Documentation/Nightly/Extensions/BabyBrain
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
|
This work was partially funded by CAPES and CNPq, a Brazillian Agencies. Information on CAPES can be obtained on the CAPES website and CNPq website. | |||||||||
|
Extension Description
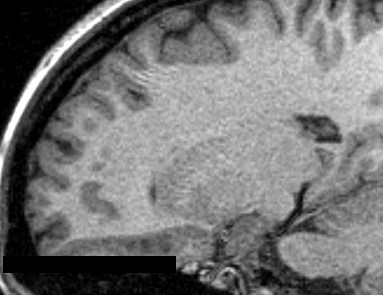
This module offers a set of algorithms to biomedical image data preparation, which is focused on the neonate and fetal MRI analysis. The methods used are optimized to structural MRI images, namely T2 and T1. Further improvements are under development stage in order to address new imaging modalities (e.g. diffusion-weighted images). The general procedure assumes that the MRI image was already reconstructed in a volume representation and also brain extracted.
Basically, there are two different modules: BabyBrainPreparation and BabyBrainSegmentation, which each of them are optimized for a specific set of image processing. More details are elucidated below.
Modules
- General image enhancement procedure: Baby Brain Preparation
- Brain tissue segmentation: Baby Brain Segmentation
Use Cases
Baby Brain Preparation:
- Use Case 1: Image preparation to further tissue segmentation
- The structural MRI images (fetal or neonate) often have a strong level of image artefacts (e.g. noise and grey level inhomogeneity), hence a previous data preparation is needed for further segmentation process.
- Use Case 2: Volume rendering
- Image preprocessing frequently increases the quality of volume renderings
Baby Brain Segmentation:
- Use Case 1: Brain morphological information
- The measurement of morphological features in brain images (e.g. volume of white or grey matter, brain atrophy, etc) are commonly used in several clinical studies.
Similar Extensions
References
- PAPER
Information for Developers
| Section under construction. |
Repositories:
- Source code: GitHub repository
- Issue tracker: open issues and enhancement requests