Slicer3:UIDesign:WorkingProblems:BCAnalysis:Dynamic
Back to Project Overview
Contents
- 1 Perfusion Analysis for Breast Cancer
Perfusion Analysis for Breast Cancer
Goal
Goal is to accommodate basic RECIST assessment for cancer treatment.
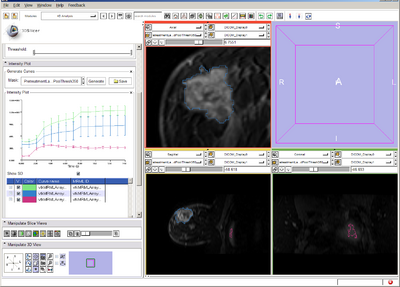
Specifically for Perfusion analysis: Develop a module that loads perfusion datasets and performs fits intensity timecourse data to Tofts Kinetic Model.
Perfusion Analysis Workflow
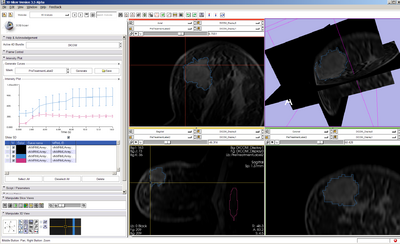
- Visualize the dataset dynamically
- Specify an ROI that includes tumor and apply to all timepoints Question: ROI or VOI? (either -- if slice, must be same slice)
- Plot contrast dilution curve
- show time to peak in tumor
- Specify an ROI that includes blood pool and apply to all timepoints Question: again, ROI or VOI?
- Plot contrast dilution curve for this (on same graph)
- show time to peak in blood pool
- show transit time (distance between peaks in each plot)
- save out timepoints ( Intensity(t) for each ROI -- include multiple tumors if desired)
Meeting notes
08/20/2009 meeting with Ron: Jeff will provide:
- indication (screen shot) of the blood pool VOI to use
- indication of the mapping between dicom files and slice, timepoint, channel to use during the load experiments
- paper describing kinetic curve fitting, based on Toft's model extracellular volume fraction and permeability used by Kasia Macura at JHU.
Systems integration notes
Loading Data:
From Jeff -- first dataset contains 15 timepoints, each 256x256x63 -- so we've found a bug in either:
- the timeseries loader in Slicer (probably in the thing that parses dicom header), or
- we have a malformed DICOM header.
This is causing Slicer to load 63x15 volumes containing a single 256x256 slice. I'll dig in to this today and see what's going on. Shouldn't be too hard to find. Junichi has given me a DCE dataset that loads properly for comparison.
- Fixed with changes to automatic parsing of DICOM data. Will stub out, check into svn today, and ask Junichi to test it on his datasets -- make sure it works for all cases before exposing in the repository.
- Changes don't work with Junichi's DCE datasets -- checked these in Slicer and they do not contain the "Number Of Temporal Points" metadata info in the DICOM header. So, have hybridized the automated load to:
- look for this tag
- if tag exists, use it to sort volumes into bundles (worked for JHU DCE, doesn't work for Meso_MRI datasets)
- if tag does not exist, use junichi's method to sort volumes into bundles (works for Meso_MRI datasets, didn't work for JHU DCE)
- this is implemented now, testing.... if works for both, will check into svn.
CHANGES TO CODE & BUILD INFRASTRUCTURE to get FourDImage and FourDAnalysis to compile
The 4D Image analysis package is required for plotting -- need to expose this in Slicer3/Modules/CMakelists.txt file. Discussed with Junichi about possibility of having both 4DImage and 4DImageAnalysis modules exposed in the trunk for the demo; he sees no problem in this *without* the Scipy modeling part exposed. Too challenging now to build python/numpy/scipy by default.
- Exposed FourDImage and FourDAnalysis modules in Slicer3/Modules/CMakelists.txt
- Changed the code in vtkKWPlotGraph.cxx: replaced finite() function is replaced code that conforms to ISO C++/2003 so it would compile on win32
- Changed some code in vtkFourDAnalysisLogic.cxx because there is no equivalent to the isnormal() function. Instead, just used the same def for finite()... but this will not test for subnormal floats.
