Documentation/Nightly/Modules/TrainModel
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
|
Extension: LesionSegmentation | |||
|
Module Description
This module is used to train new segmentation models for white matter lesion segmentation. In order to use this tool your data must include a T1, T2, FLAIR, brain mask, and expert lesion segmentation for each subject. All data must be preprocessed including intra-subject co-registration, AC-PC alignment, bias correction, consistent spacing between sequences, and brain mask creation.
Use Cases
- Use Case: Segmenting white matter lesions in a disorder for which there is no current lesion segmentation model
If a lesion segmentation model already exists for the disorder of interest you most likely do not need to train a new one. However, if a model does not exist or the existing model(s) give poor performance on your data you can use this module to train a new model, and then use that model to automatically segment white matter lesions.
In order to train a new model you must first have preprocessed data on a number of subjects. The data required includes T1, T2, FLAIR, brain masks, and lesion masks. All data must be preprocessed including intra-subject co-registration, AC-PC alignment, bias correction, consistent spacing between sequences (i.e all 1mm isotropic), and brain mask creation. Subjects do not need to be registered to each other. A model can be created on a single subject, but greater than 6 is recommended and between 10 and 15 is best. The more subjets included in the model the slower model creation will be and the slower segmentation using that model will be. However, models using more subjects will almost always be more accurate. When selecting subjects to use for the model try to pick the largest variety that you can. This means, if appropriate for your dataset, multiple ages, multiple brain sizes, multiple lesion loads, a balance between genders, multiple stages of disease progression, etc. Then identify the set of images for a single subject that has the cleanest and highest contrast scans and use that subject's scans as the first set of inputs. They will be used as the reference for intensity standardization. You will need to use those images as the reference scans when predicting as well.
Tutorials
Coming soon!
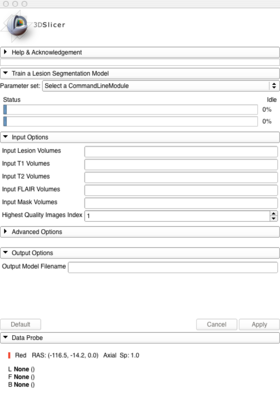
Panels and their use
A list panels in the interface, their features, what they mean, and how to use them.
|
Similar Modules
References
- Scully M, Anderson B, Lane T, Gasparovic C, Magnotta V, Sibbitt W, Roldan C, Kikinis R and Bockholt HJ (2010) An automated method for segmenting white matter lesions through multi-level morphometric feature classification with application to lupus. Front. Hum. Neurosci. doi:10.3389/fnhum.2010.00027
http://frontiersin.org/neuroscience/humanneuroscience/paper/10.3389/fnhum.2010.00027/
Information for Developers
| Section under construction. |