Difference between revisions of "Slicer3.4:Training"
From Slicer Wiki
| (93 intermediate revisions by 9 users not shown) | |||
| Line 1: | Line 1: | ||
| − | = | + | =Slicer 3.4 Tutorials= |
| − | * | + | * For tutorials for other versions of Slicer, please visit the [[Training| Slicer training portal]]. |
| + | * The following table contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment (version 3.4 release) to accomplish certain tasks. | ||
| + | * For questions related to the Slicer3.4 Compendium, please send an e-mail to Sonia Pujol, Ph.D. (spujol at bwh.harvard.edu). | ||
| − | |||
| − | + | {| border="1" cellpadding="5" width="1000px" | |
| − | + | |- style="background:#FFD700; color:#4A2A0A; font-size:130%" align="center" | |
| − | = | + | | '''Category''' |
| − | + | | '''Tutorial''' | |
| − | + | | '''Sample Data''' | |
| − | + | | '''Image''' | |
| − | + | |- | |
| − | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"|<span id="1.1"></span> '''Basic''' | |
| − | + | | style="background:#FDFC6D; color:black" valign="top"| [[Media:Slicer3Minute_SPujol.pdf | '''Slicer3Minute Tutorial''']] <br><br> The Slicer3Minute tutorial is an introduction to the advanced 3D visualization capabilities of Slicer3.4.<br> <br>'''Audience:''' First time users. | |
| − | + | | style="background:#FDFC6D; color:black" valign="top"| [[Media:Slicer3MinuteDataset.zip | '''Slicer3Minute Data''']]<br><br> The Slicer3Minute dataset contains an MR scan of the brain and 3D reconstructions of the anatomy | |
| − | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:Slicer3Minute.PNG|250px]] | |
| − | | | + | |- |
| − | | style=" | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"|<span id="1.1"></span> '''Basic''' |
| − | | style=" | + | | style="background:#FDFC6D; color:black" valign="top"| [[Media:Slicer3_Tutorial_ManualRegistration.pdf| '''Slicer3 Manual Registration Tutorial''']] <br><br> shows how to manually/interactively align two images in Slicer3.4 or 3.5<br> <br>'''Audience:''' First time & early users. |
| − | | style=" | + | | style="background:#FDFC6D; color:black" valign="top"| [[Media:Slicer3_Tutorial_ManualRegistration_ExampleDataset.zip | '''Manual Registration Data''']]<br><br> This dataset contains two brain MRI of a single subject, obtained in different orientations. |
| − | | style=" | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:Slicer3_ManualRegistrationTutorial.gif|250px]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"|<span id="1.2"></span> '''Core''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top"| [[Media:Slicer3Course_DataLoading_3DVisualization_SoniaPujol.pdf | '''Slicer3Visualization Tutorial ''']] <br><br> The Slicer3Visualization tutorial guides through 3D data loading and visualization in Slicer3.4.<br> <br>'''Audience:''' All beginning users including clinicians, scientists, engineers and programmers. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top"| [[Media:Slicer3VisualizationDataset.zip | '''Slicer3Visualization Data''']] <br><br> The Slicer3VisualizationDataset contains two MR scans of the brain, a pre-computed labelmap and 3D reconstructions of the anatomy. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:DataLoadingAndVisualization_SPujol.png|250px]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Core''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:ProgrammingIntoSlicer3_SoniaPujol.pdf | '''Programming in Slicer3 Tutorial''']] <br><br> The Programming in Slicer3 tutorial is an introduction to the the integration of stand-alone programs outside of the Slicer3 source tree.<br> <br>'''Audience:''' Programmers and Engineers. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:HelloWorld_Plugin.zip| '''HelloWorld Plugin''']]<br><br> The HelloWorld tutorial dataset contains an MR scan of the brain and pre-computed xml and C++ files for integrating the Hello World plug-in to Slicer3. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:ProgrammingCourse_Logo.PNG|250px|Programming]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:FreeSurferCourse_Slicer3-4_SoniaPujol.pdf | '''3D Visualization of FreeSurfer Data''']] <br><br> The course guides through 3D visualization of FreeSurfer brain segmentations, surface reconstruction and parcellation results in Slicer3.4.<br> <br>'''Audience:''' All users. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [http://www.na-mic.org/Wiki/index.php/File:FreeSurferTutorialData.zip '''FreeSurfer Tutorial Data''']<br><br> The FreeSurfer dataset contains an MR scan of the brain and pre-computed FreeSurfer segmentation and cortical surface reconstructions. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:FreeSurferTutorial.PNG|250px]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:AutomaticSegmentation_SoniaPujol.pdf | '''Automatic Segmentation Tutorial''']] <br><br> The course guides through the process of using the Expectation-Maximization Segmentation algorithm to automatically segment brain structures from MRI data.<br> <br>'''Audience:''' Programmers and Engineers. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:AutomaticSegmentation.zip| '''Automatic Segmentation Data''']] |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:EMSegmentationTutorial.PNG|250px]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:Slicer3_Tutorial_AtlasMerging_RegLib_C11-mj.ppt | '''Atlas Label Merging & Surface Based Registration Tutorial ''']] <br><br> This tutorial guides through the creation and co-registration of surface models of atlas structures (thalamus & thalamic nuclei) and subsequent merging of two labelmaps. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:Slicer3_Tutorial_AtlasMerging_DataSet.zip| '''Atlas Merging Data''' (contains 2 full brain atlases + intermediate result files, 72MB)]] |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:Slicer3_SurfaceRegistrationTutorial_RegLib_C11.gif|250px]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top"| [http://wiki.na-mic.org/Wiki/index.php/IGT:ToolKit/Neurosurgical-Planning '''Neurosurgical Planning Tutorial'''] <br><br> This tutorial takes the trainee through a complete workup for neurosurgical patient-specific mapping. Also see this tutorial for information on how to use Slicer's affine registration, simple region growing, model maker and tractography modules.<br> <br>'''Audience:''' All users interested in image-guided therapy. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:NeurosurgicalPlanningTutorialData.zip| '''Neurosurgical Planning Data ''']] |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:NeurosurgicalPlanningOverview.png|250px]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:DiffusionMRITutorial_SFN2009_SPujol.pdf| '''Diffusion MRI Tutorial''']] <br><br> This tutorial guides you through the process of loading diffusion MR data, estimating diffusion tensors, and performing tractography of white matter bundles.<br> <br>'''Audience:''' All users and developers. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:DiffusionDataset.zip | '''Diffusion Data''']] |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" align="center"| [[Image:cc.PNG |250px]] |
|- | |- | ||
| − | | style="background:#FFD700; color: | + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:Slicer3.4.1-ChangeTrackerTutorial.pdf | '''ChangeTracker Tutorial''']] <br><br> This tutorial describes the use of ChangeTracker module to detect changes in tumor volume from two MRI scans. |
| − | | style="background:# | + | '''Audience:''' All users interested in longitudinal analysis of pathology. |
| − | | style="background:# | + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:Slicer3.4.1-ChangeTrackerTutorial.pdf | '''Training Data download is integrated with the ChangeTracker module (see Tutorial)''']] |
| + | | style="background:#FDFC6D; color:black" align="center"| [[Image:Slicer3.4.1-ChangeTracker.jpg |250px]] | ||
|- | |- | ||
