Modules:PlastimatchLANDWARP
Return to Slicer 3.6 Documentation
Plastimatch > Landmark-based Registartion
General Information
Module Type & Category
Type: CLI
Category: Plastimatch
Authors, Collaborators & Contact
- Authors: See COPYRIGHT.TXT contained within the package
- Contact: Nadya Shusharina, Department of Radiation Oncology, Massachusetts General Hospital (nshsuharina@partners.org)
- Web page: http://plastimatch.org
Module Description
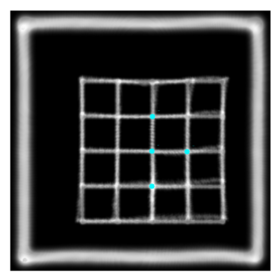
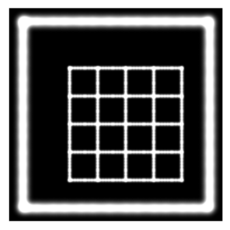
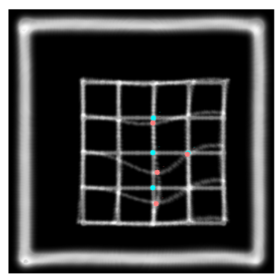
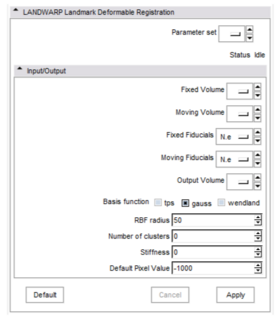
This is the plastimatch landmark-based deformable image registration module. The intended application of this method is rapid, interactive correction of registration failures with a small number of mouse clicks. Compared to other landmark-based methods, the plastimatch registration method might offer:
- both local and global registration
- regularization of the deformation field
Examples of how this module is being used:
- Intra-subject registration for adaptive radiotherapy
- Inter-subject registration for automatic segmentation
Usage
Tutorials
Quick Tour of Features and Use
|
Development
Notes from the Developer(s)
Developer-oriented documentation is found on the plastimatch web site: http://plastimatch.org
Dependencies
This module has no dependencies.
Tests
Plastimatch features approximately 100 test cases.
Known bugs
Usability issues
Please report usability issues to the bug tracker.
Source code & documentation
We recommended to download the latest source code from subversion:
Documentation:
More Information
About plastimatch
Plastimatch is an open source software for deformable image registration. It is designed for high-performance volumetric registration of medical images, such as X-ray computed tomography (CT), magnetic resonance imaging (MRI), and positron emission tomography (PET). Software features include:
- B-spline method for deformable image registration (GPU and multicore accelerated)
- Demons method for deformable image registration (GPU accelerated)
- ITK-based algorithms for translation, rigid, affine, demons, and B-spline registration
- Pipelined, multi-stage registration framework with seamless conversion between most algorithms and transform types
- Landmark-based deformable registration using thin-plate splines for global registration
- Landmark-based deformable registration using radial basis functions for local corrections
- Broad support for 3D image file formats (using ITK), including Dicom, Nifti, NRRD, MetaImage, and Analyze
- Dicom and DicomRT import and export
- XiO import and export
- Plugins for 3D Slicer
Plastimatch also features two handy utilities which are not directly related to image registration:
- FDK cone-beam CT reconstruction (GPU and multicore accelerated)
- Digitally reconstructed radiograph (DRR) generation (GPU and multicore accelerated)
Acknowledgment
National Institutes of Health
NIH / NCI 6-PO1 CA 21239
Federal share of program income earned by MGH on C06CA059267
Progetto Rocca Foundation
A collaboration between MIT and Politecnico di Milano
References
- G Sharp et al. "Plastimatch - An open source software suite for radiotherapy image processing," Proceedings of the XVIth International Conference on the use of Computers in Radiotherapy, May, 2010.
- N. Shusharina, G. Sharp "Landmark-based image registration with analytic regularization", IEEE Trans. Med. Imag., submitted, 2011.