Home < Documentation < Nightly < Modules < BoneTextureSerializer
Introduction and Acknowledgements
|
Extensions: BoneTextureExtesion
Author: Jean-Baptise Vimort, Kitware Inc.
Contributors: Beatriz Paniagua (Kitware Inc), Lucia Cevidanes (University of Michigan - School of Dentistry), Erika Benavides (University of Michigan - School of Dentistry), Antônio Carlos de Oliveira Ruellas (University of Michigan - School of Dentistry)
Contact: Jean-Baptiste Vimort, <email>jb.vimort@kitware.com</email>
Acknowledgments: This work was supported by the National Institute of Health (NIH) National Institute for Dental and Craniofacial Research (NIDCR) grant R21DE025306 (Textural Biomarkers of Arthritis for the Subchondral Bone in the Temporomandibular Joint), NIDCR grant R01DE024450 (Quantification of 3D bony Changes in
Temporomandibular Joint Osteoarthritis) and National Institute of Biomedical Imaging and Bioengineering NIBIB) grant R01EB021391 (Shape Analysis Toolbox for Medical Image Computing Projects).
License: Apache License, Version 2.0
|
Module Description
This module provides a simple way to serialize the BoneTexture module.
In order to make it work, all the input data should be in the same folder and named as following:
- The input scan should be named ScanXXX.nrrd, where XXX is the ID of the case
- If it exists, the input mask should be named SegmCXXX.nrrd, where XXX is the ID corresponding to the the input scan
Interface
Use Cases
Inputs
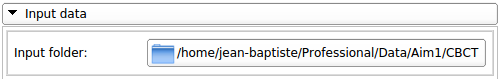 BoneTextureSerializer Input |
This section allows the user to specify the input folder, the folder specified should contain all the inputs (scans and segmentations) as described previously.
|
Parameters
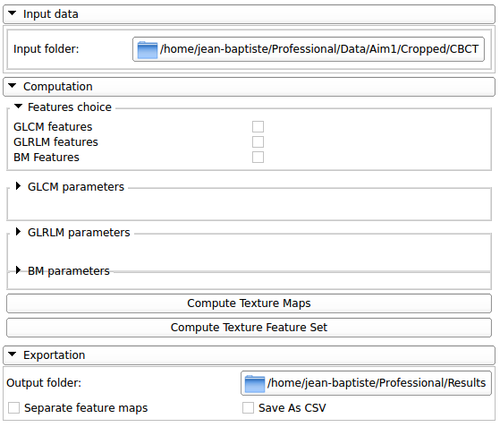
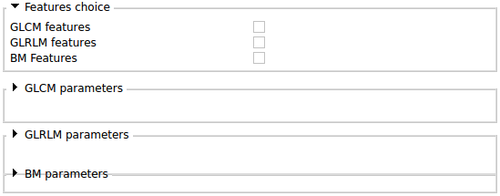 BoneTextureSerializer parameters |
This section allows the user to choose:
- The Type of features wanted: Run length features or co-occurrence features
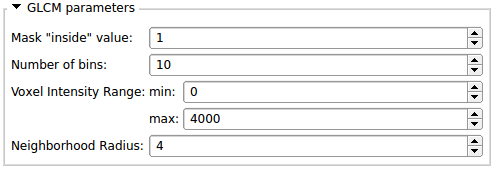
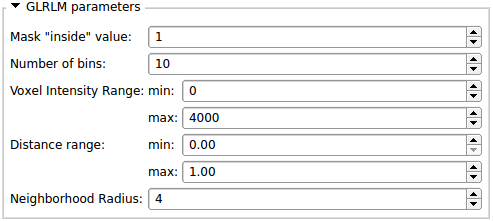
- All the parameters necessary to run each algorithms
- The type of outputs wanted: texture features or texture maps
|
Exportation
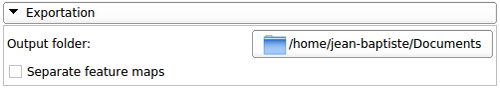 Image:BoneTextureSerializer Image:BoneTextureSerializer |
This section allows the user to specify the folder where all the results will be saved:
- The texture features for the whole image will be saved in a csv file
- The texture maps can either be saved all in a same file (a diffusion weighted image containing all the feature maps of a case) or separated (each feature map will be saved as a single volume)
|
Additional Information
Similar Modules
N/A
Information for Developers
The source code is available on github