Difference between revisions of "Documentation/4.3/Extensions/SlicerRT"
(Nightly -> 4.3) |
m |
||
| (5 intermediate revisions by 2 users not shown) | |||
| Line 12: | Line 12: | ||
Contributors: Greg Sharp (Massachusetts General Hospital), Steve Pieper (Isomics)<br> | Contributors: Greg Sharp (Massachusetts General Hospital), Steve Pieper (Isomics)<br> | ||
Contacts: | Contacts: | ||
| + | * Csaba Pinter, <email>csaba.pinter@queensu.ca</email> | ||
* Andras Lasso, <email>lasso@cs.queensu.ca</email> | * Andras Lasso, <email>lasso@cs.queensu.ca</email> | ||
* Mailing list, <email>slicerrt@alerts.assembla.com</email> (need to register to [http://www.assembla.com Assembla] using the email address from which the email is sent) | * Mailing list, <email>slicerrt@alerts.assembla.com</email> (need to register to [http://www.assembla.com Assembla] using the email address from which the email is sent) | ||
| − | * [ | + | * [[Documentation/SlicerRT/HowToReportAnError|How to report an error]] |
Website: [http://slicerrt.github.io slicerrt.org]<br> | Website: [http://slicerrt.github.io slicerrt.org]<br> | ||
License: [http://www.slicer.org/pages/LicenseText Slicer license]<br> | License: [http://www.slicer.org/pages/LicenseText Slicer license]<br> | ||
| − | '''Download/install:''' install 3D Slicer, start 3D Slicer, open the Extension Manager, install the SlicerRT extension (see more details [ | + | '''Download/install:''' install 3D Slicer, start 3D Slicer, open the Extension Manager, install the SlicerRT extension (see more details [http://slicerrt.github.io/Download.html on the download page]) |
<!-- | <!-- | ||
| Line 31: | Line 32: | ||
[[Image:SlicerRT_Logo_2.0_128x128.png]] | [[Image:SlicerRT_Logo_2.0_128x128.png]] | ||
| | | | ||
| − | SlicerRT is one of the themes of the SparKit (Software Platform and Adaptive Radiotherapy Kit) project with the goal of making 3D Slicer a powerful radiotherapy research platform. | + | * SlicerRT is one of the themes of the SparKit (Software Platform and Adaptive Radiotherapy Kit) project with the goal of making 3D Slicer a powerful radiotherapy research platform.<br>SparKit is a project funded by Cancer Care Ontarioand the Ontario Consortium for Adaptive Interventions in Radiation Oncology (OCAIRO) to provide free, open-source toolset for radiotherapy and related image-guided interventions. |
| − | SparKit is a project funded by Cancer Care Ontarioand the Ontario Consortium for Adaptive Interventions in Radiation Oncology (OCAIRO) to provide free, open-source toolset for radiotherapy and related image-guided interventions. | ||
| − | The SlicerRT extension incorporates [[Documentation/{{documentation/version}}/Extensions/Plastimatch|Plastimatch]] modules and algorithms. | + | * The SlicerRT extension incorporates [[Documentation/{{documentation/version}}/Extensions/Plastimatch|Plastimatch]] modules and algorithms. |
| + | |||
| + | * Additional information for users can be found on the [[Documentation/SlicerRT/UsersGuide|User's guide]] page <br> | ||
| | | | ||
|} | |} | ||
| Line 45: | Line 47: | ||
*[[Documentation/{{documentation/version}}/Modules/DicomRtImport|DICOM-RT import]] | *[[Documentation/{{documentation/version}}/Modules/DicomRtImport|DICOM-RT import]] | ||
| + | *[[Documentation/{{documentation/version}}/Modules/DicomRtExport|DICOM-RT export]] | ||
*[[Documentation/{{documentation/version}}/Modules/Contours|Contours]] | *[[Documentation/{{documentation/version}}/Modules/Contours|Contours]] | ||
