Documentation/4.3/Modules/AirwaySegmentation
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
|
Extension: AirwaySegmentation | |||||||||
|
Module Description
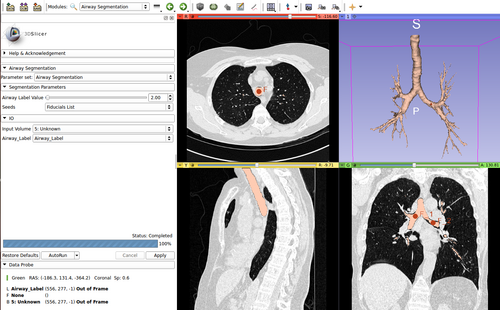
AirwaySegmentation is a simple CLI module for airway segmentation starting from chest CT images. This CLI uses a modified version of ITK's itkConnectedThresholdImageFilter to segment all the pixels with an intensity below a threshold. The threshold is automatically identified by the module. The user has to specify three fiducial points: one in the trachea, and two in the main bronchi of the left and right lungs. These fiducials are the starting points for the region growing segmentation. No more than 3 seed points are allowed.
Use Cases
- Airway Segmentation starting from chest CT datasets
Tutorials
N/A
Panels and their use
- Segmentation Parameters: Input parameters for segmentation.
- Airway Label Value: The integer value (0-255) to use for the segmentation results. This will determine the color of the segmentation that will be generated by the algorithm.
- Seeds: Seed points for the algorithm. Three seeds points must be placed. The first one in the trachea, the others in the main bronchi of the left and right lungs.
- IO: Input and Output parameters.
- Input Volume: Input chest CT dataset to be segmented.
- Output Parameters: Output label.
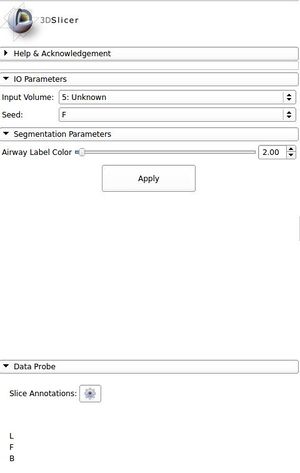
The user interface panel:
Similar Modules
N/A
References
N/A
Information for Developers
The code is available at Github.