESULTS: The root changes mainly occurred in the apical third of the maxillary incisor root and did not exceed 2 mm. CONCLUSIONS: Root resorption of the maxillary incisors after the traction of bicortically impacted canines located in a complex position was observed mainly in the apex region, and the amount of root resorption was smaller than 2 mm in all root surfaces. |
Automated 3-Dimensional Magnetic Resonance Imaging Allows for Accurate Evaluation of Glenoid Bone Loss Compared With 3-Dimensional Computed Tomography
|
Publication: Arthroscopy. 2019 Mar;35(3):734-40. PMID: 30733040 Authors: Lansdown DA, Cvetanovich GL, Verma NN, Cole BJ, Bach BR, Nicholson G, Romeo A, Dawe R, Yanke AB. Institution: Department of Orthopedic Surgery, University of California, San Francisco, San Francisco, CA, USA. Abstract: PURPOSE: To evaluate clinical measurements of glenoid bone loss based on 3-dimensional (3D) computed tomography (CT) and automatically segmented 3D reconstructions from Dixon fat-water magnetic resonance (MR) imaging. METHODS: Available CT and MR studies from 16 patients with recurrent anterior shoulder instability were retrospectively reviewed. Three-dimensional reconstructions were formed independently by 2 observers using freely available software and a simple threshold-based segmentation (3D Slicer, v.4.8.0). Bone loss was estimated with the perfect-circle method. Intra-user and interuser reproducibility was determined with intraclass correlation coefficients. Bland-Altman plots were used to evaluate the similarity between imaging modalities. RESULTS: Differences between MR and CT estimates of bone loss ranged from 0% to 6%. The individual intraclass correlation coefficients showed good to excellent reliability, with intraobserver comparisons between MR- and CT-based bone loss estimates ranging from 0.94 to 0.99. Bland-Altman plots showed 95% confidence intervals from -5% to 6% for differences between MR and CT estimates, with 88% of all measurements (42 of 48) showing a less than 2% difference between MR and CT estimates. CONCLUSIONS: The described methodology for obtaining an MR-based 3D reconstruction of the glenoid can evaluate glenoid bone loss similarly to the performance of a 3D CT reconstruction. The results may allow surgeons to simplify the preoperative imaging protocol for patients with recurrent shoulder stabilization and limit the number of shoulder CT scans. LEVEL OF EVIDENCE: Level III, retrospective therapeutic trial. |
Investigation of F-BAR Domain PACSIN Proteins Uncovers Membrane Tubulation Function in Cilia Assembly and Transport
|
Publication: Nat Commun. 2019 Jan 25;10(1):428. PMID: 30683896 | PDF Authors: Insinna C, Lu Q, Teixeira I, Harned A, Semler EM, Stauffer J, Magidson V, Tiwari A, Kenworthy AK, Narayan K, Westlake CJ. Institution: Laboratory of Cellular and Developmental Signaling, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Frederick, MD, USA. Abstract: The intracellular ciliogenesis pathway requires membrane trafficking, fusion, and reorganization. Here, we demonstrate in human cells and zebrafish that the F-BAR domain containing proteins PACSIN1 and -2 play an essential role in ciliogenesis, similar to their binding partner and membrane reorganizer EHD1. In mature cilia, PACSINs and EHDs are dynamically localized to the ciliary pocket membrane (CPM) and transported away from this structure on membrane tubules along with proteins that exit the cilium. PACSINs function early in ciliogenesis at the ciliary vesicle (CV) stage to promote mother centriole to basal body transition. Remarkably, we show that PACSIN1 and EHD1 assemble membrane t7ubules from the developing intracellular cilium that attach to the plasma membrane, creating an extracellular membrane channel (EMC) to the outside of the cell. Together, our work uncovers a function for F-BAR proteins and membrane tubulation in ciliogenesis and explains how the intracellular cilium emerges from the cell. "...Three-dimensional volume segmentation models were generated using 3D Slicer... " Funding:
|
Reproducibility and Non-Redundancy of Radiomic Features Extracted From Arterial Phase CT Scans in Hepatocellular Carcinoma Patients: Impact of Tumor Segmentation Variability
|
Publication: Quant Imaging Med Surg. 2019 Mar;9(3):453-64. PMID: 31032192 | PDF Authors: Qiu Q, Duan J, Duan Z, Meng X, Ma C, Zhu J, Lu J, Liu T, Yin Y. Institution: Department of Radiation Oncology, Shandong Cancer Hospital Affiliated to Shandong University, Jinan, China. Abstract: BACKGROUND: The reproducibility and non-redundancy of radiomic features are challenges in accelerating the clinical translation of radiomics. In this study, we focused on the robustness and non-redundancy of radiomic features extracted from computed tomography (CT) scans in hepatocellular carcinoma (HCC) patients with respect to different tumor segmentation methods. METHODS: Arterial enhanced CT images were retrospectively randomly obtained from 106 patients. As a training data set, 26 HCC patients were used to calculate the features' reproducibility and redundancy. Another data set (55 HCC patients and 25 healthy volunteers) was used for classification. The GrowCut and GraphCut semiautomatic segmentation methods were implemented in 3D Slicer software by two independent observers, and manual delineation was performed by five abdominal radiation oncologists to acquire the gross tumor volume (GTV). Seventy-one radiomic features were extracted from GTVs using Imaging Biomarker Explorer (IBEX) software, including 17 tumor intensity statistical features, 16 shape features and 38 textural features. For each radiomic feature, intraclass correlation coefficient (ICC) and hierarchical clustering were used to quantify its reproducibility and redundancy. Features with ICC values greater than 0.75 were considered reproducible. To generate the number of non-redundancy feature