*[[Documentation/{{documentation/version}}/Modules/DoseVolumeHistogram|Dose volume histogram]] | *[[Documentation/{{documentation/version}}/Modules/DoseVolumeHistogram|Dose volume histogram]] | ||
| Line 52: | Line 55: | ||
*[[Documentation/{{documentation/version}}/Modules/ContourComparison|Contour comparison]] | *[[Documentation/{{documentation/version}}/Modules/ContourComparison|Contour comparison]] | ||
*[[Documentation/{{documentation/version}}/Modules/ContourMorphology|Contour morphology]] | *[[Documentation/{{documentation/version}}/Modules/ContourMorphology|Contour morphology]] | ||
| − | |||
| − | |||
* Modules from [[Documentation/{{documentation/version}}/Extensions/Plastimatch|Plastimatch]] (Greg Sharp) | * Modules from [[Documentation/{{documentation/version}}/Extensions/Plastimatch|Plastimatch]] (Greg Sharp) | ||
**[[Documentation/{{documentation/version}}/Modules/PlmBSplineDeformableRegistration|Plastimatch Automatic deformable image registration]] (Greg Sharp) | **[[Documentation/{{documentation/version}}/Modules/PlmBSplineDeformableRegistration|Plastimatch Automatic deformable image registration]] (Greg Sharp) | ||
| − | |||
| − | |||
**[[Documentation/{{documentation/version}}/Modules/PlmLANDWARP|Plastimatch LANDWARP Landmark]] (Greg Sharp) | **[[Documentation/{{documentation/version}}/Modules/PlmLANDWARP|Plastimatch LANDWARP Landmark]] (Greg Sharp) | ||
**[[Documentation/{{documentation/version}}/Modules/PlmXFORMWARP|Plastimatch XFORMWARP]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | **[[Documentation/{{documentation/version}}/Modules/PlmXFORMWARP|Plastimatch XFORMWARP]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | ||
<!-- | <!-- | ||
| + | **[[Documentation/{{documentation/version}}/Modules/PlmDICOMRTImport|Plastimatch DICOM-RT import]] (Greg Sharp) | ||
| + | **[[Documentation/{{documentation/version}}/Modules/PlmDICOMRTExport|Plastimatch DICOM-RT export]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | ||
**[[Documentation/{{documentation/version}}/Modules/PlmDoseComparison|Plastimatch Dose Comparison]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | **[[Documentation/{{documentation/version}}/Modules/PlmDoseComparison|Plastimatch Dose Comparison]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | ||
**[[Documentation/{{documentation/version}}/Modules/PlmDoseVolumeHistogram|Plastimatch Dose Volume Histogram]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | **[[Documentation/{{documentation/version}}/Modules/PlmDoseVolumeHistogram|Plastimatch Dose Volume Histogram]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | ||
| Line 67: | Line 68: | ||
**[[Documentation/{{documentation/version}}/Modules/PlmThreshBox|Plastimatch Threshold in a box]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | **[[Documentation/{{documentation/version}}/Modules/PlmThreshBox|Plastimatch Threshold in a box]] (Greg Sharp)[[image:UnderConstruction.png|tumb|10px]] | ||
--> | --> | ||
| + | |||
| + | * Former SlicerRT modules integrated to Slicer core | ||
| + | **[[Documentation/{{documentation/version}}/Modules/SubjectHierarchy|Subject hierarchy]] | ||
| + | **[[Documentation/{{documentation/version}}/Modules/DeformationFieldVisualizer|Transform visualizer]] | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
| Line 73: | Line 78: | ||
* Evaluation of the effect of image-based non-rigid patient motion compensation | * Evaluation of the effect of image-based non-rigid patient motion compensation | ||
* Dose accumulation with motion compensation | * Dose accumulation with motion compensation | ||