| + | | style="background:#FFD700; color:#4A2A0A; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' | ||
| + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:3DVisualizationLiverSegments_SoniaPujol_RSNA2009.pdf | '''LiverSegmentation Tutorial''']] <br><br> This tutorial introduces translational clinical scientists to the capabilities of the 3D Slicer software through the segmentation of the liver. | ||
| + | '''Audience:''' All users and developers. | ||
| + | | style="background:#FDFC6D; color:black" valign="top" | [[Media:LiverData_RSNA2009.tar.gz | '''Liver Segmentation Data''']] | ||
| + | | style="background:#FDFC6D; color:black" align="center"| [[Image:LiverSegmentation.png |250px]] | ||
|} | |} | ||
= Slicer Tutorial Contest = | = Slicer Tutorial Contest = | ||
| − | ''' | + | The following tutorials were part of the [http://www.na-mic.org/Wiki/index.php/AHM2010:Tutorial_Contest January 2010 Slicer tutorial contest], [http://www.na-mic.org/Wiki/index.php/Events:TutorialContestJune2009 June 2009 Slicer tutorial contest] and [http://www.na-mic.org/Wiki/index.php/Events:TutorialContestJan2009 January 2009 Slicer tutorial contest]. |
| + | |||
| + | == '''January 2010 Slicer Tutorial Contest'''== | ||
| − | + | {| border="1" cellpadding="5" width="1000px" | |
| + | |- style="background:#849f66; color:#3b3f33; font-size:130%" align="center" | ||
| + | |'''DBPs/Collaboration''' | ||
| + | |'''Tutorial''' | ||
| + | |'''Sample Data''' | ||
| + | |'''Image''' | ||
| + | |- | ||
| + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Fiber Clustering''' | ||
| + | | style="background:#CCFF99; color:black" valign="top" | [http://wiki.na-mic.org/Wiki/images/c/c7/FiberClusteringTrainingTutorial_Winter2010AHM.pdf '''EM Fiber Clustering'''] | ||
| + | | style="background:#CCFF99; color:black" valign="top" | [http://www.nitrc.org/projects/quantitativedti/ EM Fiber Clustering Data ] | ||
| + | | style="background:#CCFF99; color:black" align="center"| [[Image:Shot2.png |250px]] | ||
| + | |- | ||
| + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''VMTK''' | ||
| + | | style="background:#CCFF99; color:black" valign="top" | [http://wiki.na-mic.org/Wiki/images/4/40/TutorialVMTKCoronariesCenterlinesMRI_Winter2010AHM.pdf '''VMTK Coronary Arteries centerlines Extraction'''] | ||
| + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/index.php/File:TutorialVMTKCoronariesCenterlinesMRI_Data_Winter2010AHM.zip VMTK Centrelines Data] | ||
| + | | style="background:#CCFF99; color:black" align="center"| [[Image:Vmtkcloseupvoronoicenterlinewithreference.png|250px]] | ||
| + | |} | ||
| + | == '''June 2009 Slicer Tutorial Contest'''== | ||
| − | {| border="1" cellpadding="5" | + | {| border="1" cellpadding="5" width="1000px" |
| − | |- style="background:# | + | |- style="background:#849f66; color:#3b3f33; font-size:130%" align="center" |
| − | | style=" | + | |'''DBPs/Collaboration''' |
| − | | style=" | + | |'''Tutorial''' |
| − | | style=" | + | |'''Sample Data''' |
| − | | style=" | + | |'''Image''' |
| + | |- | ||
| + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Confocal Microscopy''' | ||
| + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/images/4/4d/Microscopy-Confocal-TrainingTutorial-2009JUNE.pdf '''Confocal Microscopy'''] | ||
| + | | style="background:#CCFF99; color:black" valign="top" | [http://www.ccdb.ucsd.edu/index.shtm Microscopy Confocal Data] | ||
| + | | style="background:#CCFF99; color:black" align="center"| [[Image:MicroscopyTutorial.png| 250px]] | ||
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Autism''' |
| − | | style="background:#CCFF99; color:black" | | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/images/3/33/ARCTIC-Slicer3-Tutorial.pdf '''ARCTIC: Automatic Regional Cortical ThICkness'''] |
| − | | style="background:#CCFF99; color:black" | [http://www. | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.nitrc.org/projects/arctic/ ARCTIC Data] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image:Corticalthickness.png| 250px ]] |
