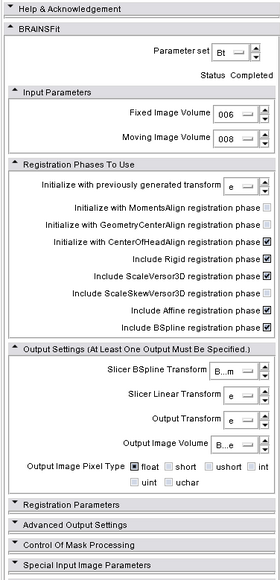
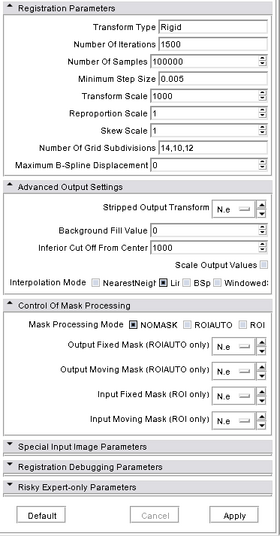
Modules:BRAINSFit
Return to Slicer 3.6 Documentation
Module Name
BRAINSFit
Example Applications: from the Registration Library
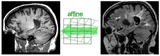
|
Affine Intra Subject Brain |
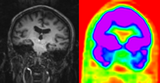
|
PET Brain |
| lleft link=http://www.na-mic.org/Wiki/index.php/Projects:RegistrationLibrary:RegLib_C29 | Diffusion MRI/DTI |
- And many more, see the Slicer Registration Library
(you can sort the table by method)
General Information
Module Type & Category
Type: CLI
Category: Registration
Authors, Collaborators & Contact
Author: Hans J. Johnson, hans-johnson -at- uiowa.edu, http://www.psychiatry.uiowa.edu
Contributors: Hans Johnson(1,3,4); Kent Williams(1); Gregory Harris(1), Vincent Magnotta(1,2,3); Andriy Fedorov(5), fedorov -at- bwh.harvard.edu (Slicer integration); (1=University of Iowa Department of Psychiatry, 2=University of Iowa Department of Radiology, 3=University of Iowa Department of Biomedical Engineering, 4=University of Iowa Department of Electrical and Computer Engineering, 5=Surgical Planning Lab, Harvard)
Module Description
Insight/Examples/Registration/ImageRegistration8.cxx This program is the most functional example of multi-modal 3D rigid image registration provided with ITK. ImageRegistration8 is in the Examples directory, and also sec. 8.5.3 in the ITK manual. We have modified and extended this example in several ways:- defined a new ITK Transform class, based on itkScaleSkewVersor3DTransform which has 3 dimensions of scale but no skew aspect.
- implemented a set of functions to convert between Versor Transforms and the general itk::AffineTransform and deferred converting from specific to more general representations to preserve transform information specificity as long as possible. Our Rigid transform is the narrowest, a Versor rotation plus separate translation.
- Added a template class itkMultiModal3DMutualRegistrationHelper which is templated over the type of ITK transform generated, and the optimizer used.
- Added image masks as an optional input to the Registration algorithm, limiting the volume considered during registration to voxels within the brain.
- Added image mask generation as an optional input to the Registration algorithm when meaningful masks such as for whole brain are not available, allowing the fit to at least be focused on whole head tissue.
- Added the ability to use one transform result, such as the Rigid transform, to initialize a more adaptive transform
- Defined the command line parameters using tools from the Slicer [ 3] program, in order to conform to the Slicer3 Execution model.