| + | * Evaluation of gel dosimetry | ||
| + | * Testing of treatment planning algorithms | ||
| − | | [[File: | + | | [[File:SlicerRt_Montage.jpg|1024px|SlicerRT highlights]] |
|} | |} | ||
| Line 85: | Line 92: | ||
** Workshop material: [http://www.na-mic.org/Wiki/index.php/File:Pinter_MedUni2013_Workshop.pdf download] from NA-MIC.org | ** Workshop material: [http://www.na-mic.org/Wiki/index.php/File:Pinter_MedUni2013_Workshop.pdf download] from NA-MIC.org | ||
* RSNA 2012 tutorial | * RSNA 2012 tutorial | ||
| − | ** Tutorial description: [https://www.assembla.com/spaces/slicerrt/wiki/20121127_Slicer_tutorials_at_RSNA_2012 | + | ** Tutorial description: [http://www.donotlink.com/bEo SlicerRT wiki: Slicer tutorials at RSNA 2012] |
| + | <!-- Original link: https://www.assembla.com/spaces/slicerrt/wiki/20121127_Slicer_tutorials_at_RSNA_2012 --> | ||
** Sample data: [http://slicer.kitware.com/midas3/folder/859 download] SlicerRT ART dose verification data from Midas server | ** Sample data: [http://slicer.kitware.com/midas3/folder/859 download] SlicerRT ART dose verification data from Midas server | ||
* Summer NA-MIC week 2012 tutorial | * Summer NA-MIC week 2012 tutorial | ||
| − | ** Tutorial presentation: [https://www.assembla.com/spaces/slicerrt/documents/aAuMq81Eqr4ztvacwqjQXA/download/aAuMq81Eqr4ztvacwqjQXA download] from the SlicerRT website | + | ** Tutorial presentation: [http://www.donotlink.com/bEp download] from the SlicerRT website |
| − | + | <!-- Original link: https://www.assembla.com/spaces/slicerrt/documents/aAuMq81Eqr4ztvacwqjQXA/download/aAuMq81Eqr4ztvacwqjQXA --> | |
| + | ** Sample data: [http://www.donotlink.com/bEq download] from the SlicerRT website | ||
| + | <!-- Original link: https://www.assembla.com/spaces/slicerrt/documents/bMuwgYTKur4yP-acwqjQWU/download/bMuwgYTKur4yP-acwqjQWU --> | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/extension-section|Similar Extensions}} | {{documentation/{{documentation/version}}/extension-section|Similar Extensions}} | ||
* [[Documentation/{{documentation/version}}/Extensions/Plastimatch|Plastimatch]]: SlicerRT and Plastimatch are complementary software libraries. Plastimatch focuses on delivering new computational methods radiotherapy, while SlicerRT aims for providing an easy-to-use interface for a wide range of stable, well-tested radiotherapy related features. SlicerRT uses Plastimatch internally for certain operations. | * [[Documentation/{{documentation/version}}/Extensions/Plastimatch|Plastimatch]]: SlicerRT and Plastimatch are complementary software libraries. Plastimatch focuses on delivering new computational methods radiotherapy, while SlicerRT aims for providing an easy-to-use interface for a wide range of stable, well-tested radiotherapy related features. SlicerRT uses Plastimatch internally for certain operations. | ||
| + | * [[Documentation/{{documentation/version}}/Modules/GelDosimetry|Gel Dosimetry]]: Slicelet facilitating a streamlined workflow to perform gel dosimetry analysis for commissioning linacs and evaluating new dose calculation procedures | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/extension-section|References}} | {{documentation/{{documentation/version}}/extension-section|References}} | ||
| − | + | ==How to cite== | |
| + | Please cite the following paper when referring to SlicerRt in your publication:<br> | ||
