Slicer3:DTMRI
From Slicer Wiki
Home < Slicer3:DTMRI
<flash>file=dti_glyphs.swf|width=800|height=600|quality=best|loop=true|play=true</flash>
Contents
- 1 Goals
- 2 Core infrastructure for DT-MRI processing and visualization, fiber processing and visualization
- 3 DT-MRI processing and visualization Modules
- 3.1 Volumes Module
- 3.2 Tractography Module
- 3.3 Tensor Estimation from DWI Module
- 3.4 Diffusion Tensor Scalar Measurements Module
- 3.5 Rician LMMSE Filter Module
- 3.6 Tractography ROI Seeding Module
- 3.7 Tractography Fiducial Seeding Module
- 3.8 Tractography ROI Select Module
- 3.9 Stochastic Tractography Filter
- 3.10 ROI Tract Filter
- 3.11 Generate Connectivity Map
- 4 Future development plans
- 5 Development Screenshots
- 6 Notes on general diffusion framework (ODF/2 tensor) support
Goals
- Development of the core infrastructure for DT-MRI processing and visualization.
- Development of the core infrastructure for fiber tracks processing and visualization.
- Integration of new and existing methods and algorithms for DT-MRI processing using the provided infrastructure.
- Porting of the current DT-MRI capabilities existing in Slicer 2.x
Core infrastructure for DT-MRI processing and visualization, fiber processing and visualization
- MRML nodes for data representation, storage, and display. MRML nodes store data and the state of Slicer modules.
- Data display and processing logic components.
- GUI components.
Data Representation
MRML nodes for different data representations involved in DTI analysis:
- Diffusion Weighted Images, represented as vtkImageData: vtkMRMLDiffusionWeightedVolumeNode.
- Diffusion Tensor Images, represented as vtkImageData: vtkMRMLDiffusionTensorVolumeNode.
- Fiber Bundles, represented as vtkPolyData: vtkMRMLFiberBundleNode.
Data Storage and I/O
- DWI and DTI I/O: NRRD is the format supported by Slicer3 for storing DWI and DTI images.
- NNRD reader/writer: vtkNRRDReader and vtkNRRDWriter.
- MRML Storage node: vtkMRMLNRRDStorageNode.
- Fiber I/O: vtkPolyData has been the format adopted for the description of fibers
- MRML Storage node: vtkMRMLFiberBundleStorageNode
Visualization/Display
MRML nodes for DWI, DTI, Tractography data visualization:
- Diffusion Weighted Images: vtkMRMLDiffusionWeightedVolumeDisplayNode.
- Diffusion Tensor Images: vtkMRMLDiffusionTensorVolumeDisplayNode, and vtkMRMLDiffusionTensorDisplayPropertiesNode.
- Fiber Bundles: vtkMRMLFiberBundleDisplayNode, vtkMRMLFiberBundleLineDisplayNode, vtkMRMLFiberBundleTubeDisplayNode, and vtkMRMLFiberBundleGlyphDisplayNode.
Displaying Logic
- Visualization pipelines for DTI, DWI volumes, and fiber bundles are incorporated into the corresponding display nodes.
- DWI volumes are displayed as separate components.
- DTI volumes are displayed as computed scalar properties (such as FA, Linear Measure, Color Orientation, etc.)
- Fiber bundles are displayed as lines, tubes, and glyphs with their own properties and colors
Diffusion Processing Toolbox
vtkTeem-library provides tools for:
- Tensor estimation
- Computation of scalar measurements from tensor fields
- Fast rendering of tensor fields using glyphs: line, box, ellipsoid
- Fiber Tracking using integration techniques
- Multiple ROI seeding and logic interconnections between ROIs
DT-MRI processing and visualization Modules
Volumes Module
- Volume module is capable of loading and saving DWI and DTI images in the NRRD file format