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Prostate''' |
| − | | style="background:#CCFF99; color:black" | | + | | style="background:#CCFF99; color:black" valign="top"| [http://www.na-mic.org/Wiki/images/0/06/DBP2JohnsHopkinsTransRectalProstateBiopsy_TutorialPres2009June.pdf '''Trans-rectal MR guided prostate biopsy'''] |
| − | | style="background:#CCFF99; color:black" | [ | + | | style="background:#CCFF99; color:black" valign="top" | [[Media:TransRectalProstateBiopsyTutorialDataset.zip| MR-guided Prostate Biopsy Data]] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image:TransRectalBiopsy.png| 250px]] |
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Schizophrenia''' |
| − | | style="background:#CCFF99; color:black" | ''' | + | | style="background:#CCFF99; color:black" valign="top" | [[media:Stochastic_June09_3.ppt| '''Python Stochastic Tractography Module''']] |
| − | | style="background:#CCFF99; color:black" | [ | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/index.php/File:IJdata.tar.gz Stochastic Tractography Data] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image:StochasticTractographyTutorial2.png|250px]] |
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Lupus''' |
| − | | style="background:#CCFF99; color:black" | | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/images/7/7c/Slicer3Training_WhiteMatterLesions_v2.2.1.pdf '''White Matter Lesions Segmentation'''] |
| − | | style="background:#CCFF99; color:black" | [http://www.na-mic.org/Wiki/index.php/File: | + | | style="background:#CCFF99; color:black" valign="top"| [http://www.na-mic.org/Wiki/index.php/File:LesionSegmentationTutorialData.zip Lesion Segmentation Tutorial Data] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image:LupusTutorial2.PNG| 250px]] |
|- | |- | ||
| − | | | + | |} |
| − | | style="background:# | + | |
| − | | | + | == '''January 2009 Slicer Tutorial Contest'''== |
| − | | | + | |
| + | {| border="1" cellpadding="5" width="1000px" | ||
| + | |- style="background:#849f66; color:#3b3f33; font-size:130%" align="center" | ||
| + | |'''DBPs/Collaboration''' | ||
| + | |'''Tutorial''' | ||
| + | |'''Sample Data''' | ||
| + | |'''Image''' | ||
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Automated FE Mesh''' |
| − | | style="background:#CCFF99; color:black" | | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/images/b/b8/IA-FEMesh-Tutorial-Notes.pdf '''IA FEMesh Tutorial'''] |
| − | | style="background:#CCFF99; color:black" | [http://www.na-mic.org/Wiki/index.php/File:MeshTutorialExampleData.zip Mesh Tutorial Example Data] | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/index.php/File:MeshTutorialExampleData.zip Mesh Tutorial Example Data] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image:Femesh-in-trunk-120808.png|250px]] |
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Alcohol and Stress Interaction''' |
| − | | style="background:#CCFF99; color:black" | | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/images/8/83/EMSegment_TrainingTutorial.pdf '''Non-Human Primate Segmentation Tutorial'''] |
| − | | style="background:#CCFF99; color:black" | [ | + | | style="background:#CCFF99; color:black" valign="top" | [[media:SlicerTutorialData_Vervet.tar.gz | Non-Human Primate Segmentation Data]] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image: Non-HumanPrimateSegmentation.png|250px]] |
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Prostate''' |
| − | | style="background:#CCFF99; color:black" | | + | | style="background:#CCFF99; color:black" valign="top"| [http://wiki.na-mic.org/Wiki/index.php/IGT:ToolKit/Prostate-Planning '''MR-guided Prostate Interventions'''] |
