Difference between revisions of "Modules:AtlasCreator:CongealingCLI"
| Line 5: | Line 5: | ||
=== Graphical User Interface in 3D Slicer === | === Graphical User Interface in 3D Slicer === | ||
| + | {| | ||
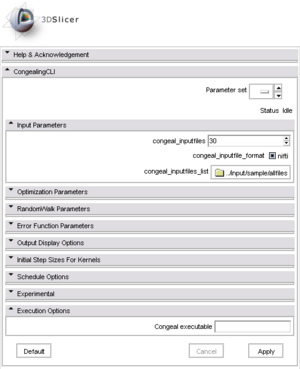
| + | |[[Image:CongealingCLIdefault.png|thumb|300px|The '''simple GUI''' mask, directly usable to create a default configuration file or to launch Congeal after specifying the path.]] | ||
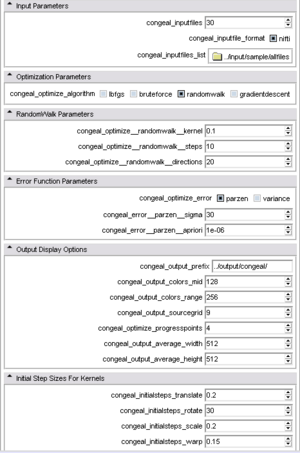
| + | |[[Image:CongealingCLItop.png|thumb|300px|The '''advanced GUI''' after the first couple of panels were expanded.]] | ||
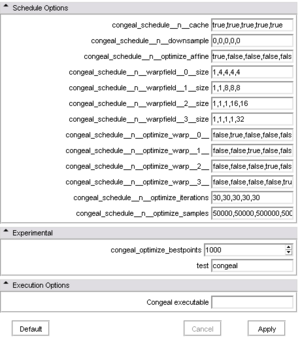
| + | |[[Image:CongealingCLIbottom.png|thumb|300px|The '''advanced GUI''' after the last couple of panels were expanded.]] | ||
| + | |} | ||
=== Command Line Interface === | === Command Line Interface === | ||
Revision as of 01:38, 7 April 2011
Home < Modules:AtlasCreator:CongealingCLIContents
CongealingCLI
CongealingCLI is a wrapper to access the Congealing Un-biased Groupwise Registration tool. It is now possible to generate a configuration file for Congealing by using a GUI or command line arguments. In fact, by not specifying any arguments and just running the wrapper, a default configuration file for congeal is generated.
Graphical User Interface in 3D Slicer
Command Line Interface
The option --help prints the possible command line arguments:
$ ./CongealingCLI --help
USAGE:
./CongealingCLI [--returnparameterfile <std::string>]
[--processinformationaddress <std::string>] [--xml]
[--echo] [--launch <std::string>] [--test
<std::string>] [--congeal_optimize_bestpoints <int>]
[--congeal_schedule__n__optimize_samples
<std::vector<int>>]
[--congeal_schedule__n__optimize_iterations
<std::vector<int>>]
[--congeal_schedule__n__optimize_warp__3__
<std::vector<std::string>>]
[--congeal_schedule__n__optimize_warp__2__
<std::vector<std::string>>]
[--congeal_schedule__n__optimize_warp__1__
<std::vector<std::string>>]
[--congeal_schedule__n__optimize_warp__0__
<std::vector<std::string>>]
[--congeal_schedule__n__warpfield__3__size
<std::vector<int>>]
[--congeal_schedule__n__warpfield__2__size
<std::vector<int>>]
[--congeal_schedule__n__warpfield__1__size
<std::vector<int>>]
[--congeal_schedule__n__warpfield__0__size
<std::vector<int>>]
[--congeal_schedule__n__optimize_affine
<std::vector<std::string>>]
[--congeal_schedule__n__downsample <std::vector<int>>]
[--congeal_schedule__n__cache
<std::vector<std::string>>]
[--congeal_initialsteps_warp <float>]
[--congeal_initialsteps_scale <float>]
[--congeal_initialsteps_rotate <float>]
[--congeal_initialsteps_translate <float>]
[--congeal_output_average_height <int>]
[--congeal_output_average_width <int>]
[--congeal_optimize_progresspoints <int>]
[--congeal_output_sourcegrid <int>]
[--congeal_output_colors_range <int>]
[--congeal_output_colors_mid <int>]
[--congeal_output_prefix <std::string>]
[--congeal_error__parzen__apriori <float>]
[--congeal_error__parzen__sigma <float>]
[--congeal_optimize_error <parzen|variance>]
[--congeal_optimize__randomwalk__directions <int>]
[--congeal_optimize__randomwalk__steps <int>]
[--congeal_optimize_randomwalk_kernel <float>]
[--congeal_optimize_algorithm <lbfgs|bruteforce
|randomwalk|gradientdescent>]
[--congeal_inputfiles_list <std::string>]
[--congeal_inputfile_format <nifti>]
[--congeal_inputfiles <int>] [--] [--version] [-h]
Where:
--returnparameterfile <std::string>
Filename in which to write simple return parameters (int, float,
int-vector, etc.) as opposed to bulk return parameters (image,
geometry, transform, measurement, table).
