Home < Documentation < Nightly < Extensions < PBNRR
Introduction and Acknowledgements
|
Extension: Physics-Based Non-Rigid Registration (PBNRR)
Acknowledgments:
This work is funded mainly by the ARRA funds for the ITK-v4 implementation with grant number:NLMA2D2201000586P. In addition, this work is supported in part by the NSF grant CCF-1439079 and by the John Simon Guggenheim Foundation and the Richard T. Cheng Endowment.
Maintainer: Angelos Angelopoulos (CRTC)
Author: Fotis Drakopoulos
Contributors: Angelos Angelopoulos (CRTC), Fotis Drakopoulos (CRTC), Yixun Liu (CRTC), Andriy Kot (CRTC), Andrey Fedorov (SPL B&W Harvard), Olivier Clatz (Asclepios INRIA), Nikos Chrisochoides (CRTC)
Contact: Nikos Chrisochoides (<email>npchris@gmail.com</email>), Angelos Angelopoulos, (<email>aangelos28@gmail.com</email>)
Website: https://crtc.cs.odu.edu/
License: BSD
|
| Center for Real-time Computing
|
|
|
Module Description
The module Non-Rigid Registers a moving to a fixed MRI. It uses a linear homogeneous bio-mechanical model to compute a dense deformation field that defines a transformation for every point in the fixed image to the moving image. The method includes three components (Feature Point Selection, Block Matching and Finite Element Solver) combine together to provide a user-friendly interface.
Use Cases
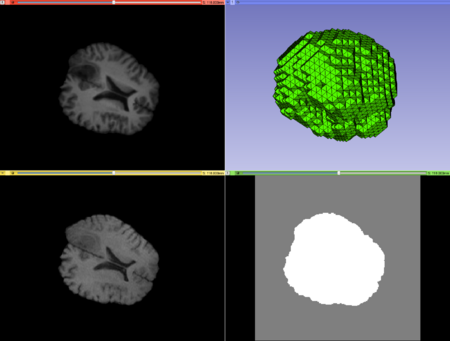 Case1 Input; top left: moving image, top right: mesh, bottom left: fixed image, bottom right: mask image. |
 Case1 Output; top left: warped image, top right: warped mesh, bottom left: direct deformation field, bottom right: inverse deformation field. |
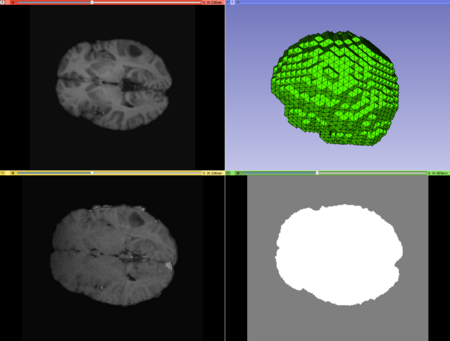 Case2 Input; top left: moving image, top right: mesh, bottom left: fixed image, bottom right: mask image. |
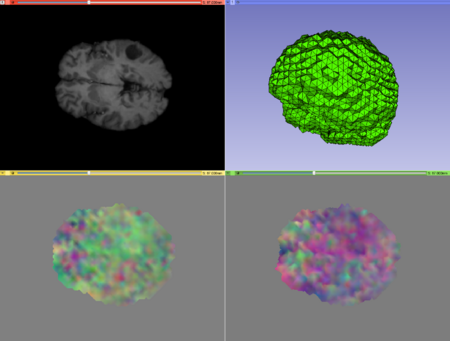 Case2 Output; top left: warped image, top right: warped mesh, bottom left: direct deformation field, bottom right: inverse deformation field. |
Tutorials
Panels and their use
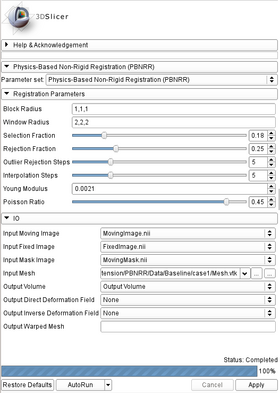
- Registration Parameters:
- Block Radius: Radius (in image voxels) of the selected image blocks in each dimension (default: 1,1,1).
- Window Radius: Radius (in image voxels) of the Block Matching window in each dimension (default: 5,5,5).
- Selection Fraction: Fraction of the selected blocks from the total number of image blocks. Value should be between [0.01,1] (default: 0.05).
- Rejection Fraction: Fraction of the rejected blocks from the number of the selected image blocks. Value should be between [0.01,1) (default: 0.25).
- Outlier Rejection Steps: Number of outlier rejection steps. Value should be between [1,20] (default: 5).
- Interpolation Steps: Number of interpolation steps. Value should be between [1,20] (default: 5).
- Young Modulus: Young Modulus of the linear bio-mechanical model (default: 0.0021 N/mm2).
- Poisson Ratio: Poisson Ratio of the linear bio-mechanical model. Value should be between [0.10,0.49] (default: 0.45).
- Input/Output:
- Input Moving Image: The input moving image (preoperative).
- Input Fixed Image: The input fixed image (intraoperative).
- Input Mask Image: The input moving mask.
- Input Mesh: The input mesh.
- Output Volume: Moving image to the fixed image coordinate frame (optional).
- Output Direct Deformation Field: Transform calculated that aligns the fixed and moving image. Maps positions in the moving coordinate frame to the fixed coordinate frame (optional).
- Output Inverse Deformation Field: Transform calculated that aligns the fixed and moving image. Maps positions in the fixed coordinate frame to the moving coordinate frame (optional).
- Output Warped Mesh: The warped tetrahedral mesh in vtk file format (optional).
|
|
Similar Modules
N/A
References
- Olivier Clatz, Hervé Delingette, Ion-Florin Talos, Alexandra J. Golby, Ron Kikinis, Ferenc Jolesz, Nicholas Ayache, and Simon Warfield,“Robust Non-Rigid Registration to Capture Brain Shift from Intra-Operative MRI“, IEEE Transactions on Medical Imaging, 24(11):1417-1427, Nov. 2005.
- Liu Y, Kot A, Drakopoulos F, Yao C, Fedorov A, Enquobahrie A, Clatz O and Chrisochoides NP (2014), ”An ITK implementation of a physics-based non-rigid registration method for brain deformation in image-guided neurosurgery”, Frontiers in Neuroinformatics 8:33. doi: 10.3389/fninf.2014.00033
Information for Developers