Difference between revisions of "Documentation/Nightly/Modules/FiberBundleLabelSelect"
From Slicer Wiki
Zhangfanmark (talk | contribs) |
m (Text replacement - "\bhttps:\/\/www\.slicer\.org\/slicerWiki\/index\.php\/([^ ]+)\b " to "https://www.slicer.org/wiki/$1 ") |
||
| (7 intermediate revisions by 3 users not shown) | |||
| Line 8: | Line 8: | ||
{{documentation/{{documentation/version}}/module-introduction-start|{{documentation/modulename}}}} | {{documentation/{{documentation/version}}/module-introduction-start|{{documentation/modulename}}}} | ||
{{documentation/{{documentation/version}}/module-introduction-row}} | {{documentation/{{documentation/version}}/module-introduction-row}} | ||
| − | + | ||
| − | + | {{documentation/{{documentation/version}}/module-acknowledgements}} | |
| − | + | Contact: <email>slicer-users@bwh.harvard.edu</email><br> | |
| − | Contact: | + | Website: http://slicerdmri.github.io/ |
| + | |||
{{documentation/{{documentation/version}}/module-introduction-row}} | {{documentation/{{documentation/version}}/module-introduction-row}} | ||
{{documentation/{{documentation/version}}/module-introduction-logo-gallery | {{documentation/{{documentation/version}}/module-introduction-logo-gallery | ||
| Line 17: | Line 18: | ||
|Image:Logo-splnew.jpg|Surgical Planning Laboratory | |Image:Logo-splnew.jpg|Surgical Planning Laboratory | ||
|Image:NAC-logo.png|NAC | |Image:NAC-logo.png|NAC | ||
| − | |Image: | + | |Image:Wholebraintractography.png|Whole brain tractography |
| + | |Image:CCSelectedTracts.png|Corpus callosum (CC) tract selection | ||
}} | }} | ||
{{documentation/{{documentation/version}}/module-introduction-end}} | {{documentation/{{documentation/version}}/module-introduction-end}} | ||
| Line 24: | Line 26: | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Module Description}} | {{documentation/{{documentation/version}}/module-section|Module Description}} | ||
| − | + | {{documentation/{{documentation/version}}/module-description}} | |
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
| Line 36: | Line 38: | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Tutorials}} | {{documentation/{{documentation/version}}/module-section|Tutorials}} | ||
| − | + | Links to tutorials that use this module | |
| − | + | * Slicer4 Fiber Bundle Selection and Scalar Measurements Tutorial: https://www.slicer.org/wiki/Documentation/4.5/Training#Fiber_Bundle_Selection_and_Scalar_Measurements * Tractography selection and measurements in command line interface (CLI) mode: http://dmri.slicer.org/tutorials/tractography_measurement | |
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Panels and their use}} | {{documentation/{{documentation/version}}/module-section|Panels and their use}} | ||
| − | + | {{documentation/{{documentation/version}}/module-parametersdescription}} | |
| − | |||
| − | { | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Similar Modules}} | {{documentation/{{documentation/version}}/module-section|Similar Modules}} | ||
* Tractography Label Map Seeding | * Tractography Label Map Seeding | ||
| + | |||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|References}} | {{documentation/{{documentation/version}}/module-section|References}} | ||
| − | + | * http://slicerdmri.github.io/ | |
| − | |||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-section|Information for Developers}} | {{documentation/{{documentation/version}}/module-section|Information for Developers}} | ||
| − | + | * https://github.com/SlicerDMRI | |
| − | |||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
{{documentation/{{documentation/version}}/module-footer}} | {{documentation/{{documentation/version}}/module-footer}} | ||
<!-- ---------------------------- --> | <!-- ---------------------------- --> | ||
Latest revision as of 13:59, 27 November 2019
Home < Documentation < Nightly < Modules < FiberBundleLabelSelect
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
|
Contact: <email>slicer-users@bwh.harvard.edu</email> | |||||||||||
|
Module Description
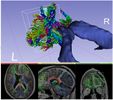
Fiber bundle label select allows a user to select tracts passing or not passing through single or multiple labels. (Selection region label maps may be created with the Editor module or other segmentation tools.)
Use Cases
Most frequently used for these scenarios:
- Use Case 1: Filter out a subset of DTI fiber tracts which are passing through selected region(s) defined in the label map volume.
- Use Case 2: Filter out a subset of DTI fiber tracts which are NOT passing through selected region(s) defined in the label map volume.
- Use Case 3: A combination of Case 1 and Case 2.
Tutorials
Links to tutorials that use this module
- Slicer4 Fiber Bundle Selection and Scalar Measurements Tutorial: https://www.slicer.org/wiki/Documentation/4.5/Training#Fiber_Bundle_Selection_and_Scalar_Measurements * Tractography selection and measurements in command line interface (CLI) mode: http://dmri.slicer.org/tutorials/tractography_measurement
Panels and their use
Parameters:
- IO: Input/output parameters
- Selection Region Label Map (InputLabel_A): Label map volume defining selection region
- Input Fiber Bundle (InputFibers): Input tractography for region-based selection.
- Output Fiber Bundle (OutputFibers): Selected tractography result from region-based selection.
- Tract selection region labels:
- Inclusion labels (comma-separated) (PassLabel): Comma-separated list of label values defining inclusion region. Fibers passing through these labels will be retained.
- Inclusion label combination logic (PassOperation): AND: Fiber must pass through all specified labels.
OR: Fiber must pass through any specified label (at least one). - Exclusion labels (comma-separated) (NotPassLabel): Comma-separated list of label values defining exclusion region. Fibers passing through these labels will be removed.
- Exclusion label combination logic (NoPassOperation): AND: Any fiber passing through all specified labels is excluded.
OR: Fibers passing through any specified label (at least one) are excluded.
- Advanced Settings: Advanced settings
- Sampling distance (mm) (SamplingDistance): Sampling Distance
List of parameters generated transforming this XML file using this XSL file. To update the URL of the XML file, edit this page.
Similar Modules
- Tractography Label Map Seeding