--processinformationaddress <std::string>
Address of a structure to store process information (progress, abort,
etc.). (default: 0)
--xml
Produce xml description of command line arguments (default: 0)
--echo
Echo the command line arguments (default: 0)
--launch <std::string>
The path to the congeal executable. Congeal will only be executed, if
this is set.
--test <std::string>
Currently unused. 'Must be congeal.' (default: congeal)
--congeal_optimize_bestpoints <int>
(default: 1000)
--congeal_schedule__n__optimize_samples <std::vector<int>>
number of samples to be compared in each transformed input volume.
Separated by comma for each schedule run. (default: 50000,50000,500000
,500000,500000)
--congeal_schedule__n__optimize_iterations <std::vector<int>>
number of optimzation iterations to be taken in the schedules.
Separated by comma for each schedule run. (default: 30,30,30,30,30)
--congeal_schedule__n__optimize_warp__3__ <std::vector<std::string>>
determines if the B-spline parameters for B-spline field 3 should be
optimized or left fixed. Separated by comma for each schedule run.
(default: false,false,false,false,true)
--congeal_schedule__n__optimize_warp__2__ <std::vector<std::string>>
determines if the B-spline parameters for B-spline field 2 should be
optimized or left fixed. Separated by comma for each schedule run.
(default: false,false,false,true,false)
--congeal_schedule__n__optimize_warp__1__ <std::vector<std::string>>
determines if the B-spline parameters for B-spline field 1 should be
optimized or left fixed. Separated by comma for each schedule run.
(default: false,false,true,false,false)
--congeal_schedule__n__optimize_warp__0__ <std::vector<std::string>>
determines if the B-spline parameters for B-spline field 0 should be
optimized or left fixed. Separated by comma for each schedule run.
(default: false,true,false,false,false)
--congeal_schedule__n__warpfield__3__size <std::vector<int>>
determines number of support points in each dimension of of B-Spline
mesh for field 3. Separated by comma for each schedule run. (default:
1,1,1,1,32)
--congeal_schedule__n__warpfield__2__size <std::vector<int>>
determines number of support points in each dimension of of B-Spline
mesh for field 2. Separated by comma for each schedule run. (default:
1,1,1,16,16)
--congeal_schedule__n__warpfield__1__size <std::vector<int>>
determines number of support points in each dimension of of B-Spline
mesh for field 1. Separated by comma for each schedule run. (default:
1,1,8,8,8)
--congeal_schedule__n__warpfield__0__size <std::vector<int>>
determines number of support points in each dimension of of B-Spline
mesh for field 0. Separated by comma for each schedule run. (default:
1,4,4,4,4)
--congeal_schedule__n__optimize_affine <std::vector<std::string>>
determines if affine parameters should be optimized or left fixed.
Separated by comma for each schedule run. (default: true,false,false
,false,false)
--congeal_schedule__n__downsample <std::vector<int>>
determines how many times the input data should be downsampled (by
factor of 2 in each dimension) prior to congealing. Separated by comma
for each schedule run. (default: 0,0,0,0,0)
--congeal_schedule__n__cache <std::vector<std::string>>
determines whether or not the schedules results can be retrieved from
the previous run. Separated by comma for each schedule run. (default:
true,true,true,true,true)
--congeal_initialsteps_warp <float>
relative scaling of warp control point displacement when computing
kernels and step sizes. Scale: Warp as fraction of control point's
region (default: 0.15)
--congeal_initialsteps_scale <float>
relative scaling of scaling parameters when computing kernels and step
sizes. Scale: Scale as fraction of image size (default: 0.2)
--congeal_initialsteps_rotate <float>
relative scaling of rotation parameters when computing kernels and
step sizes. Scale: rotation in degrees (default: 30)
--congeal_initialsteps_translate <float>
relative scaling of translation parameters when computing kernels and
step sizes. Scale: translation as fraction of image size (default:
0.2)
--congeal_output_average_height <int>
determines the height of the congealing average visualization
(default: 512)
--congeal_output_average_width <int>
determines the width of the congealing average visualization (default:
512)
--congeal_optimize_progresspoints <int>
determines how many output file sets will be generated during each
schedule (default: 4)
--congeal_output_sourcegrid <int>
determines how many of the transformed source values are shown in the
*-inputs* images (default: 9)
--congeal_output_colors_range <int>
color equalization slope. This value determines the relationship
between changes in data value and changes in output image gray value
(default: 256)
--congeal_output_colors_mid <int>
color equalization intercept. This value determines which data value
will be mapped to mid gray (default: 128)
--congeal_output_prefix <std::string>
string prepended to the filenames of the outputfiles. This value can
include an absolute or relative path, as well as a file prefix
(default: ../output/congeal/)
--congeal_error__parzen__apriori <float>
constant factor added to each Parzen estimate (default: 1e-06)
--congeal_error__parzen__sigma <float>
sigma of Gaussian used as kernel in Parzen density estimator. Measured
in voxel intensity (default: 30)
--congeal_optimize_error <parzen|variance>
selects the error metric to be used. parzen -- entropy estimate based
on Parzen density estimator. variance -- variance of voxel stack.