subgroups, the R2 statistic method was used. Then, a classification model was built using a support vector machine (SVM) algorithm with 10-fold cross validation, and area under ROC curve (AUC) was used to evaluate the utility of non-redundant feature extraction by hierarchical clustering. RESULTS: The percentages of excellent reproducible features in the manual delineation group, GraphCut and GrowCut segmentation group were 69% [49], 73% [52] and 79% [56], respectively. Sixty-five percent [46] of the features showed strong robustness for all segmentation methods. The optimal number of cluster subgroup were 9, 13 and 11 for manual delineation, GraphCut and GrowCut segmentation, respectively. The optimal cluster subgroup number was 6 for all groups when the collectively high reproducibility features were selected for clustering. The receiver operating characteristic (ROC) analysis of radiomics classification model with and without feature reduction for healthy liver and HCC had an AUC value of 0.857 and 0.721 respectively. CONCLUSIONS: Our study demonstrates that variations exist in the reproducibility of quantitative imaging features extracted from tumor regions segmented using different methods. The reproducibility and non-redundancy of the radiomic features rely greatly on the tumor segmentation in HCC CT images. We recommend that the most reliable and uniform radiomic features should be selected in the clinical use of radiomics. Classification experiments with feature reduction showed that radiomic features were effective in identifying healthy liver and HCC. |
Real-Time Adaptive Planning Method for Radiotherapy Treatment Delivery for Prostate Cancer Patients, Based on a Library of Plans Accounting for Possible Anatomy Configuration Changes
|
Publication: PLoS One. 2019 Feb 28;14(2):e0213002. PMID: 30818345 | PDF Authors: Antico M, Prinsen P, Cellini F, Fracassi A, Isola AA, Cobben D, Fontanarosa D. Institution: School of Chemistry, Physics and Mechanical Engineering, Queensland University of Technology, Brisbane, Queensland, Australia. Abstract: BACKGROUND AND PURPOSE: In prostate cancer treatment with external beam radiation therapy (EBRT), prostate motion and internal changes in tissue distribution can lead to a decrease in plan quality. In most currently used planning methods, the uncertainties due to prostate motion are compensated by irradiating a larger treatment volume. However, this could cause underdosage of the treatment volume and overdosage of the organs at risk (OARs). To reduce this problem, in this proof of principle study we developed and evaluated a novel adaptive planning method. The strategy proposed corrects the dose delivered by each beam according to the actual position of the target in order to produce a final dose distribution dosimetrically as similar as possible to the prescribed one. MATERIAL AND METHODS: Our adaptive planning method was tested on a phantom case and on a clinical case. For the first, a pilot study was performed on an in-silico pelvic phantom. A "library" of intensity modulated RT (IMRT) plans corresponding to possible positions of the prostate during a treatment fraction was generated at planning stage. Then a 3D random walk model was used to simulate possible displacements of the prostate during the treatment fraction. At treatment stage, at the end of each beam, based on the current position of the target, the beam from the library of plans, which could reproduce the best approximation of the prescribed dose distribution, was selected and delivered. In the clinical case, the same approach was used on two prostate cancer patients: for the first a tissue deformation was simulated in-silico and for the second a cone beam CT (CBCT) taken during the treatment was used to simulate an intra-fraction change. Then, dosimetric comparisons with the standard treatment plan and, for the second patient, also with an isocenter shift correction, were performed. "...The CT was then elastically deformed on the CBCT using the B-spline method in 3D Slicer." RESULTS: For the phantom case, the plan generated using the adaptive planning method was able to meet all the dosimetric requirements and to correct for a misdosage of 13% of the dose prescription on the prostate. For the first clinical case, the standard planning method caused underdosage of the seminal vesicles, respectively by 5% and 4% of the prescribed dose, when the position changes for the target were correctly taken into account. The proposed adaptive planning method corrected any possible missed target coverage, reducing at the same time the dose on the OARs. For the second clinical case, both with the standard planning strategy and with the isocenter shift correction target coverage was significantly worsened (in particular uniformity) and some organs exceeded some toxicity objectives. While with our approach, the most uniform coverage for the target was produced and systematically the lowest toxicity values for the organs at risk were achieved. CONCLUSIONS: In our proof of principle study, the adaptive planning method performed better than the standard planning and the isocenter shift methods for prostate EBRT. It improved the coverage of the treatment volumes and lowered the dose to the OARs. This planning method is particularly promising for hypofractionated IMRT treatments in which a higher precision and control on dose deposition are needed. Further studies will be performed to test more extensively the proposed adaptive planning method and to evaluate it at a full clinical level. |
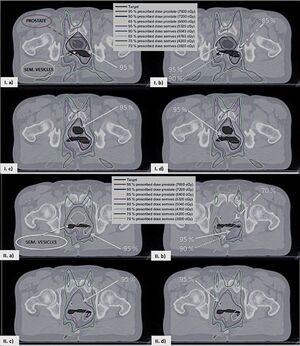 Isodose lines for two pelvic slices In the top and the bottom figure a, b, c and d correspond respectively to the PTV planning method without/with intra-fraction motion and the adaptive planning method with/without MU rescaling (both with intra-fraction motion). In the top figure in a and b the PTV is shown as a grey shaded area surrounding the prostate. The prostate is visible only in the slice shown in the top figure. |
