Slicer 3.6:Training
From Slicer Wiki
Home < Slicer 3.6:Training
Contents
Slicer 3.6 Tutorials
- The following table contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment (version 3.6 release) to accomplish certain tasks.
- For tutorials for other versions of Slicer, please visit the Slicer training portal.
- For questions related to the Slicer3 Compendium, please send an e-mail to Sonia Pujol, Ph.D
| Category | Tutorial | Sample Data | Image |
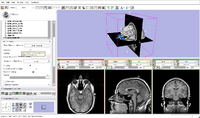
| Core | Slicer3Minute Tutorial The Slicer3Minute tutorial is an introduction to the advanced 3D visualization capabilities of Slicer3.6. Audience: First time users. |
Slicer3Minute Data The Slicer3Minute dataset contains an MR scan of the brain and 3D reconstructions of the anatomy |
|
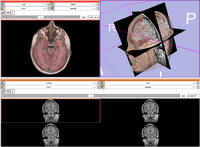
| Core | Slicer3Visualization Tutorial The Slicer3Visualization tutorial guides through 3D data loading and visualization in Slicer3.6. Audience: All beginners including clinicians, scientists, engineers and programmers. |
Slicer3Visualization Data The Slicer3VisualizationDataset contains two MR scans of the brain, a pre-computed labelmap and 3D reconstructions of the anatomy. |
|
| Core | Programming in Slicer3 Tutorial The Programming in Slicer3 tutorial is an introduction to the the integration of stand-alone programs outside of the Slicer3 source tree. Audience: Programmers and Engineers. |
HelloWorld Plugin The HelloWorld tutorial dataset contains an MR scan of the brain and pre-computed xml and C++ files for integrating the Hello World plug-in to Slicer3. |
|
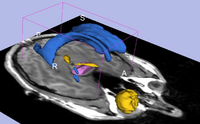
| Core | Interactive Editor-PDF Interactive Editor-PPT Shows how to use the interactive editing tools in Slicer. Audience: All users and developers. |
Editor Data This dataset contains a MR dataset of the brain. |
|
| Core | Manual Registration-PDF Shows how to manually/interactively align two images in Slicer3.6 Audience: First time & early users. |
Manual Registration Data This dataset contains two brain MRI of a single subject, obtained in different orientations. |
|
| Specialized | Diffusion MRI tutorial-PDF Diffusion MRI tutorial-PPT This tutorial guides you through the process of loading diffusion MR data, estimating diffusion tensors, and performing tractography of white matter bundles. Audience: All users and developers. |
Diffusion Data |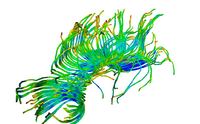 
|
| Specialized | Change Tracker Tutorial This tutorial describes the use of ChangeTracker module to detect changes in tumor volume from two MRI scans. Audience: All users interested in longitudinal analysis of pathology. |
Training Data download is integrated with the ChangeTracker module (see Tutorial) |
Summer 2010 Tutorial Contest Entries (under construction)
| Category | Tutorial | Sample Data | Image |
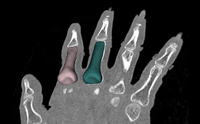
| Specialized | Meshing Workflow tutorial Audience: All users and developers. |
Meshing Workflow Data |
|
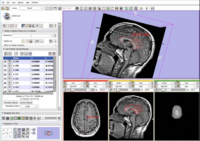
| Specialized | Fiducials tutorial Audience: All users and developers. |
Fiducials Data |
|
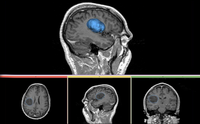
| Specialized | Robust Statistic Segmenter Audience: All users and developers. |
Robust Statistic Segmenter Data |
|
| Specialized | Longitudinal lesion comparison Audience: All users and developers. |
Longitudinal Lesion Comparison Data | 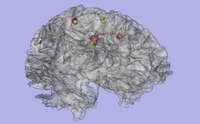
|
| Specialized | Robot-Assisted MRI-Guided Prostate Biopsy Audience: All users and developers. |
Robot-Assisted MRI-Guided Prostate Biopsy | 
|
| Specialized | Atlas Label Fusion & Surface Registration Audience: All users and developers. |
Atlas Label Fusion & Surface Registration | 
|
Software Installation
- The Slicer download page contains information on how to obtain a compiled version of Slicer for a variety of platforms and where to find the source code for Slicer 3.
Software Documentation
- For the Slicer 3.6 manual pages please click here. These pages are the reference manual for Slicer 3.6 and briefly explain the functionality found in panels and modules.