(default: parzen)
--congeal_optimize__randomwalk__directions <int>
number of beams to try during each iteration (default: 20)
--congeal_optimize__randomwalk__steps <int>
maximum number of steps to take along any beam (default: 10)
--congeal_optimize_randomwalk_kernel <float>
size of support to use for computing initial stepsize. This factor is
multiplied by *.initialsteps to establish a maximum step radius for
each dimensions (default: 0.1)
--congeal_optimize_algorithm <lbfgs|bruteforce|randomwalk
|gradientdescent>
determines the optimization algorithm used (default: randomwalk)
--congeal_inputfiles_list <std::string>
a path to a file containing a list of input data files. The list file
should contain one filename per line. Only congeal_inputfiles files
will be used as input unless congeal_inputfiles is set to '0' in which
case all the files in the list will be used (default:
../input/sample/allfiles)
--congeal_inputfile_format <nifti>
format of input files. Currently only 'nifti' is supported (default:
nifti)
--congeal_inputfiles <int>
number of input files to use. Use '0' for all files when used in
conjuctions with congeal_inputfiles.list (default: 30)
--, --ignore_rest
Ignores the rest of the labeled arguments following this flag.
--version
Displays version information and exits.
-h, --help
Displays usage information and exits.
Description: Generates a configuration file for the Congealing
Registration tool.
Author(s): Daniel Haehn and Kilian Pohl, University of
Pennsylvania
Acknowledgements: The research was funded by an ARRA supplement to NIH
NCRR (P41 RR13218).
Examples
The following commands are possible:
$ ./CongealingCLI Configuration file written to /var/tmp/tmp.0.YQNbrV
or
$ ./CongealingCLI --congeal_inputfiles 30 --congeal_inputfile_format nifti --congeal_inputfiles_list ../input/sample/allfiles --congeal_optimize_algorithm randomwalk --congeal_optimize_randomwalk_kernel 0.1 --congeal_optimize__randomwalk__steps 10 --congeal_optimize__randomwalk__directions 20 --congeal_optimize_error parzen --congeal_error__parzen__sigma 30 --congeal_error__parzen__apriori 1e-06 --congeal_output_prefix ../output/congeal/ --congeal_output_colors_mid 128 --congeal_output_colors_range 256 --congeal_output_sourcegrid 9 --congeal_optimize_progresspoints 4 --congeal_output_average_width 512 --congeal_output_average_height 512 --congeal_initialsteps_translate 0.2 --congeal_initialsteps_rotate 30 --congeal_initialsteps_scale 0.2 --congeal_initialsteps_warp 0.15 --congeal_schedule__n__cache true,true,true,true,true --congeal_schedule__n__downsample 0,0,0,0,0 --congeal_schedule__n__optimize_affine true,false,false,false,false --congeal_schedule__n__warpfield__0__size 1,4,4,4,4 --congeal_schedule__n__warpfield__1__size 1,1,8,8,8 --congeal_schedule__n__warpfield__2__size 1,1,1,16,16 --congeal_schedule__n__warpfield__3__size 1,1,1,1,32 --congeal_schedule__n__optimize_warp__0__ false,true,false,false,false --congeal_schedule__n__optimize_warp__1__ false,false,true,false,false --congeal_schedule__n__optimize_warp__2__ false,false,false,true,false --congeal_schedule__n__optimize_warp__3__ false,false,false,false,true --congeal_schedule__n__optimize_iterations 30,30,30,30,30 --congeal_schedule__n__optimize_samples 50000,50000,500000,500000,500000 --congeal_optimize_bestpoints 1000 --test congeal Configuration file written to /var/tmp/tmp.0.YQNbrV
and the output is the same for both (thanks to the default values, they also equal a press on the Apply button in the GUI without any changes to the input fields):
$ cat /var/tmp/tmp.0.YQNbrV
# experimental
congeal.optimize.bestpoints 1000
test congeal
# input
congeal.inputfiles 30
congeal.inputfile.format nifti
congeal.inputfiles.list ../input/sample/allfiles
# optimization
congeal.optimize.algorithm randomwalk
