Documentation:Nightly:Registration:RegistrationLibrary:RegLib C01
Contents
Slicer Registration Library Case #1: Follow-up MRI of Brain Tumor
Input
| fixed image | moving image |
Description
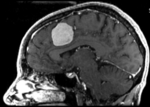
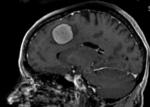
This case contains a baseline-followup pair of brain MRI. Goal is to align the follow-up image with the reference/baseline scan to assess changes, particularly in the tumor region.
Approach: Since image contrast and content is very similar in both images, and there is a good amount of initial overlap, the standard Slicer automated registration should work well for this. If possible, and affine registration would be preferable to a rigid one to address the typical distortions that occur in MRI and often vary between scans. However to assure the tumor and tumor changes do not influence the result, we will build a mask that excludes the tumor region from participating (i.e. actively contributing toward the result transform). We also use this case as an example to illustrate how to quickly generate masks for registration.
Modules used
Download (from NAMIC MIDAS)
Keywords
MRI, brain, head, intra-subject, meningioma
Video Screencasts
- Movie/screencast showing generating a registration mask to exclude the tumor
- Movie/screencast showing rigid and affine registration
Procedure
- Rigid Registration: open the General Registration (BRAINS) module
- Input Images: Fixed Image Volume: MR_1_baseline image
- 'Input Images: Moving Image Volume: MR_2_followup image
- Output Settings:
- Slicer Linear Transform (create new transform, rename to: "Xf1_Rigid")
- Output Image Volume none
- Registration Phases: select/check Rigid
- Initialization Phases: none
- Main Parameters:
- Leave all other settings at default
- click: Apply; runtime < 10 sec
- Building Registration Masks: We have now generate two masks that exclude the tumor region from contributing to the registration.
- open the Editor module
- As Master Volume, select the MR_1_baseline image
- click on the threshold tool and lower the threshold until the entire volume is selected (about a threshold of 5). Then click Apply.
- select the Brush tool. Increase the radius so that the brush completely covers the tumor (about 18mm).
- set the paint color to 0 (background).
- place the brush on an axial slice over the tumor. Use a left click to paint and the arrow keys to switch to the next slice. Repeat for all slices showing the tumor.
- save the mask as "MR_1_baseline_mask.nrrd"
- Now we generate a mask for MR_2. We can repeat the above or we can generate a mask by sending the first one through the inverse of the rigid transform we already computed.
- Go to the Transforms module.
- Active Transform: select the "Xf1_Rigid" transform we created above
- scroll down and click on the Invert button. This inverts the transform, which now points from MR_1 to MR_2. So now we can use it to generate a mask for MR_2 from the mask of MR_1. We must not forget to undo the invert afterwards!
- Go to the BRAINSResample module
- Image To Warp: select MR_1_baseline_mask
- Reference Image: select MR_2_followup
- Outputs:OutputImage: create & rename new, rename to "MR_2_followup_mask"
- Pixel Type: select uint or uchar
- Transform File: select "Xf1_Rigid"
- Interpolation Mode: select Nearest Neighbor
- Click Apply.
- Return to the Transforms module and invert the "Xf1_Rigid" transform again to return it to its original state.
- Go to the Volumes module
- Active Volume: select the new "MR_2_followup_mask"
- scroll down and check the 'labelmap checkbox.
- save the new mask.
- we now have two masks for use in the affine registration
- Affine registration: We have now increase the DOF to 12 in the form of an affine transform that also corrects scale and allows shear.
- Input Images: Fixed Image Volume: MR_1_baseline
- 'Input Images: Moving Image Volume: MR_2_followup
- Output Settings:
- Slicer Linear Transform (create new transform, rename to: "Xf2_Affine")
- Output Image Volume none
- Registration Phases: uncheck Rigid, select "Rigid+Scale" and "Affine"
- Initialization Transform: select "Xf1_Rigid"
- Initialization Transform Mode: off
- Main Parameters:
- Number Of Samples: increase to 300,000
- Masking Option: select "ROI" button
- Masking input fixed: select "MR_1_baseline_mask"
- Masking input moving: select "MR_2_followup_mask"
- click: Apply
- Verify
- place "MR_1_baseline" in the background and "MR_2_followup" into the foreground.
- place fade slider halfway
- Go to the Data module
- Drag "MR_2_followup" into the "Xf2_Affine" transform node, if it isn't there already.
- Apply the transforms without resampling / Save result images
- for linear (rigid to affine) transforms you have two options for generating a result image in the new orientation: with and without resampling. The method without resampling leaves the original image data untouched and writes the new orientation into the image header. This preserves the full original image quality and is the preferred method, provided that the software using the result image can read the spatial information in the header correctly. If you open an image with a software that does not account for this header information, the image will appear in its original orientation. The resampling method will create a new image volume by interpolating the data into the new space. This has the advantage that your registered volume will have matching size and resolution as your reference image, but it is associated with some data loss from the interpolation during resampling.
- Apply Transform without resampling:
- Go to the Data module
- left click on the moving image node and drag it into/onto the transform node. You should see a "+" sign appear next to the transform node indicating the volume is now placed under that transform. If the image is in either foreground or background and visible in the slice views or 3D view, you will see the image update to its new position immediately.
- right click on the image node and from the pop-up menu, select Harden Transform. This will apply the transform and move the image node back outside onto the main level.
- double click on the image node and rename it to document that this is no longer the original image but the reoriented one. Note that the image in its original position is replaced in memory and no longer accessible. There is no "undo" for this step, so if you wish to return to the original image, you must reload it from the original file. Be careful not to accidentally overwrite the original image file with the resampled one. Use a new filename when saving the image.
- Alternative: Apply Transform with resampling:
- Go to the BRAINSResample module
- Image To Warp: select MR_2_followup
- Reference Image: select MR_1_baseline
- Outputs:OutputImage: create & rename new, rename to "MR_2_followup_Xf2"
- Pixel Type: select short , or match the datatype of the original image.
- Transform File: select "Xf2_Affine"
- Interpolation Mode: select "WindowedSinc" (recommended for 3D MRI). For labelmaps select Nearest Neighbor
- Click Apply. This will generate a new volume with the same resolution as the reference image.
- Go to the BRAINSResample module
Registration Results
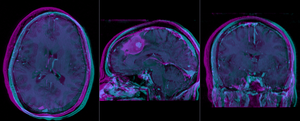 |
baseline and follow-up before registration (click to enlarge) |
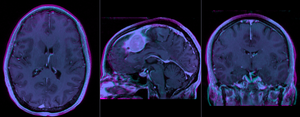 |
after rigid alignment. Note the residual shape differences (click to enlarge) |
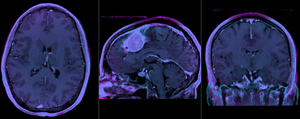 |
after affine registration (click to enlarge) |
Discussion: Key Strategies
- The affine registration seeks to address residual distortions that are typical for longitudinal MRI. However to prevent the tumor (and putative tumor change) from affecting the registration, we mask out the tumor so that it does not contribute to the registration. The assumption is thus that aside from the tumor we expect the two serial scans to be matching exactly, i.e. any differences in size or shape is considered an artifact we wish to remove by registration.