Towards an Advanced Virtual Ultrasound-guided Renal Biopsy Trainer
|
Publication: Proc SPIE Int Soc Opt Eng. 2019 Feb;10951.. PMID: 31474785 | PDF Authors: Horvath S, Arikatla S, Cleary K, Sharma K, Rosenberg A, Enquobahrie A. Institution: Kitware, Inc., Clifton Park, NY, USA. Abstract: Ultrasound (US)-guided renal biopsy is a critically important tool in the evaluation and management of non-malignant renal pathologies with diagnostic and prognostic significance. It requires a good biopsy technique and skill to safely and consistently obtain high yield biopsy samples for tissue analysis. This project aims to develop a virtual trainer to help clinicians to improve procedural skill competence in real-time ultrasound-guided renal biopsy. This paper presents a cost-effective, high-fidelity trainer built using low-cost hardware components and open source visualization and interactive simulation libraries: interactive medical simulation toolkit (iMSTK) and 3D Slicer. We used a physical mannequin to simulate the tactile feedback that trainees experience while scanning a real patient and to provide trainees with spatial awareness of the US scanning plane with respect to the patient's anatomy. The ultrasound probe and biopsy needle were modeled using commonly used clinical tools and were instrumented to communicate with the simulator. 3D Slicer was used to visualize an image sliced from a pre-acquired 3-D ultrasound volume based on the location of the probe, with a realistic needle rendering. The simulation engine in iMSTK modeled the interaction between the needle and the virtual tissue to generate visual deformations on the tissue and tactile forces on the needle which are transmitted to the needle that the user holds. Initial testing has shown promising results with respect to quality of simulated images and system responsiveness. Further evaluation by clinicians is planned for the next stage. |
Methods for Quantitative Characterization of Bone Injury From Computed-Tomography Images
|
Publication: Proc SPIE Int Soc Opt Eng. 2019 Feb;10953. PMID: 31031512 | PDF Authors: Hernandez-Cerdan P, Paniagua B, Prothero J, Marron JS, Livingston E, Bateman T, McCormick M. Institution: Kitware, Inc., Carrboro, NC, USA. Abstract: Computed tomography (CT) images can potentially provide insights into bone structure for diagnosis of disorders and diseases. However, evaluation of trabecular bone structure and whole bone shape is often qualitative or semi-quantitative. This limits inter-study comparisons and the ability to detect subtle bone quality variations during early disease onset or in response to new treatments. In this work, we enable quantitative characterization of bone diseases through bone morphometry, texture analysis, and shape analysis methods. The potential of our analysis methods to identify the impact of hemophilia is validated in a mouse femur wound model. In our results, shape localizes and characterizes the formation of spurious bone, and our texture and bone morphometry analysis results provide extra information about the composition of that bone. Some of our one-dimensional (1D) textural features were able to significantly differentiate our injured femurs from our healthy femurs, even with this small sample size demonstrating the potential of the proposed analysis framework. While trabecular bone morphometrics have been a pillar in 3D microCT bone research for decades, the proposed analysis framework augments how we define and understand phenotypical presentation of bone disease. The contributed open source software is exposed to the medical image analysis community through 3D Slicer extensions to ensure both robustness and reproducibility. |
Predicting Breast Cancer Molecular Subtype with MRI Dataset Utilizing Convolutional Neural Network Algorithm
|
Publication: J Digit Imaging. 2019 Apr;32(2):276-82. PMID: 30706213 Authors: Ha R, Mutasa S, Karcich J, Gupta P, Pascual Van Sant E, Nemer J, Sun M, Chang P, Liu MZ, Jambawalikar S. Institution: Department of Radiology, Columbia University Medical Center, New York, NY, USA. Abstract: To develop a convolutional neural network (CNN) algorithm that can predict the molecular subtype of a breast cancer based on MRI features. An IRB-approved study was performed in 216 patients with available pre-treatment MRIs and immunohistochemical staining pathology data. First post-contrast MRI images were used for 3D segmentation using 3D Slicer. A CNN architecture was designed with 14 layers. Residual connections were used in the earlier layers to allow stabilization of gradients during backpropagation. Inception style layers were utilized deeper in the network to allow learned segregation of more complex feature mappings. Extensive regularization was utilized including dropout, L2, feature map dropout, and transition layers. The class imbalance was addressed by doubling the input of underrepresented classes and utilizing a class sensitive cost function. Parameters were tuned based on a 20% validation group. A class balanced holdout set of 40 patients was utilized as the testing set. Software code was written in Python using the TensorFlow module on a Linux workstation with one NVidia Titan X GPU. Seventy-four luminal A, 106 luminal B, 13 HER2+, and 23 basal breast tumors were evaluated. Testing set accuracy was measured at 70%. The class normalized macro area under receiver operating curve (ROC) was measured at 0.853. Non-normalized micro-aggregated AUC was measured at 0.871, representing improved discriminatory power for the highly represented Luminal A and Luminal B subtypes. Aggregate sensitivity and specificity was measured at 0.603 and 0.958. MRI analysis of breast cancers utilizing a novel CNN can predict the molecular subtype of breast cancers. Larger data sets will likely improve our model. |