| − | | style="background:#CCFF99; color:black" | [http://www.na-mic.org/Wiki/index.php/File:MRGuidedProstateInterventions.zip MR Guided Prostate Interventions Data] | + | | style="background:#CCFF99; color:black" valign="top"| [http://www.na-mic.org/Wiki/index.php/File:MRGuidedProstateInterventions.zip MR Guided Prostate Interventions Data] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"|[[Image:ProstatePlanningOverview.png | 250px]] |
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''IGT''' |
| − | | style="background:#CCFF99; color:black" | | + | | style="background:#CCFF99; color:black" valign="top" | [http://www.na-mic.org/Wiki/index.php/IGT:ToolKit/Navigation-tutorial '''Basic Navigation Tutorial'''] |
| − | | style="background:#CCFF99; color:black" | [http://www.na-mic.org/publications/item/view/1265 Basic Navigation Data] | + | | style="background:#CCFF99; color:black" valign="top"| [http://www.na-mic.org/publications/item/view/1265 Basic Navigation Data] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image:300px-IGTBasicNavigationSummary.png| 250px]] |
|- | |- | ||
| − | | style="background:# | + | | style="background:#849f66; color:#3b3f33; font-size:110%" align="center"| <span id="1.1"></span> '''Orbit Segmentation''' |
| − | | style="background:#CCFF99; color:black" | ''' | + | | style="background:#CCFF99; color:black" valign="top"| [[Media:OrbitSegmentationTutorial.pdf| '''Segmentation of the Orbit''']] |
| − | | style="background:#CCFF99; color:black" | [ | + | | style="background:#CCFF99; color:black" valign="top"| [[Media:OrbiteSegmentationData.zip | Orbit Segmentation Data ]] |
| − | | style="background:#CCFF99; color:black" align=" | + | | style="background:#CCFF99; color:black" align="center"| [[Image:OrbitSegmentationt.png |250px]] |
|- | |- | ||
| − | | style="background:# | + | |} |
| − | + | ||
| − | | | + | = Slicer 3.5 Tutorials= |
| − | | | + | |
| + | *The following table contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment (version 3.5) to accomplish certain tasks. | ||
| + | |||
| + | |||
| + | |||
| + | {| border="1" cellpadding="5" width="1000px" | ||
| + | |- style="background:#547389; color:#994139; font-size:130%" align="center" | ||
| + | |'''Contest''' | ||
| + | |'''Tutorial''' | ||
| + | |'''Sample Data''' | ||
| + | |'''Image''' | ||
|- | |- | ||
| − | | style="background:# | + | | style="background:#547389; color:#994139; font-size:110%" align="center"| <span id="1.1"></span> '''Specialized''' |
| − | | style="background:# | + | | style="background:#b7cfd7; color:black" valign="top"| [http://www.na-mic.org/Wiki/index.php/RobustStatisticsSegmentation '''Robust Statistics Segmentation Tutorial'''] |
| − | | style="background:# | + | | style="background:#b7cfd7; color:black" valign="top" | [[Media:Patient1.tar.gz | '''Robust Statistics Segmentation Data''']] |
| − | | style="background:# | + | | style="background:#b7cfd7; color:black" align="center"| [[Image:RobustStatisticsSegmentation LiverSpleenKidney.png |250px]] |
|- | |- | ||
| − | | | + | |} |
| − | + | ||
| − | + | =Software Installation= | |
| − | + | *The [http://www.slicer.org/pages/Special:SlicerDownloads '''Slicer download page'''] contains information on how to obtain a compiled version of Slicer for a variety of platforms and where to find the source code for Slicer 3. | |
| − | + | ||
| + | =Software Documentation= | ||
| + | *For the Slicer 3.4 manual pages please click [[Documentation-3.4|here]]. These pages are the reference manual for Slicer 3.4 and briefly explain the functionality found in panels and modules. | ||
| + | |||
| + | =Older Tutorials= | ||
| + | *Visit the [http://wiki.na-mic.org/Wiki/index.php/Slicer3.2:Training Slicer 3.2 training page]. | ||
| + | *Visit the [http://wiki.na-mic.org/Wiki/index.php/Slicer:Workshops:User_Training_101 Slicer 2 training page]. | ||
Latest revision as of 17:28, 30 January 2012
Home < Slicer3.4:TrainingContents
Slicer 3.4 Tutorials
- For tutorials for other versions of Slicer, please visit the Slicer training portal.
- The following table contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment (version 3.4 release) to accomplish certain tasks.