- Extensive testing as part of the BRAINS2 application suite, determined reasonable defaults for registration algorithm parameters. http://testing.psychiatry.uiowa.edu/CDash/
Usage
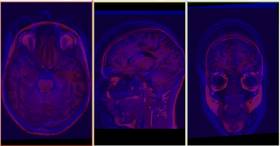
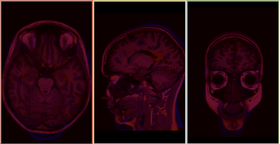
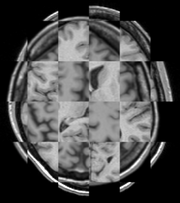
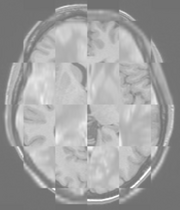
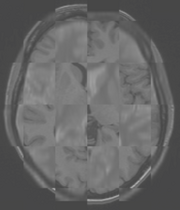
The BRAINSFit distribution contains a directory named TestData, which contains two example images. The first, test.nii.gz is a NIfTI format image volume, which is used the input for the CTest-managed regression test program. The program makexfrmedImage.cxx, included in the BRAINSFit distribution was used to generate test2.nii.gz, by scaling, rotating and translating test.nii.gz. You can see representative Sagittal slices of test.nii.gz (in this case, the fixed image, test2.nii.gz (the moving image), and the two images ’checkerboarded’ together to the right. To register test2.nii.gz to test.nii.gz, you can use the following command:
BRAINSFit --fixedVolume test.nii.gz \ --movingVolume test2.nii.gz \ --outputVolume registered.nii.gz \ --transformType Affine
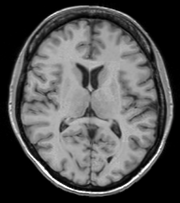
A representative slice of the registered results image registered.nii.gz is to the right. You can see from the Checkerboard of the Fixed and Registered images that the fit is quite good with Affine transform-based registration. The blurring of the registered images is a consequence of the initial scaling used in generating the moving image from the fixed image, compounded by the interpolation necessitated by the transform operation.
You can see the differences in results if you re-run BRAINSFit using Rigid, ScaleVersor3D, or ScaleSkewVersor3D as the ----transformType parameter. In this case, the authors judged Affine the best method for registering these particular two images; in the BRAINS2 automated processing pipeline, Rigid usually works well for registering research scans.
Use Cases, Examples
This module is especially appropriate for these use cases:
- Use Case 1: Same Subject: Longitudinal
For this use case we're registering a baseline T1 scan with a follow-up T1 scan on the same subject a year later. The two images are again available in the Slicer3/Applications/CLI/BRAINSTools/BRAINSCommonLib/TestData directory as testT1.nii.gz and testT1Longitudinal.nii.gz
First we set the fixed and moving volumes as well as the output transform and output volume names.
--fixedVolume testT1.nii.gz \ --movingVolume testT1Longitudinal.nii.gz \ --outputVolume testT1LongRegFixed.nii.gz \ --outputTransform longToBase.xform \
Since these are the same subject and very little has likely changed in the last year we'll use a Rigid registration. If the registration is poor or there are reasons to expect anatomical changes then additional transforms may be needed, in which case they can be added in a comma separated list, such as "Rigid,ScaleVersor3D,ScaleSkewVersor3D,Affine,BSpline".
--transformType Rigid \
The scans are the same modality so we'll use --histogramMatch to match the intensity profiles as this tends to help registration. If there are lesions or tumors that vary between images this may not be a good idea, as it will make it harder to detect differences between the images.
--histogramMatch \
To start with the best possible initial alignment we use --initializeTransformMode. We're working with human heads so we pick useCenterOfHeadAlign, which detects the center of head even with varying amounts of neck or shoulders present.
--initializeTransformMode useCenterOfHeadAlign \
ROI masks normally improve registration but we haven't generated any so we turn on --maskProcessingMode ROIAUTO.
--maskProcessingMode ROIAUTO \
The registration generally performs better if we include some background in the mask that way the tissue boundary is very clear. The parameter that expands the mask outside the brain is ROIAutoDilateSize (under Registration Debugging Parameters if using the GUI). These values are in millimeters so a good starting value is 3.
--ROIAutoDilateSize 3 \
Last we set the interpolation mode to be Linear, which is a decent tradeoff between quality and speed. If the best possible interpolation is needed regardless of processing time, select WindowedSync instead of linear.
--interpolationMode Linear
The full command is:
BRAINSFit --fixedVolume testT1.nii.gz \ --movingVolume testT1Longitudinal.nii.gz \ --outputVolume testT1LongRegFixed.nii.gz \ --outputTransform longToBase.xform \ --transformType Rigid \ --histogramMatch \ --initializeTransformMode useCenterOfHeadAlign \ --maskProcessingMode ROIAUTO \ --ROIAutoDilateSize 3 \ --interpolationMode Linear
| Program title | BRAINSFit |
| Program description | Uses the Mattes Mutual Registration algorithm to register a three-dimensional volume to a reference volume. Described in BRAINSFit: Mutual Information Registrations of Whole-Brain 3D Images, Using the Insight Toolkit, Johnson H.J., Harris G., Williams K., The Insight Journal, 2007. http://hdl.handle.net/1926/1291 |
| Program version | 3.0.0 |
| Program documentation-url | https://www.slicer.org/wiki/Modules:BRAINSFit |
- Use Case 2: Same Subject: MultiModal
For this use case we're registering a T1 scan with a T2 scan collected in the same sesson. The two images are again available in the Slicer3/Applications/CLI/BRAINSTools/BRAINSCommonLib/TestData directory as testT1.nii.gz and testT2.nii.gz
First we set the fixed and moving volumes as well as the output transform and output volume names.