| + | C. Pinter, A. Lasso, A. Wang, D. Jaffray and G. Fichtinger, [http://perk.cs.queensu.ca/sites/perk.cs.queensu.ca/files/Pinter2012_0.pdf "SlicerRT – Radiation therapy research toolkit for 3D Slicer"], Med. Phys., 39(10) pp. 6332-6338, 2012 | ||
| + | <pre> | ||
| + | @ARTICLE{Pinter2012, | ||
| + | author = {Pinter, C. and Lasso, A. and Wang, A. and Jaffray, D. and Fichtinger, G.}, | ||
| + | title = {SlicerRT – Radiation therapy research toolkit for 3D Slicer}, | ||
| + | journal = {Med. Phys.}, | ||
| + | year = {2012}, | ||
| + | volume = {39}, | ||
| + | number = {10}, | ||
| + | pages = {6332-6338}, | ||
| + | } | ||
| + | </pre> | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/extension-section|Information for Developers}} | {{documentation/{{documentation/version}}/extension-section|Information for Developers}} | ||
| + | * [http://www.donotlink.com/bEr SlicerRT developers wiki page] | ||
| + | <!-- Original link: https://www.assembla.com/spaces/slicerrt/wiki/SlicerRt_developers_page --> | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/extension-footer}} | {{documentation/{{documentation/version}}/extension-footer}} | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
Latest revision as of 22:04, 23 September 2014
Home < Documentation < 4.3 < Extensions < SlicerRT
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
Authors: Csaba Pinter (PerkLab, Queen's University), Andras Lasso (PerkLab, Queen's University), Kevin Wang (Radiation Medicine Program, Princess Margaret Hospital, University Health Network Toronto)
Contributors: Greg Sharp (Massachusetts General Hospital), Steve Pieper (Isomics)
Contacts:
- Csaba Pinter, <email>csaba.pinter@queensu.ca</email>
- Andras Lasso, <email>lasso@cs.queensu.ca</email>
- Mailing list, <email>slicerrt@alerts.assembla.com</email> (need to register to Assembla using the email address from which the email is sent)
- How to report an error
Website: slicerrt.org
License: Slicer license
Download/install: install 3D Slicer, start 3D Slicer, open the Extension Manager, install the SlicerRT extension (see more details on the download page)
Extension Description
|
Modules
Use Cases
|
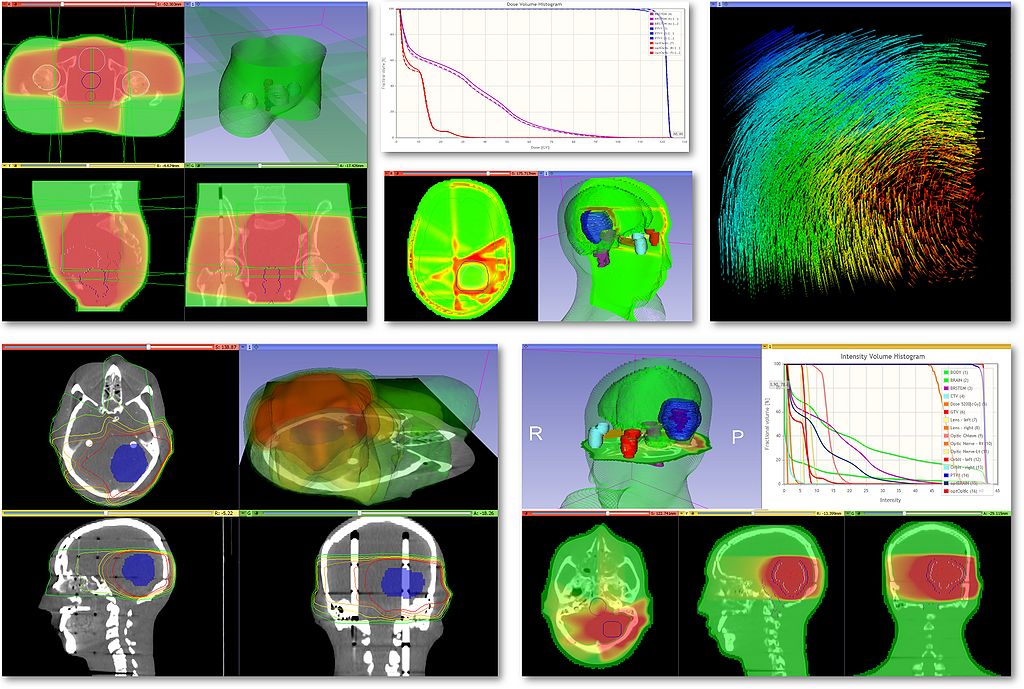
|
Tutorials
- Summer NA-MIC week 2013 tutorial (recommended)
- ECR 2013 - Medical University Vienna workshop
- Workshop material: download from NA-MIC.org
- RSNA 2012 tutorial
- Tutorial description: SlicerRT wiki: Slicer tutorials at RSNA 2012
- Sample data: download SlicerRT ART dose verification data from Midas server
- Summer NA-MIC week 2012 tutorial
Similar Extensions
- Plastimatch: SlicerRT and Plastimatch are complementary software libraries. Plastimatch focuses on delivering new computational methods radiotherapy, while SlicerRT aims for providing an easy-to-use interface for a wide range of stable, well-tested radiotherapy related features. SlicerRT uses Plastimatch internally for certain operations.
- Gel Dosimetry: Slicelet facilitating a streamlined workflow to perform gel dosimetry analysis for commissioning linacs and evaluating new dose calculation procedures
References
How to cite
Please cite the following paper when referring to SlicerRt in your publication:
C. Pinter, A. Lasso, A. Wang, D. Jaffray and G. Fichtinger, "SlicerRT – Radiation therapy research toolkit for 3D Slicer", Med. Phys., 39(10) pp. 6332-6338, 2012
@ARTICLE{Pinter2012,
author = {Pinter, C. and Lasso, A. and Wang, A. and Jaffray, D. and Fichtinger, G.},
title = {SlicerRT – Radiation therapy research toolkit for 3D Slicer},
journal = {Med. Phys.},
year = {2012},
volume = {39},
number = {10},
pages = {6332-6338},
}



