Difference between revisions of "Modules:ARCTIC-Documentation-3.6"
| Line 88: | Line 88: | ||
===Dependencies=== | ===Dependencies=== | ||
| − | Slicer3 modules: RegisterImages | + | * Slicer3 modules: |
| + | ** RegisterImages | ||
| + | ** ResampleScalarVectorDWIVolume | ||
| + | * Atlas: | ||
| + | ** | ||
| + | * Libraries: | ||
===Tests=== | ===Tests=== | ||
Revision as of 21:40, 31 March 2010
Home < Modules:ARCTIC-Documentation-3.6Return to Slicer 3.6 Documentation
Module Name
MyModule
General Information
Module Type & Category
Type: CLI
Category: Pipeline
Authors, Collaborators & Contact
- Author1: Cedric Mathieu, UNC-Chapel Hill
- Contributor1: Clement Vachet, UNC-Chapel Hill
- Contributor2: Martin Styner, UNC-Chapel Hill
- Contributor3: Heather Cody Hazlett, UNC-Chapel Hill
- Contact: Clement Vachet, cvachet[at]email[dot]unc[dot]edu
Module Description
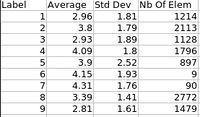
ARCTIC (Automatic Regional Cortical ThICkness) is an end-to-end application developped at UNC-Chapel Hill allowing individual analysis of cortical thickness. This cross-platform tool can be run within Slicer3 as an external module, or directly as a command line.
Usage
Use Cases, Examples
This module is especially appropriate when one wants to perform individual regional cortical thickness analysis. ARCTIC allows efficient QC via precomputed 3D-Slicer scenes.
Tutorials
- Tutorials:
- Tutorial Dataset
Quick Tour of Features and Use
A list panels in the interface, their features, what they mean, and how to use them. For instance:
|
Development
Notes from the Developer(s)
Algorithms used, library classes depended upon, use cases, etc.
Dependencies
- Slicer3 modules:
- RegisterImages
- ResampleScalarVectorDWIVolume
- Atlas:
- Libraries:
Tests
On the Dashboard, these tests verify that the module is working on various platforms:
- MyModuleTest1 MyModuleTest1.cxx
- MyModuleTest2 MyModuleTest2.cxx
Known bugs
Links to known bugs in the Slicer3 bug tracker
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Download release source code via svn:
More Information
Acknowledgment
This work is part of the National Alliance for Medical Image Computing (NAMIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 EB005149. Information on the National Centers for Biomedical Computing can be obtained from http://nihroadmap.nih.gov/bioinformatics
References
Publications related to this module go here. Links to pdfs would be useful.