Unique Metasomal Musculature in Sweat Bees (Hymenoptera, Apoidea, Halictidae) Revealed by Micro-CT Scanning
|
Publication: American Museum Novitates 2019 Feb; 3920:28 | PDF Authors: Herhold, Hollister W.; Davis, Steven R.; Smith, Corey Shepard.; Engel, Michael S.; Grimaldi, David A. Institution: American Museum of Natural History, New York, NY Abstract: Bees of the family Halictidae (Apoidea: Anthophila) have three pairs of thick, bundled muscles that are circular to subcircular in cross section within the first metasomal segment, as revealed by micro-CT scanning of 16 species in 15 genera of five bee families. In nonhalictids and the basal halictid subfamily Rophitinae, these muscles are planar (flat and sheetlike), typically lying between the anterior air sacs and abdominal wall. In Nomiinae and Halictinae, these muscles, especially the dorsal-ventral pair, bulge into air-sac space, partly enveloped by air-sac membrane. A possible function may be to facilitate metasomal compression and contraction, and thus air flow. The bundled shape of these derived halictid muscles is similar to that of flight muscles, but further data is needed to determine if they are fibrillar, which would suggest a completely different function. "Segmentation and volume rendering was done using 3D Slicer v4.9." |
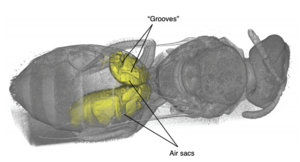 Volume rendering of Lasioglossum (Dialictus) sp. (Halictinae: Halictini), showing location of abdominal air sacs, displayed as a yellow solid, and “grooves” where metasomal muscles extend into the air-sac space. The insect’s exoskeleton and internal anatomy is rendered translucent, allowing examination of air-sac morphology. The bilateral asymmetry shown here is not uncommon, and usually has to do with how recently the specimen has fed. Distension of the gut can occupy space normally taken up by air sacs. |
Quantitative Features Can Predict Further Growth of Persistent Pure Ground-Glass Nodule
|
Publication: Quant Imaging Med Surg. 2019 Feb;9(2):283-91. PMID: 30976552 | PDF Authors: Shi Z, Deng J, She Y, Zhang L, Ren Y, Sun W, Su H, Dai C, Jiang G, Sun X, Xie D, Chen C. Institution: Department of Thoracic Surgery, Shanghai Pulmonary Hospital, Tongji University School of Medicine, Shanghai, China. Abstract: BACKGROUND: To evaluate whether quantitative features of persistent pure ground-glass nodules (PGGN) on the initial computed tomography (CT) scans can predict further nodule growth. METHODS: This retrospective study included 59 patients with 101 PGGNs from 2011 to 2012, who received regular CT follow-up for lung nodule surveillance. Nineteen quantitative image features consisting of 8 volumetric and 11 histogram parameters were calculated to detect lung nodule growth. For the extraction of the quantitative features, semi-automatic GrowCut segmentation was implemented on chest CT images in 3D Slicer platform. Univariate and multivariate analyses were performed to identify risk factors for nodule growth. RESULTS: With a median follow-up of 52 months, nodule growth was detected in 10 nodules by radiological assessment and in 16 nodules by quantitative features. In univariate analysis, 3D maximum diameter (MD), volume, mass, surface area, 90% percentile, and standard deviation value (SD) of PGGN on the initial CT scan were significantly different between stable nodules and nodules with further growth. In multivariate analysis, MD [hazard ratio (HR), 3.75; 95% confidence interval (CI), 2.14-6.55] and SD (HR, 2.06; 95% CI, 1.35-3.14) were independent predictors of further nodule growth. Also, the area under the curve was 0.896 (95% CI: 0.820-0.948) and 0.813 (95% CI: 0.723-0.883) for MD with a cut-off value of 10.2mm and SD of 50.0 Hounsfield Unit (HU). Besides, the growth rate was 55.6% (n=15) of PGGNs with MD >10.2 mm and SD >50.0 HU. CONCLUSIONS: Based on the initial CT scan, the quantitative features can predict PGGN growth more precisely. PGGN with MD >10.2 mm and SD >50.0 HU may require close follow-up or surgical intervention for the high incidence of growth. |
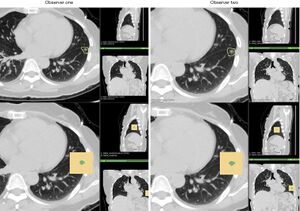 A representative image is displaying the process of nodule segmentation in 3D Slicer between two observers. |
3D Reconstruction of MR-Visible Fe3O4-Mesh Implants: Pelvic Mesh Measurement Techniques and Preliminary Findings
|
Publication: Neurourol Urodyn. 2019 Jan;38(1):369-78. PMID: 30387537 | PDF Authors: Brocker KA, Mokry T, Alt CD, Kauczor HU, Lenz F, Sohn C, DeLancey JO, Chen L. Institution: Department of Obstetrics and Gynecology, Medical School, University of Heidelberg, Heidelberg, Germany. Abstract: AIMS: To develop MR-based measurement technique to evaluate the postoperative dimension and location of implanted magnetic resonance (MR)-visible meshes. METHODS: This technique development study reports findings of six patients (A-F) with cystoceles treated with anterior vaginal MR-visible Fe3O4 -polypropylene implants. Implanted meshes were reconstructed from 3 months and/or 1 year postsurgical MR-images using 3D Slicer. Measurements including mesh length, distance to the ischial spines, pudendal, and obturator neurovascular bundles and urethra were obtained using software Rhino® and a custom Matlab® program. The range of implanted mesh length and their placements were reported and compared with mesh design and implantation recommendations. With the anterior/posterior-mesh-segment-ratio mesh shrinkage localization was evaluated. RESULTS: Examinations were possible for patients A-D 3 months and for A, C, E, and F 1 year postsurgical. The mesh was at least 40% shorter in all patients 3 months and/or 1 year postoperatively. A, B showed shrinkage in the anterior segment, D, E in the posterior segment (Patients C, F not applicable due to intraoperative mesh trimming). Patient E presented pain in the area of mesh shrinkage. In Patient C posterior mesh fixations were placed in the iliococcygeal muscle rather than sacrospinous ligaments. Arm placement less than 20 mm from the pudendal neurovascular bundles was seen in all cases. The portion of the urethra having mesh underneath it ranged from 19% to 55%. CONCLUSIONS: MRI-based measurement techniques have been developed to quantify implanted mesh location and dimension. Mesh placement variations possibly correlating with postoperative complications can be illustrated. Funding:
|
A Complete Workflow for Utilizing Monte Carlo Toolkits in Clinical Cases for a Double-Scattering Proton Therapy System
|
Publication: J Appl Clin Med Phys. 2019 Jan;20(1):23-30. PMID: 30426669 | PDF Authors: Muller L, Prusator M, Ahmad S, Chen Y. Institution: Department of Radiation Oncology, University of Oklahoma Health Sciences Center, Oklahoma City, OK. Abstract: The methods described in this paper allow end users to utilize Monte Carlo (MC) toolkits for patient-specific dose simulation and perform analysis and plan comparisons for double-scattering proton therapy systems. The authors aim to fill two aspects of this process previously not explicitly published. The first one addresses the modeling of field-specific components in simulation space. Patient-specific compensator and aperture models are exported from treatment planning system and converted to STL format using a combination of software tools including Matlab and Autodesk's Netfabb. They are then loaded into the MC geometry for simulation purpose. The second details a method for easily visualizing and comparing simulated doses with the dose calculated from the treatment planning system. This system is established by utilizing the open source software 3D Slicer. The methodology was demonstrated with a two-field proton treatment plan on the IROC lung phantom. Profiles and two-dimensional (2D) dose planes through the target isocenter were analyzed using our in-house software tools. This present workflow and set of codes can be easily adapted by other groups for their clinical practice. |
Accuracy Assessment of 3D-Printed Tooth Replicas
|
Publication: Int J Comput Dent. 2019;22(4):321-29. PMID: 31840140 Authors: Sokolowski AA, Kammerhofer J, Madreiter-Sokolowski CT, Payer M, Koller M, Jakse N, Wegscheider WA. Abstract: AIM: The production of individual tooth replicas has two applications in dental practice: tooth autotransplantations and dental root analogue implants. These applications require a particularly high degree of precision. The purpose of this study was to establish and evaluate a method for fabricating individual 3D-printed tooth replicas. MATERIALS AND METHODS: 10 patients requiring extraction of a wisdom tooth and a preoperative cone beam computed tomography (CBCT) scan were included; exclusion criteria were intraoperative fragmentation or fracture of the tooth. 3D Slicer v4.6.2 was used for tooth segmentation and model generation based on CBCT data. The tooth replicas were manufactured by selective laser melting (SLM). The extracted teeth and 3D-printed replicas were scanned and tested for surface deviations in CloudCompare 2.8.1. RESULTS: The mean absolute surface deviation between the 3D-printed teeth and the corresponding extracted teeth ranged from 0.13 to 0.25 mm, with standard deviations of 0.10 to 0.21 mm; 95% of the measured surface points deviated less than 0.474 mm; the surface area was reduced by -6.0% and the volume by -3.4%. The root mean square was 0.238 mm and the mean maximum absolute surface deviation was 0.927 mm. The SLM technique showed a high precision with a mean absolute deviation of 0.045 mm and a standard deviation of 0.04 mm. CONCLUSION: 3D-printed tooth replicas with a very high accuracy could be produced based on CBCT data. The described method is suitable for manufacturing tooth replicas for use in tooth autotransplantations or for fabricating root analogue implants. |
Glottic Configuration Changes and Outcomes of Endoscopic Arytenoid Abduction Lateropexy
|
Publication: Eur Arch Otorhinolaryngol. 2019 Jan;276(1):167-173. PMID: 30483943 Authors: Szakács L, Sztanó B, Matievics V, Bere Z, Castellanos PF, Rovó L. Institution: Department of Otorhinolaryngology-Head and Neck Surgery, University of Szeged, Szeged, Hungary. Abstract: Endoscopic arytenoid abduction lateropexy (EAAL) is an effective glottis enlarging procedure for the treatment of bilateral vocal cord palsy (BVCP). The postoperative glottic configuration changes can be evaluated by modern, high-resolution, 3D image reconstructions. Functional results are described by spirometry as well as objective and subjective phoniatric tests. METHODS: Unilateral EAAL was performed in ten malignant thyroid gland tumor patients (eight women, two men), who had BVCP after thyroid surgery. 3D Slicer software was used for morphometric analysis. Pre- and postoperative peak inspiratory flow (PIF) and standard phoniatric parameters were compared. RESULTS: The glottic gap improved significantly (+ 60%). Significant improvement of PIF was found in all cases. Phoniatric tests revealed better quality of voice and patient satisfaction. Their voices changed from a severely impaired to a socially acceptable, almost normal, quality. CONCLUSION: The results support our clinical observations that the ideal position of the lateralization sutures is the one which provides a physiological abduction position of the arytenoid cartilage. Considering these good results, the surgical indications for minimally invasive endoscopic arytenoid lateropexy may be extended. |
Pyramidal Neuron Growth and Increased Hippocampal Volume During Labor and Birth in Autism
|
