Modules:FastMarchingSegmentation-Documentation-3.6
Return to Slicer 3.6 Documentation
Fast Marching Segmentation
FastMarchingSegmentation
General Information
Module Type & Category
Type: Interactive
Category: Segmentation
Authors, Collaborators & Contact
- Andriy Fedorov, Ron Kikinis (BWH)
- Eric Pichon
- Contact: Andriy Fedorov, fedorov at bwh
Module Description
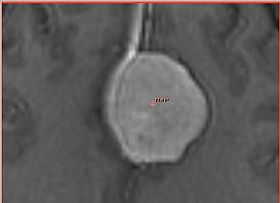
This module implements active contour segmentation based on fast marching method.
Usage
Use Cases, Examples
- This module is most useful to segment image regions that have similar intensity
- Initialization is very simple: points within the region to be segmented and expected volume of the segmented structure
- The segmentation is completed relatively quickly for typical images, facilitating experimentation with the selection of optimum parameters
- The resulting volume can be adjusted interactively by scrolling through the evolution of the label contour
Tutorials
Not available at this time
Quick Tour of Features and Use
|
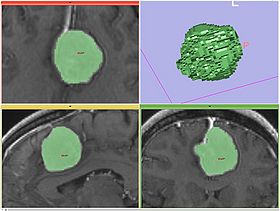
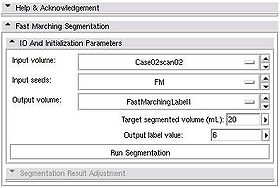
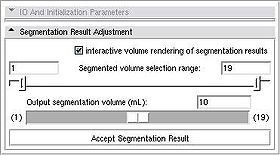
Segmentation workflow of this module consists of the two steps: Initialization and adjustment of the segmentation result.
Once these items have been specified, push Run Segmentation button to initiate the segmentation process.
Once the satisfactory result is achieved, Accept Segmentation Result button will finalize the segmentation. |
|
Usage tips:
- Put bigger number as the estimate volume It is better to overestimate the volume of the structure in the initialization, than underestimate. Remember, that you can always scroll back in the contour evolution, but cannot exceed predefined volume limit.
- Play with the fiducial points If the segmentation result is not satisfactory, try adding more fiducials. If the region to be segmented has regions of different intensities, put fiducials in each of such regions. If all fails, try to segment separate regions one after another, and use ImageLabelCombine module to merge them together later.
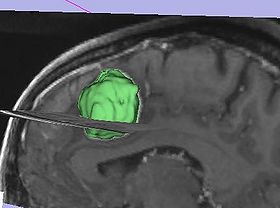
- Take advantage of volume rendering You can quickly see if there are "leaks", without the need to scroll through the slices. However, volume rendering may not show small "leaks".
IMPORTANT: Memory usage warning
The code allocates something like
16*sizeof(int)*dX*dY*dZ + dX*dY*dZ + (sizeof(int)+2*sizeof(double))*(max_intensity-min_intensity)
bytes in RAM. So, depending on what is the size of your image, you may run out of memory, which would cause the problem you describe. Chances of this happening depend on the size of the image, scalar type, scalar range, OS you use, whether you use 32- or 64- bit system, and how much memory you already have in use.
In order to reduce the chances of running out of memory, you can reduce the size of the required memory as follows:
1) use CropVolume module to extract a smaller VOI around the structure you are trying to segment (use resample constant 1.0). This will reduce dimensions of the input image.
2) you can rescale the intensities of your image to a smaller range
3) you can do both 1) and 2)
Development
Notes from the Developer(s)
This is a scripted Tcl module. The core fast marching algorithm is implemented in VTK classes vtkPichonFastMarching and vtkPichonFastMarchingPDF in Slicer/Base/Logic.
In the cases when the range of input intensities exceeds 300, the module internally rescales the input image intensities to be in [0,300]. Perhaps this should be exposed at the user level, or reconsidered.
Dependencies
Core modules: Fiducials, Volumes
Tests
The following tests verify the function of this module on the 3D Slicer dashboard:
vtkPichonFastMarchingTest1: exercise basic functionality of the vtkPichonFastMarching class
vtkPichonFastMarchingTest2: performs segmentation of the test meningioma image from one seed placed inside the tumor
Known bugs
#835: Fast marching will usually fail if you try to segment an object with uniform or very close to uniform intesity
#823: Volume rendering performance in Fast Marching
Follow this link to the Slicer3 bug tracker.
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Source code can accessed here
Links to documentation generated by doxygen.
More Information
Acknowledgment
References
- Wikipedia: Fast Marching method
- The algorithm implemented in this module is described in the following publication: Pichon E, Tannenbaum A, Kikinis R. A statistically based flow for image segmentation. Med Image Anal. 2004 Sep;8(3):267-74. PMID: 15450221. [1]
- The module has been transferred from Slicer2, legacy documentation from GATech here