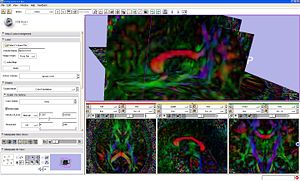
|
DTI volume slices colored by orientaion. |
- Volume Display GUI provides different display controls based on the type of the volume
- Built-in Slicer3 module
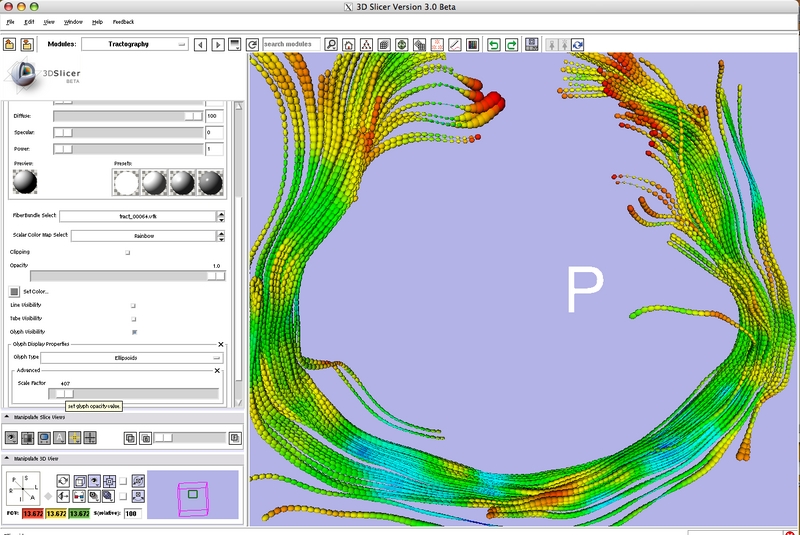
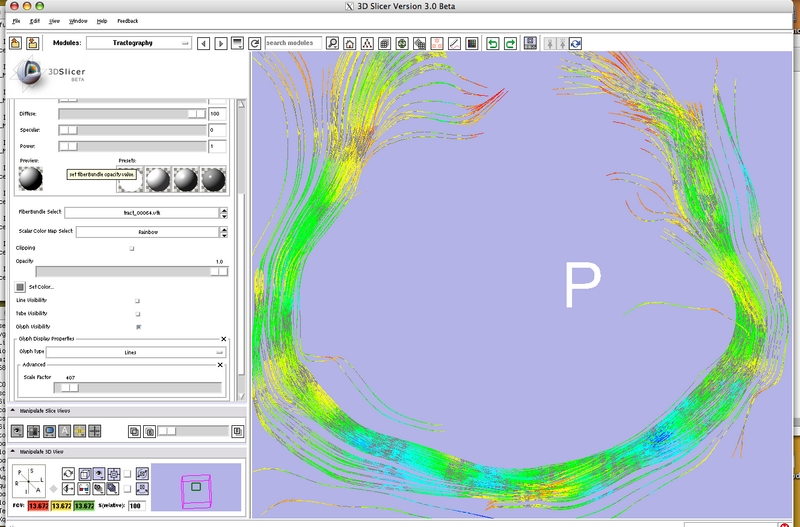
Tractography Module
- Tractography Display-Load-Save module is capable of loading and saving fiber tracts in the vtkPolyData file formats.
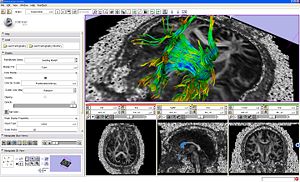
|
Fiber bundles displayed as lines and glyph ellipsoids. |
- Slicer Models module also can be used to load, save, display tracts as lines, but it does not provide tensor data display
- Built-in Slicer3 module
Tensor Estimation from DWI Module
- CLI Module: DiffusionTensorEstimation.
Teem currently provides a clean interface to do this estimation in a voxel by voxel fashion. Collaboration with Gordon Kindlmann for a vtk filter implementation that encapsulates the estimation process (vtkTeemEstimateDiffusionTensor).
Diffusion Tensor Scalar Measurements Module
- CLI Module: DiffusionTensorMathematics.
- Implemented in vtkTeem library vtkDiffusionTensorMathematics.
Rician LMMSE Filter Module
- CLI Module: dwiNoiseFilter.
- Filters a set of diffusion weighted images in the mean squared error sense using a Rician noise model. The noise parameter is automatically estimated.
- Contributed by Santiago Aja Fernandez and Marc Niethammer
- Additional Rician filtering module provided by Sylvain Gouttard et al
Tractography ROI Seeding Module
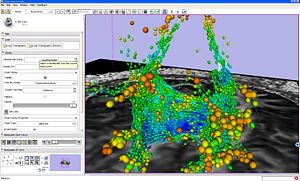
|
Seeding from ROI example. |
Tractography Fiducial Seeding Module
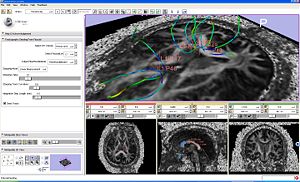
|
Seeding from fiducials example. |
Tractography ROI Select Module
- Select tracts passing or not passing through ROIs
- CLI Module: ROI Select.
Stochastic Tractography Filter
- CLI Module: StochasticTractographyFilter.
- Generates a map of connectivity probabilities from a DWI volume.).
- Contributed by Tri Ngo (tringo@gmail.com)
ROI Tract Filter
- CLI Module: ROITractFilter.
- Creates a new tract container containing only tracts which pass through the selected ROI's.
- Contributed by Tri Ngo (tringo@gmail.com)
Generate Connectivity Map
- CLI Module: GenerateConnectivityMap.
- Generates a volume where the value of each voxel is the number of fibers which pass through that voxel divided by the total number of sampled fibers. This value can been interpreted as the probability that a particular voxel is connected to the seed ROI by a fiber tract.
- Contributed by Tri Ngo (tringo@gmail.com)
Future development plans
- Fiber editing: enviroment for manually editing individual fibers/bundles, reassigning of fibers to bundles.
- Fiber Bundle Clustering (Core 1).
- Render glyphs in the 2D slice windows.
- Statistics along fiber tracts (Core 1).
- Quantitative measurement
- Tract-based
- Region of interest-based
- fMRI seeding
- Surgical planning
- DT-MRI segmentation/atlas creation: enviroment for segmentation of DT-MRI fields
- DT-MRI registration: enviroment for registration of DT-MRI fields (possibly via DWI registration -- work done at GE and presented in MICCAI '06).
- Tensor estimation using different methods, namely:
- Least Squares
- Weighted Least Squares
- Non-linear methods
- Maximum Likelihood approach
Development Screenshots
Notes on general diffusion framework (ODF/2 tensor) support
http://wiki.na-mic.org/Wiki/index.php/Slicer3:DTMRI:GeneralDiffusionFramework