Publication: Sci Adv. 2019 Jan 23;5(1):eaav0394. PMID: 30746473 | PDF Authors: Cloarec R, Riffault B, Dufour A, Rabiei H, Gouty-Colomer LA, Dumon C, Guimond D, Bonifazi P, Eftekhari S, Lozovaya N, Ferrari DC, Ben-Ari Y. Institution: Neurochlore, Ben-Ari Institute of Neuroarcheology (IBEN), Zone Luminy Biotech Entreprises, 13288 Cedex 09 , Marseille, France. Abstract: We report that the apical dendrites of CA3 hippocampal pyramidal neurons are increased during labor and birth in the valproate model of autism but not in control animals. Using the iDISCO clearing method, we show that hippocampal, especially CA3 region, and neocortical volumes are increased and that the cerebral volume distribution shifts from normal to lognormal in valproate-treated animals. Maternal administration during labor and birth of the NKCC1 chloride transporter antagonist bumetanide, which reduces [Cl-]i levels and attenuates the severity of autism, abolished the neocortical and hippocampal volume changes and reduced the whole-brain volume in valproate-treated animals. These results suggest that the abolition of the oxytocin-mediated excitatory-to-inhibitory shift of GABA actions during labor and birth contributes to the pathogenesis of autism spectrum disorders by stimulating growth during a vulnerable period. "...Image registration to compute average shapes of the whole brain, hippocampus, and neocortex was performed using a python script, MeshLab software (www.meshlab.net/), and the SPHARM module of 3D Slicer software. " |
Opposing CSF Hydrodynamic Trends Found in the Cerebral Aqueduct and Prepontine Cistern Following Shunt Treatment in Patients With Normal Pressure Hydrocephalus
|
Publication: Fluids Barriers CNS. 2019 Jan 22;16(1):2. PMID: 30665428 | PDF Authors: Hamilton RB, Scalzo F, Baldwin K, Dorn A, Vespa P, Hu X, Bergsneider M. Institution: Neural Systems and Dynamics Laboratory, Department of Neurosurgery, The David Geffen School of Medicine, University of California-Los Angeles, Los Angeles, CA, USA. Abstract: BACKGROUND: This study investigated cerebrospinal fluid (CSF) hydrodynamics using cine phase-contrast MRI in the cerebral aqueduct and the prepontine cistern between three distinct groups: pre-shunt normal pressure hydrocephalus (NPH) patients, post-shunt NPH patients, and controls. We hypothesized that the hyperdynamic flow of CSF through the cerebral aqueduct seen in NPH patients was due to a reduction in cisternal CSF volume buffering. Both hydrodynamic (velocity, flow, stroke volume) and peak flow latency (PFL) parameters were investigated. METHODS: Scans were conducted on 30 pre-treatment patients ranging in age from 58 to 88 years along with an additional 12 controls. Twelve patients also received scans following either ventriculoatrial (VA) or ventriculoperitoneal (VP) shunt treatment (9 VP, 3 VA), ranging in age from 74 to 89 years with a mean follow up time of 6 months. RESULTS: Significant differences in area, velocity, flow, and stroke volume for the cerebral aqueduct were found between the pre-treatment NPH group and the healthy controls. Shunting caused a significant decrease in both caudal and cranial mean flow and stroke volume in the cerebral aqueduct. No significant changes were found in the prepontine cistern between the pre-treatment group and healthy controls. For the PFL, no significant differences were seen in the cerebral aqueduct between any of the three groups; however, the prepontine cistern PFL was significantly decreased in the pre-treatment NPH group when compared to the control group. CONCLUSIONS: Although several studies have quantified the changes in aqueductal flow between hydrocephalic groups and controls, few studies have investigated prepontine cistern flow. Our study was the first to investigate both regions in the same patients for NPH pre- and post- treatment. Following shunt treatment, the aqueductal CSF metrics decreased toward control values, while the prepontine cistern metrics trended up (not significantly) from the normal values established in this study. The opposing trend of the two locations suggests a redistribution of CSF pulsatility in NPH patients. Furthermore, the significantly decreased latency of the prepontine cisternal CSF flow suggests additional evidence for CSF pulsatility dysfunction. "...For the nine patients that had pre- and post- treatment scans, the total lateral and third ventricle volumes were calculated (3D Slicer). " |
Computer Simulations Suggest That Prostate Enlargement Due to Benign Prostatic Hyperplasia Mechanically Impedes Prostate Cancer Growth
|
Publication: Proc Natl Acad Sci U S A. 2019 Jan 22;116(4):1152-61. PMID: 30617074 | PDF Authors: Lorenzo G, Hughes TJR, Dominguez-Frojan P, Reali A, Gomez H. Institution: Departamento de Matemáticas, Universidade da Coruña, Spain. Abstract: Prostate cancer and benign prostatic hyperplasia are common genitourinary diseases in aging men. Both pathologies may coexist and share numerous similarities, which have suggested several connections or some interplay between them. However, solid evidence confirming their existence is lacking. Recent studies on extensive series of prostatectomy specimens have shown that tumors originating in larger prostates present favorable pathological features. Hence, large prostates may exert a protective effect against prostate cancer. In this work, we propose a mechanical explanation for this phenomenon. The mechanical stress fields that originate as tumors enlarge have been shown to slow down their dynamics. Benign prostatic hyperplasia contributes to these mechanical stress fields, hence further restraining prostate cancer growth. We derived a tissue-scale, patient-specific mechanically coupled mathematical model to qualitatively investigate the mechanical interaction of prostate cancer and benign prostatic hyperplasia. This model was calibrated by studying the deformation caused by each disease independently. Our simulations show that a history of benign prostatic hyperplasia creates mechanical stress fields in the prostate that impede prostatic tumor growth and limit its invasiveness. The technology presented herein may assist physicians in the clinical management of benign prostate hyperplasia and prostate cancer by predicting pathological outcomes on a tissue-scale, patient-specific basis. "...used 3D Slicer to generate a triangular surface model of the prostate from the contours of the organ and the urethra drawn on the T2-weighted MR images, using the provided prostate segmentation as guidance." |