- For questions related to the Slicer3.4 Compendium, please send an e-mail to Sonia Pujol, Ph.D. (spujol at bwh.harvard.edu).
| Category | Tutorial | Sample Data | Image |
| Basic | Slicer3Minute Tutorial The Slicer3Minute tutorial is an introduction to the advanced 3D visualization capabilities of Slicer3.4. Audience: First time users. |
Slicer3Minute Data The Slicer3Minute dataset contains an MR scan of the brain and 3D reconstructions of the anatomy |
|
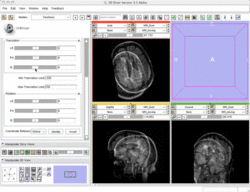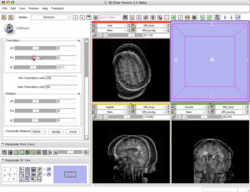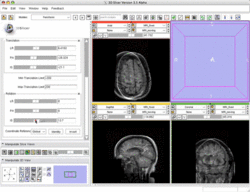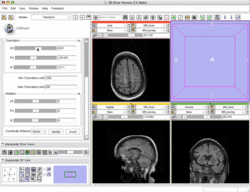
| Basic | Slicer3 Manual Registration Tutorial shows how to manually/interactively align two images in Slicer3.4 or 3.5 Audience: First time & early users. |
Manual Registration Data This dataset contains two brain MRI of a single subject, obtained in different orientations. |
|
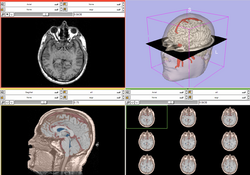
| Core | Slicer3Visualization Tutorial The Slicer3Visualization tutorial guides through 3D data loading and visualization in Slicer3.4. Audience: All beginning users including clinicians, scientists, engineers and programmers. |
Slicer3Visualization Data The Slicer3VisualizationDataset contains two MR scans of the brain, a pre-computed labelmap and 3D reconstructions of the anatomy. |
|
| Core | Programming in Slicer3 Tutorial The Programming in Slicer3 tutorial is an introduction to the the integration of stand-alone programs outside of the Slicer3 source tree. Audience: Programmers and Engineers. |
HelloWorld Plugin The HelloWorld tutorial dataset contains an MR scan of the brain and pre-computed xml and C++ files for integrating the Hello World plug-in to Slicer3. |
|
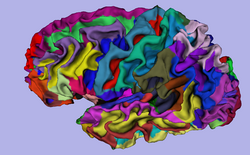
| Specialized | 3D Visualization of FreeSurfer Data The course guides through 3D visualization of FreeSurfer brain segmentations, surface reconstruction and parcellation results in Slicer3.4. Audience: All users. |
FreeSurfer Tutorial Data The FreeSurfer dataset contains an MR scan of the brain and pre-computed FreeSurfer segmentation and cortical surface reconstructions. |
|
| Specialized | Automatic Segmentation Tutorial The course guides through the process of using the Expectation-Maximization Segmentation algorithm to automatically segment brain structures from MRI data. Audience: Programmers and Engineers. |
Automatic Segmentation Data | 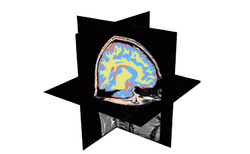
|
| Specialized | Atlas Label Merging & Surface Based Registration Tutorial This tutorial guides through the creation and co-registration of surface models of atlas structures (thalamus & thalamic nuclei) and subsequent merging of two labelmaps. |
Atlas Merging Data (contains 2 full brain atlases + intermediate result files, 72MB) | 
|
| Specialized | Neurosurgical Planning Tutorial This tutorial takes the trainee through a complete workup for neurosurgical patient-specific mapping. Also see this tutorial for information on how to use Slicer's affine registration, simple region growing, model maker and tractography modules. Audience: All users interested in image-guided therapy. |
Neurosurgical Planning Data | 
|
| Specialized | Diffusion MRI Tutorial This tutorial guides you through the process of loading diffusion MR data, estimating diffusion tensors, and performing tractography of white matter bundles. Audience: All users and developers. |
Diffusion Data | 
|
| Specialized | ChangeTracker Tutorial This tutorial describes the use of ChangeTracker module to detect changes in tumor volume from two MRI scans. Audience: All users interested in longitudinal analysis of pathology. |
Training Data download is integrated with the ChangeTracker module (see Tutorial) | |
| Specialized | LiverSegmentation Tutorial This tutorial introduces translational clinical scientists to the capabilities of the 3D Slicer software through the segmentation of the liver. Audience: All users and developers. |
Liver Segmentation Data | 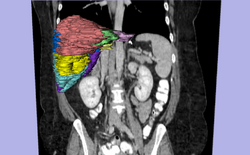
|
Slicer Tutorial Contest
The following tutorials were part of the January 2010 Slicer tutorial contest, June 2009 Slicer tutorial contest and January 2009 Slicer tutorial contest.