- Had to move pyconfig.h to Slicer3-lib/python-build/Include so FourDAnalysis module could compile (couldn't find this file, included by <Python.h> for some reason (maybe something wrong with my python install on win32.
- Building in Debug mode rather than release creates python25_d.dll. The compile in genlib only builds python in release mode. Try changing the mode to ::$VTK_BUILD_TYPE and copy the python25_d.dll (as well as python25.dll to the python-build/Lib dir) in genlib.tcl...
- Something still looking to link with python25.dll (SlicerBaseGUIPython?) compiling python in debug mode does not generate the release libarary. So, next thing to try is running the python build twice, once in release, once in debug. Sloppy, but then we have both. made this change in genlib.tcl, and did the file copy of both the dlls.
- then, in subsequent call in getbuildtest.tcl, had to put a catch around the python lib file copy -- and add a copy of the debug library too. I think the result is that both debug and release python libs will be present; the particular slicer build will just link with the required python lib. There's certainly a better way, but I'll leave that to experts to improve if this appears to work.
- FourDAnalysis module still couldn't ind python25_d.lib. in Slicer_variables.tcl changed the way PYTHON_LIB is set:
set ::PYTHON_TEST_FILE $::PYTHON_BIN_DIR/PCbuild/python.exe
if { $::VTK_BUILD_TYPE == "Release" } {
set ::PYTHON_LIB $::PYTHON_BIN_DIR/PCbuild/python25.lib
} elseif { $::VTK_BUILD_TYPE == "Debug" } {
set ::PYTHON_LIB $::PYTHON_BIN_DIR/PCbuild/python25_d.lib
} elseif { $::VTK_BUILD_TYPE == "RelWithDebInfo" } {
set ::PYTHON_LIB $::PYTHON_BIN_DIR/PCbuild/python25.lib
} else {
puts stderr "Could not match build type to set python lib \"$::VTK_BUILD_TYPE\"."
}
puts "Set python lib = $::PYTHON_LIB"
set ::PYTHON_INCLUDE $::PYTHON_BIN_DIR/include
SCIPY for Slicer and python2.5 on win32
- Had to include scipy binary -- downloaded from here. Installed this library in my non-slicer python installation (the installer didn't give me option to put it anywhere else). Then did a copy of files to Slicer installation. This will take some debugging I think.
- C:/Python25/Lib/site-packages/scipy --> (path to slicer)/Slicer3-lib/python-build/Lib/site-packages/.
- C:/Python25/Lib/site-packages/README.txt --> (path to slicer)/Slicer3-lib/python-build/Lib/site-packages/.
- C:/Python25/Lib/site-packages/scipy-0.6.0-py2.5.egg-info --> (path to slicer)/Slicer3-lib/python-build/Lib/site-packages/.
- new files in C:/Python25/Lib/ --> (path to slicer)/Slicer3-lib/python-build/Lib/.
- new files in C:/Python25/Lib/encodings --> (path to slicer)/Slicer3-lib/python-build/Lib/encodings/.
now let's see how that works.
Started slicer, opened python console, typed import scipy and got the following error:
Traceback (most recent call last): File "<console>", line 1, in <module> File "C:\Python25\Lib\site-packages\scipy\__init__.py", line 25, in <module> from numpy import show_config as show_numpy_config ImportError: No module named numpy
Looks like numpy didn't install correctly for me using the genlib.tcl script.
NUMPY for Slicer and python2.5 on win32
So let's try grabbing the precompiled binary for numpy... from here.
- C:/Python25/Scripts/* --> (path to slicer)/Slicer3-lib/python-build/Lib/. (not sure about this one...)
- C:/Python25/Lib/site-packages/numpy --> (path to slicer)/Slicer3-lib/python-build/Lib/site-packages/.
- C:/Python25/Lib/site-packages/numpy-1.3.0-pyp2.5.egg-info --> (path to slicer)/Slicer3-lib/python-build/Lib/site-packages/.
Started slicer, opened python console, typed import numpy and import scipy and seemed to get the right behavior.
>>>>>>> SHOULD NOT RELY ON SCIPY FOR BROAD USE: IMPLEMENT IN C++
First experiment
- Load pre-treatment perfusion.
- Save scene PreTreatmentPerfusion.mrml
- Reload & see if volumeBundleNode reloads properly.