Machine Learning Derived Segmentation of Phase Velocity Encoded Cardiovascular Magnetic Resonance for Fully Automated Aortic Flow Quantification
|
Publication: J Cardiovasc Magn Reson. 2019 Jan 7;21(1):1. PMID: 30612574 | PDF Authors: Bratt A, Kim J, Pollie M, Beecy AN, Tehrani NH, Codella N, Perez-Johnston R, Palumbo MC, Alakbarli J, Colizza W, Drexler IR, Azevedo CF, Kim RJ, Devereux RB, Weinsaft JW. Institution: Department of Radiology, Weill Cornell Medicine, New York, NY, USA. Abstract: BACKGROUND: Phase contrast (PC) cardiovascular magnetic resonance (CMR) is widely employed for flow quantification, but analysis typically requires time consuming manual segmentation which can require human correction. Advances in machine learning have markedly improved automated processing, but have yet to be applied to PC-CMR. This study tested a novel machine learning model for fully automated analysis of PC-CMR aortic flow. METHODS: A machine learning model was designed to track aortic valve borders based on neural network approaches. The model was trained in a derivation cohort encompassing 150 patients who underwent clinical PC-CMR then compared to manual and commercially-available automated segmentation in a prospective validation cohort. Further validation testing was performed in an external cohort acquired from a different site/CMR vendor. RESULTS: Among 190 coronary artery disease patients prospectively undergoing CMR on commercial scanners (84% 1.5T, 16% 3T), machine learning segmentation was uniformly successful, requiring no human intervention: Segmentation time was < 0.01 min/case (1.2 min for entire dataset); manual segmentation required 3.96 ± 0.36 min/case (12.5 h for entire dataset). Correlations between machine learning and manual segmentation-derived flow approached unity (r = 0.99, p < 0.001). Machine learning yielded smaller absolute differences with manual segmentation than did commercial automation (1.85 ± 1.80 vs. 3.33 ± 3.18 mL, p < 0.01): Nearly all (98%) of cases differed by ≤5 mL between machine learning and manual methods. Among patients without advanced mitral regurgitation, machine learning correlated well (r = 0.63, p < 0.001) and yielded small differences with cine-CMR stroke volume (∆ 1.3 ± 17.7 mL, p = 0.36). Among advanced mitral regurgitation patients, machine learning yielded lower stroke volume than did volumetric cine-CMR (∆ 12.6 ± 20.9 mL, p = 0.005), further supporting validity of this method. Among the external validation cohort (n = 80) acquired using a different CMR vendor, the algorithm yielded equivalently small differences (∆ 1.39 ± 1.77 mL, p = 0.4) and high correlations (r = 0.99, p < 0.001) with manual segmentation, including similar results in 20 patients with bicuspid or stenotic aortic valve pathology (∆ 1.71 ± 2.25 mL, p = 0.25). CONCLUSION: Fully automated machine learning PC-CMR segmentation performs robustly for aortic flow quantification - yielding rapid segmentation, small differences with manual segmentation, and identification of differential forward/left ventricular volumetric stroke volume in context of concomitant mitral regurgitation. Findings support use of machine learning for analysis of large scale CMR datasets. "...labeling pixels in the magnitude images as either valve or non-valve using 3D Slicer, an open-source medical image post-processing application." |
Morphological Analysis of Sigmoid Sinus Anatomy: Clinical Applications to Neurotological Surgery
|
Publication: J Otolaryngol Head Neck Surg. 2019 Jan 11;48(1):2. PMID: 30635049 | PDF Authors: Van Osch K, Allen D, Gare B, Hudson TJ, Ladak H, Agrawal SK. Institution: Schulich School of Medicine & Dentistry, Western University, London, Ontario, Canada. Abstract: OBJECTIVES: The primary objective of this study was to use high-resolution micro-CT images to create accurate three-dimensional (3D) models of several intratemporal structures, and to compare several surgically important dimensions within the temporal bone. The secondary objective was to create a statistical shape model (SSM) of a dominant and non-dominant sigmoid sinus (SS) to provide a template for automated segmentation algorithms. METHODS: A free image processing software, 3D Slicer, was utilized to create three-dimensional reconstructions of the SS, jugular bulb (JB), facial nerve (FN), and external auditory canal (EAC) from micro-CT scans. The models were used to compare several clinically important dimensions between the dominant and non-dominant SS. Anatomic variability of the SS was also analyzed using SSMs generated using the Statismo software framework. RESULTS: Three-dimensional models from 38 temporal bones were generated and analyzed. Right dominance was observed in 74% of the paired SSs. All distances were significantly shorter on the dominant side (p < 0.05), including: EAC - SS (dominant: 13.7 ± 3.4 mm; non-dominant: 15.3 ± 2.7 mm), FN - SS (dominant: 7.2 ± 1.8 mm; non-dominant: 8.1 ± 2.3 mm), 2nd genu FN - superior tip of JB (dominant: 8.7 ± 2.2 mm; non-dominant: 11.2 ± 2.6 mm), horizontal distance between the superior tip of JB - descending FN (dominant: 9.5 ± 2.3 mm; non-dominant: 13.2 ± 3.5 mm), and horizontal distance between the FN at the stylomastoid foramen - JB (dominant: 5.4 ± 2.2 mm; non-dominant: 7.7 ± 2.1). Analysis of the SSMs indicated that SS morphology is most variable at its junction with the transverse sinus, and least variable at the JB. CONCLUSIONS: This is the first known study to investigate the anatomical variation and relationships of the SS using high resolution scans, 3D models and statistical shape analysis. This analysis seeks to guide neurotological surgical approaches and provide a template for automated segmentation and surgical simulation. "In 3D Slicer, nine fiducials (F1 – F9) were placed on the 3D reconstructions of the SS, JB, EAC, and FN to analyze several surgically relevant relationships between these structures." Funding:
|
Biomarker Localization, Analysis, Visualization, Extraction, and Registration (BLAzER) Methodology for Research and Clinical Brain PET Applications
|
Publication: J Alzheimers Dis. 2019;70(4):1241-57. PMID: 31322571 | PDF Authors: Raman F, Grandhi S, Murchison CF, Kennedy RE, Landau S, Roberson ED, McConathy J; Alzheimer’s Disease Neuroimaging Initiative. Institution: Department of Radiology, University of Alabama at Birmingham, Birmingham, AL, USA. Abstract: BACKGROUND: Tools for efficient evaluation of amyloid- and tau-PET images are needed in both clinical and research settings. OBJECTIVE: This study was designed to validate a semi-automated image analysis methodology, called Biomarker Localization, Analysis, Visualization, Extraction, and Registration (BLAzER). We tested BLAzER using two different segmentation platforms, FreeSurfer (FS) and Neuroreader (NR), for regional brain PET quantification in participants in the Alzheimer's Disease Neuroimaging Initiative (ADNI) dataset. METHODS: 127 amyloid-PET and 55 tau-PET studies with volumetric MRIs were obtained from ADNI. The BLAzER methodology utilizes segmentation of MR images by FS or NR, then visualizes and quantifies regional brain PET data using FDA-cleared software (MIM), enabling quality control to ensure optimal registration and to detect segmentation errors. RESULTS: BLAzER analysis required ∼5 min plus segmentation time. BLAzER using FS segmentation showed strong agreement with ADNI for global amyloid-PET standardized uptake value ratios (SUVRs) (r = 0.9922, p < 0.001) and regional tau-PET SUVRs across all Braak staging regions (r > 0.97, p < 0.001) with high inter-operator reproducibility (ICC > 0.97) and nearly identical dichotomization as amyloid-positive or -negative (2 discrepant cases out of 127). Comparing FS versus NR segmentation with BLAzER, global SUVRs were strongly correlated for amyloid-PET (r = 0.9841, p < 0.001), but were systematically higher (4% on average) with NR, likely due to more inclusion of white matter with NR-defined regions. CONCLUSIONS: BLAzER provides an efficient methodology for regional brain PET quantification. FDA-cleared components and visualization of registration reduce barriers between research and clinical applications. "...Upon completion of segmentation, parcellated, segmented brain images were converted from MGZ to DICOM format using 3D Slicer v4.5.0." Funding:
|
Imaging as Part of a Quality Assurance Program: Predictors of Interobserver Variability for Pretreatment Image Registration for Lung SBRT
|
Publication: Technol Cancer Res Treat. 2019; 18: PMC6732858 | PDF Authors: Geoff Baran, Jay Burmeister, Peter Paximadis, Todd Bossenberg, Robert Halford, Kathryn Masi, Adrian Nalichowski, Steven Miller, Nitin Vaishampayan, Mark Zaki, Kristen Iannotti, Melanie Komajda, Kathryn Mattews, Essam Qasim, Sean Sullivan, Hassan Beydoun, Michael Dominello. Institution: Department of Radiation Oncology, Karmanos Cancer Institute, Detroit, MI, USA. Abstract: Purpose: To evaluate the magnitude of interobserver variability in pretreatment image registration for lung stereotactic body radiation therapy patients in aggregate and within 3 clinical subgroups and to determine methods to identify patients expected to demonstrate larger variability. Methods and Materials: Retrospective image registration was performed for the first and last treatment fraction for 10 lung stereotactic body radiation therapy patients by 16 individual observers (5 physicians, 6 physicists, and 5 therapists). Registration translation values were compared within and between subgroups overall and between the first and the last fractions. Four metrics were evaluated as possible predictors for large interobserver variability. Results: The mean 3-dimensional displacement vector for all patients over all comparisons was 2.4 ± 1.8 mm. Three patients had mean 3-dimensional vector differences >3 mm. This cohort of patients showed a significant interfraction difference in variance (P value = .01), increasing from first fraction to last. A significant difference in interobserver variability was observed between physicians and physicists (P value < .01) and therapists and physicists (P value < .01) but not between physicians and therapists (P value = .07). Three of the 4 quantities evaluated as potential predictive metrics showed statistical correlation with increased interobserver variation, including target excursion and local target/lung contrast. Conclusion: Variability in pretreatment image guidance represents an important treatment consideration, particularly for stereotactic body radiation therapy, which employs small margins and a small number of treatment fractions. As a result of the data presented here, we have initiated weekly “registration rounds” to familiarize all staff physicians with the target and normal anatomy for each stereotactic body radiation therapy patient and minimize interobserver variations in image registration prior to treatment. The metrics shown here are capable of identifying patients for which large interobserver variations would be anticipated. These metrics may be used in the future to develop thresholds for additional interventions to mitigate registration variations. "...The GTV0 contour and PTV* contour were exported to 3D Slicer along with the 4DCT image associated with GTV0." |
Go to 2020 :: 2019 :: 2018 :: 2017 :: 2016 :: 2015 :: 2014-2011:: 2010-2005