--fixedVolume testT1.nii.gz \ --movingVolume testT2.nii.gz \ --outputVolume testT2RegT1.nii.gz \ --outputTransform T2ToT1.xform \
Since these are the same subject, same session we'll use a Rigid registration.
--transformType Rigid \
The scans are different modalities so we absolutely DO NOT want to use --histogramMatch to match the intensity profiles as this would try to map the T2 intensities into T1 intensities, resulting in an image that was neither, and hence useless.
To start with the best possible initial alignment we use --initializeTransformMode. We're working with human heads so we pick useCenterOfHeadAlign, which detects the center of head even with varying amounts of neck or shoulders present.
--initializeTransformMode useCenterOfHeadAlign \
ROI masks normally improve registration but we haven't generated any so we turn on --maskProcessingMode ROIAUTO.
--maskProcessingMode ROIAUTO \
The registration generally performs better if we include some background in the mask that way the tissue boundary is very clear. The parameter that expands the mask outside the brain is ROIAutoDilateSize (under Registration Debugging Parameters if using the GUI). These values are in millimeters so a good starting value is 3.
--ROIAutoDilateSize 3 \
Last we set the interpolation mode to be Linear, which is a decent tradeoff between quality and speed. If the best possible interpolation is needed regardless of processing time, select WindowedSync instead of linear.
--interpolationMode Linear
The full command is:
BRAINSFit --fixedVolume testT1.nii.gz \ --movingVolume testT2.nii.gz \ --outputVolume testT2RegT1.nii.gz \ --outputTransform T2ToT1.xform \ --transformType Rigid \ --initializeTransformMode useCenterOfHeadAlign \ --maskProcessingMode ROIAUTO \ --ROIAutoDilateSize 3 \ --interpolationMode Linear
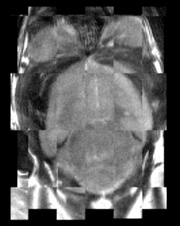
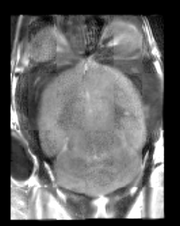
- Use Case 3: Mouse Registration
Here we'll register brains from two different mice together. The fixed and moving mouse brains used in this example are available in the Slicer3/Applications/CLI/BRAINSTools/BRAINSCommonLib/TestData directory.
First we set the fixed and moving volumes as well as the output transform and output volume names.
--fixedVolume mouseFixed.nii.gz \ --movingVolume mouseMoving.nii.gz \ --outputVolume movingRegFixed.nii.gz \ --outputTransform movingToFixed.xform \
Since the subjects are different we are going to use transforms all the way through BSpline. Again, building up transforms one type at a time can't hurt and might help, so we're including all transforms from Rigid through BSpline in the transformType parameter.
--transformType Rigid,ScaleVersor3D,ScaleSkewVersor3D,Affine,BSpline \
The scans are the same modality so we'll use --histogramMatch.
--histogramMatch \
To start with the best possible initial alignment we use --initializeTransformMode but we are't working with human heads so we can't pick useCenterOfHeadAlign. Instead we pick useMomentsAlign which does a reasonable job of selecting the centers of mass.
--initializeTransformMode useMomentsAlign \
ROI masks normally improve registration but we haven't generated any so we turn on --maskProcessingMode ROIAUTO.
--maskProcessingMode ROIAUTO \
Since the mouse brains are much smaller than human brains there are a few advanced parameters we'll need to tweak, ROIAutoClosingSize and ROIAutoDilateSize (both under Registration Debugging Parameters if using the GUI). These values are in millimeters so a good starting value for mice is 0.9.
--ROIAutoClosingSize 0.9 \ --ROIAutoDilateSize 0.9 \
Last we set the interpolation mode to be Linear, which is a decent tradeoff between quality and speed. If the best possible interpolation is needed regardless of processing time, select WindowedSync instead of linear.