- BUG: looks like timeseries bundles did not get saved; FourDAnalysis module requires user to select a timeseries Bundle for plotting; none exists in the reloaded scene. Looks like FourDImage contains methods to serialize the mrml node, so maybe save was not good.
- Yes, BUG FIXED.
- Segment tumor & bloodpool
- in editor, select either Volname_Display0 or Volname_Display1 as src volume for label map -- Select Volname_Display1 (some label map out of range error occurs with Volname_Display0 for some reason... or maybe it's just an error enocuntered for the first label map created. don't know)
- do not use white as a label (plotting can't display white, so displays black label for white label (can be confusing)
- Save scene PreTreatmentPerfusionWithSegmentation.mrml
- Load just post-treatment perfusion
- Segment tumor & bloodpool
- Save scene PostTreatmentPerfusionWithSegmentation.mrml
- Reload PreTreatmentPerfusionWithSegmentation.mrml
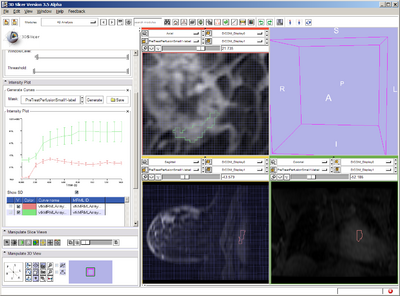
- Perform plotting on pre-treatment
- in FourDAnalysis, Mask = label map
- Perform Tofts Kinetic Model analysis on pre-treatment
- Select Tofts2 script (ignores plasma volume parameter, simpler than tofts)
- for AIF (c) arterial input function value (in multicolumn list), select the LUT number of the color of the blood pool
- for curve fitting, specify the color of the tumor label.
Not reliable scipy install on win32. module functions thru plotting, but curve fitting still not working cross-platform (only Mac).
Experiments to define best VOI
Pre-treatment VOI defined by careful tumor segmentation
Post-treatment scan appears to use different acquisition params from Pre-TX study. Suggested that Post-TX scan may not have used fat suppression. One suggestion is to carefully define VOI in Post-TX to include only scar, and to exclude anything that might look like fat. (will this not also potentially exclude tumor tissue?) Error bars too huge on Pre-Tx scan -- the analysis is weak. Try to exclude what we know to be necrotic tissue within the tumor.
Pre-treatment VOI defined as guided by functional imaging analysis slide from JHU
Using the following image provided by JHU to define a small VOI within Pre-TX tumor:
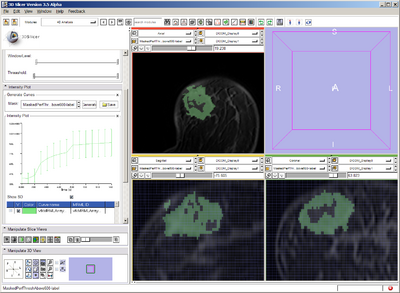
Pre-treatment VOI defined with strict intensity threshold (>1100)
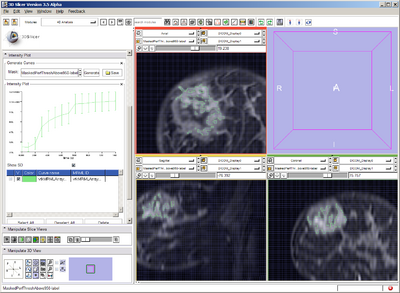
Pre-treatment VOI defined with various intensity threshold & mask
Thresholding to define 3 VOIs:
- 600 to max
- 950 to max
- 1100 to max
Masked each image with the segmentation of tumor to cull any voxels outside.
Approach & STATUS
- Need to compute VOI for tumor necrotic region and confirm flat plot
- Current status:
- FourDImage module not exposed to users in release or nightlies
- Only available in developer builds
- Need to refine and expose by November (RSNA deadline)
- FourDAnalysis module which implements Tofts Kinetic Model for curve fitting uses specialized components
- (scipy) doesn't build/install on all Slicer platforms
- (scipy) requires lots of hand-holding to install properly
- must convert module to C++ for cross-platform build (November deadline?)
- Need to implement idealized arterial input function if no AIF available in dataset (November deadline?)