Home < Modules:SpineSegmentation-Documentation-3.6Return to Slicer 3.6 Documentation
Gallery of New Features
Module Name
SpineSegmentation module for Slicer 3.6
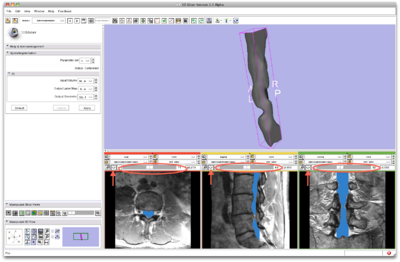
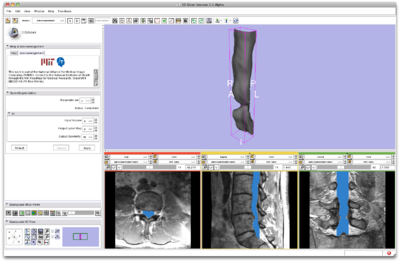
 Load volume into Slicer3.6 and select the SpineSegmentation module |
 Select input and output nodes and click Apply |
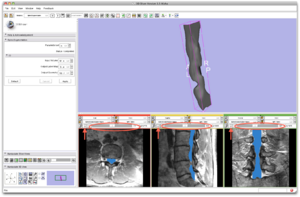 Processing takes approx. 3 minutes. The segmentation results are shown as a label map (blue) and a 3D model (3D view) |
General Information
Module Type & Category
Type: Interactive or CLI
Category: Segmentation
Authors, Collaborators & Contact
- Sylvain Jaume: MIT & logo, if desired
- Contact: Sylvain Jaume, sylvain at csail.mit.edu
Module Description
Image segmentation of the spinal cord and the cerebro-spinal fluid in T2-weighted MRI images. The SpineSegmentation module implements a model-based pattern recognition algorithm for fully automated segmentation.
Usage
Use Cases, Examples
This module is especially appropriate for these use cases:
Examples of the module in use:
Tutorials
Links to tutorials explaining how to use this module:
Tutorial and Usage
Below are the step-by-step instructions to load, process and save some sample data.
To download the sample data, you need to go to http://nitrc.org and create a username.
The data are located at
http://www.nitrc.org/plugins/scmsvn/viewcvs.php/Slicer3/Modules/SpineSegmentation/TestingData/?root=sylvainproject
|
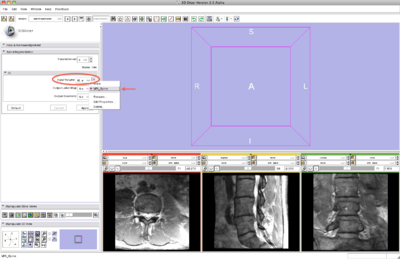
Step 1. Load the sample data
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
|
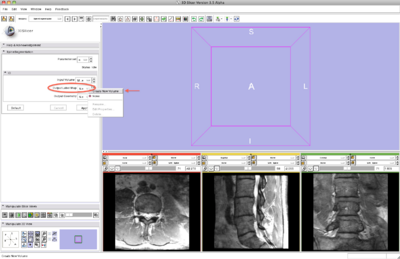
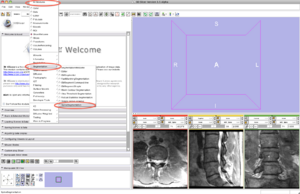
Step 2/9. Select the SpineSegmentation module
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
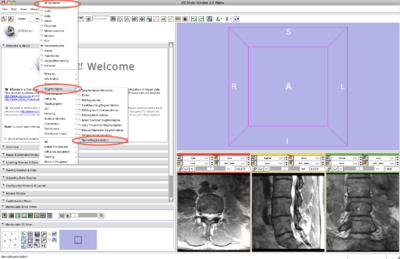 2. select SpineSegmentation module |
|
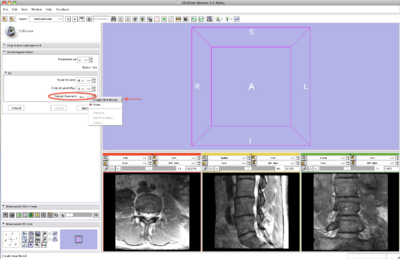
Step 3/9. Select the input image
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
|
Step 4/9. Create a Slicer node for the output image
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
|
Step 5/9. Create a Slicer node for the output model
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
|
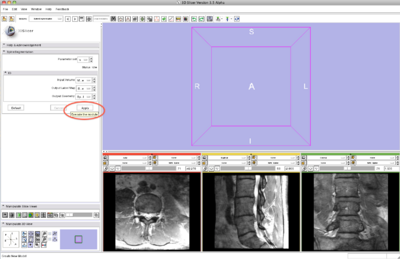
Step 6/9. Apply the segmentation algorithm
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
|
Step 7/9. Review segmentation result
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
|
Step 8/9. Save result image and model
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
|
Step 9/9. Find contact information for help and paper reference
- Input panel:
- Image input select the input image
- Image output create Slicer node for output image
- Model output create Slicer node for output model
- Command panel:
- Default reset input and output nodes to blank values
- Cancel cancel the execution of the algorithm
- Apply apply the segmentation algorithm (takes approx. 3 min)
|
|
Development
Notes from the Developer(s)
Algorithms used, library classes depended upon, use cases, etc.
Dependencies
Other modules or packages that are required for this module's use.
Tests
On the Dashboard, these tests verify that the module is working on various platforms:
Known bugs
Links to known bugs in the Slicer3 bug tracker
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Links to the module's source code:
Source code:
Doxygen documentation:
More Information
Acknowledgment
Include funding and other support here.
References
Publications related to this module go here. Links to pdfs would be useful.