# randomwalk
congeal.optimize[randomwalk].kernel 0.1
congeal.optimize[randomwalk].steps 10
congeal.optimize[randomwalk].directions 20
# error function
congeal.optimize.error parzen
# parzen error function
congeal.error[parzen].sigma 30
congeal.error[parzen].apriori 1e-06
# output
congeal.output.prefix ../output/congeal/
congeal.output.colors.mid 128
congeal.output.colors.range 256
congeal.output.sourcegrid 9
congeal.optimize.progresspoints 4
congeal.output.average.width 512
congeal.output.average.height 512
# initial steps
congeal.initialsteps.translate 0.2
congeal.initialsteps.rotate 30
congeal.initialsteps.scale 0.2
congeal.initialsteps.warp 0.15
# schedules
n -1
congeal.schedule[{++n}].cache true
congeal.schedule[{$n}].downsample 0
congeal.schedule[{$n}].optimize.affine true
congeal.schedule[{$n}].warpfield[0].size 1
congeal.schedule[{$n}].warpfield[1].size 1
congeal.schedule[{$n}].warpfield[2].size 1
congeal.schedule[{$n}].warpfield[3].size 1
congeal.schedule[{$n}].optimize.warp[0] false
congeal.schedule[{$n}].optimize.warp[1] false
congeal.schedule[{$n}].optimize.warp[2] false
congeal.schedule[{$n}].optimize.warp[3] false
congeal.schedule[{$n}].optimize.iterations 30
congeal.schedule[{$n}].optimize.samples 50000
congeal.schedule[{++n}].cache true
congeal.schedule[{$n}].downsample 0
congeal.schedule[{$n}].optimize.affine false
congeal.schedule[{$n}].warpfield[0].size 4
congeal.schedule[{$n}].warpfield[1].size 1
congeal.schedule[{$n}].warpfield[2].size 1
congeal.schedule[{$n}].warpfield[3].size 1
congeal.schedule[{$n}].optimize.warp[0] true
congeal.schedule[{$n}].optimize.warp[1] false
congeal.schedule[{$n}].optimize.warp[2] false
congeal.schedule[{$n}].optimize.warp[3] false
congeal.schedule[{$n}].optimize.iterations 30
congeal.schedule[{$n}].optimize.samples 50000
congeal.schedule[{++n}].cache true
congeal.schedule[{$n}].downsample 0
congeal.schedule[{$n}].optimize.affine false
congeal.schedule[{$n}].warpfield[0].size 4
congeal.schedule[{$n}].warpfield[1].size 8
congeal.schedule[{$n}].warpfield[2].size 1
congeal.schedule[{$n}].warpfield[3].size 1
congeal.schedule[{$n}].optimize.warp[0] false
congeal.schedule[{$n}].optimize.warp[1] true
congeal.schedule[{$n}].optimize.warp[2] false
congeal.schedule[{$n}].optimize.warp[3] false
congeal.schedule[{$n}].optimize.iterations 30
congeal.schedule[{$n}].optimize.samples 500000
congeal.schedule[{++n}].cache true
congeal.schedule[{$n}].downsample 0
congeal.schedule[{$n}].optimize.affine false
congeal.schedule[{$n}].warpfield[0].size 4
congeal.schedule[{$n}].warpfield[1].size 8
congeal.schedule[{$n}].warpfield[2].size 16
congeal.schedule[{$n}].warpfield[3].size 1
congeal.schedule[{$n}].optimize.warp[0] false
congeal.schedule[{$n}].optimize.warp[1] false
congeal.schedule[{$n}].optimize.warp[2] true
congeal.schedule[{$n}].optimize.warp[3] false
congeal.schedule[{$n}].optimize.iterations 30
congeal.schedule[{$n}].optimize.samples 500000
congeal.schedule[{++n}].cache true
congeal.schedule[{$n}].downsample 0
congeal.schedule[{$n}].optimize.affine false
congeal.schedule[{$n}].warpfield[0].size 4
congeal.schedule[{$n}].warpfield[1].size 8
congeal.schedule[{$n}].warpfield[2].size 16
congeal.schedule[{$n}].warpfield[3].size 32
congeal.schedule[{$n}].optimize.warp[0] false
congeal.schedule[{$n}].optimize.warp[1] false
congeal.schedule[{$n}].optimize.warp[2] false
congeal.schedule[{$n}].optimize.warp[3] true
congeal.schedule[{$n}].optimize.iterations 30
congeal.schedule[{$n}].optimize.samples 500000
congeal.schedules {++n}
Using --launch
When adding a --launch PATH_TO_CONGEAL_EXEC, the congeal executable gets launched rather than printing the path to the generated configuration file.
For example
$ ./CongealingCLI --launch congeal
starts the congeal executable with the configuration file shown above.