--interpolationMode Linear
The full command is:
BRAINSFit --fixedVolume mouseFixed.nii.gz \ --movingVolume mouseMoving.nii.gz \ --outputVolume movingRegFixed.nii.gz \ --outputTransform movingToFixed.xform \ --transformType Rigid,ScaleVersor3D,ScaleSkewVersor3D,Affine,BSpline \ --histogramMatch \ --initializeTransformMode useMomentsAlign \ --maskProcessingMode ROIAUTO \ --ROIAutoClosingSize 0.9 \ --ROIAutoDilateSize 0.9 \ --interpolationMode Linear
Quick Tour of Features and Use
This section was partially generated with a python script to convert Slicer Execution Model xml files into MediaWiki compatible documentation
|
Development
Notes from the Developer(s)
This is a thin wrapper program around the BRAINSFitHelper class in BRAINSCommonLib. The BRAINSFitHelper class is intended to allow all the functionality of BRAINSFit to be easily incorporated into another program by including a single header file, linking against the BRAINSCommonLib library, and adding code similar to the following to your application:
typedef itk::BRAINSFitHelper HelperType;
HelperType::Pointer myHelper=HelperType::New();
myHelper->SetTransformType(localTransformType);
myHelper->SetFixedVolume(extractFixedVolume);
myHelper->SetMovingVolume(extractMovingVolume);
myHelper->StartRegistration();
currentGenericTransform=myHelper->GetCurrentGenericTransform();
MovingVolumeType::ConstPointer preprocessedMovingVolume = myHelper->GetPreprocessedMovingVolume();
/* Optional member functions that can also be set */
myHelper->SetHistogramMatch(histogramMatch);
myHelper->SetNumberOfMatchPoints(numberOfMatchPoints);
myHelper->SetFixedBinaryVolume(fixedMask);
myHelper->SetMovingBinaryVolume(movingMask);
myHelper->SetPermitParameterVariation(permitParameterVariation);
myHelper->SetNumberOfSamples(numberOfSamples);
myHelper->SetNumberOfHistogramBins(numberOfHistogramBins);
myHelper->SetNumberOfIterations(numberOfIterations);
myHelper->SetMaximumStepLength(maximumStepSize);
myHelper->SetMinimumStepLength(minimumStepSize);
myHelper->SetRelaxationFactor(relaxationFactor);
myHelper->SetTranslationScale(translationScale);
myHelper->SetReproportionScale(reproportionScale);
myHelper->SetSkewScale(skewScale);
myHelper->SetUseExplicitPDFDerivativesMode(useExplicitPDFDerivativesMode);
myHelper->SetUseCachingOfBSplineWeightsMode(useCachingOfBSplineWeightsMode);
myHelper->SetBackgroundFillValue(backgroundFillValue);
myHelper->SetInitializeTransformMode(localInitializeTransformMode);
myHelper->SetMaskInferiorCutOffFromCenter(maskInferiorCutOffFromCenter);
myHelper->SetCurrentGenericTransform(currentGenericTransform);
myHelper->SetSplineGridSize(splineGridSize);
myHelper->SetCostFunctionConvergenceFactor(costFunctionConvergenceFactor);
myHelper->SetProjectedGradientTolerance(projectedGradientTolerance);
myHelper->SetMaxBSplineDisplacement(maxBSplineDisplacement);
myHelper->SetDisplayDeformedImage(UseDebugImageViewer);
myHelper->SetPromptUserAfterDisplay(PromptAfterImageSend);
myHelper->SetDebugLevel(debugLevel);
if(debugLevel > 7 )
{
myHelper->PrintCommandLine(true,"BF");
}
Dependencies
BRAINSFit depends on Slicer3 (for the SlicerExecutionModel support) and ITK. BRAINSFit depends on the BRAINSCommonLib library
Tests
Nightly testing of the development head can be found at: http://testing.psychiatry.uiowa.edu/CDash
Known bugs
Links to known bugs and feature requests are listed at:
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Links to the module's source code:
Source code:
More Information
Acknowledgment
This research was supported by funding from grants NS050568 and NS40068 from the National Institute of Neurological Disorders and Stroke and grants MH31593, MH40856, from the National Institute of Mental Health.
References
- BRAINSFit: Mutual Information Registrations of Whole-Brain 3D Images, Using the Insight Toolkit, Johnson H.J., Harris G., Williams K., The Insight Journal, 2007.