January 2010 Slicer Tutorial Contest
| DBPs/Collaboration | Tutorial | Sample Data | Image |
| Fiber Clustering | EM Fiber Clustering | EM Fiber Clustering Data | 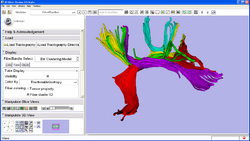
|
| VMTK | VMTK Coronary Arteries centerlines Extraction | VMTK Centrelines Data | 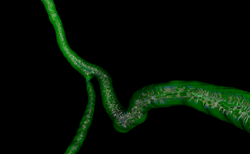
|
June 2009 Slicer Tutorial Contest
| DBPs/Collaboration | Tutorial | Sample Data | Image |
| Confocal Microscopy | Confocal Microscopy | Microscopy Confocal Data | 
|
| Autism | ARCTIC: Automatic Regional Cortical ThICkness | ARCTIC Data | 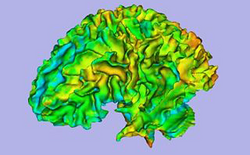
|
| Prostate | Trans-rectal MR guided prostate biopsy | MR-guided Prostate Biopsy Data | 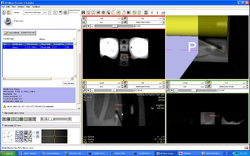
|
| Schizophrenia | Python Stochastic Tractography Module | Stochastic Tractography Data | 
|
| Lupus | White Matter Lesions Segmentation | Lesion Segmentation Tutorial Data | 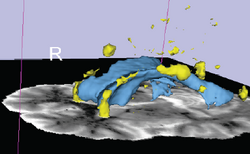
|
January 2009 Slicer Tutorial Contest
| DBPs/Collaboration | Tutorial | Sample Data | Image |
| Automated FE Mesh | IA FEMesh Tutorial | Mesh Tutorial Example Data | 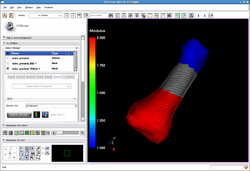
|
| Alcohol and Stress Interaction | Non-Human Primate Segmentation Tutorial | Non-Human Primate Segmentation Data | 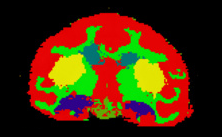
|
| Prostate | MR-guided Prostate Interventions | MR Guided Prostate Interventions Data | 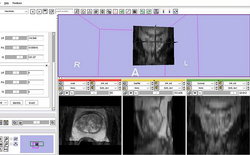
|
| IGT | Basic Navigation Tutorial | Basic Navigation Data | 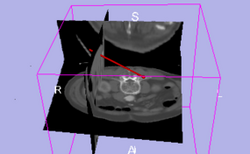
|
| Orbit Segmentation | Segmentation of the Orbit | Orbit Segmentation Data | 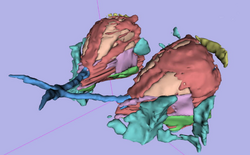
|
Slicer 3.5 Tutorials
- The following table contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment (version 3.5) to accomplish certain tasks.
| Contest | Tutorial | Sample Data | Image |
| Specialized | Robust Statistics Segmentation Tutorial | Robust Statistics Segmentation Data | 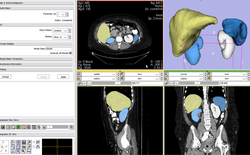
|
Software Installation
- The Slicer download page contains information on how to obtain a compiled version of Slicer for a variety of platforms and where to find the source code for Slicer 3.
Software Documentation
- For the Slicer 3.4 manual pages please click here. These pages are the reference manual for Slicer 3.4 and briefly explain the functionality found in panels and modules.
Older Tutorials
- Visit the Slicer 3.2 training page.
- Visit the Slicer 2 training page.