Documentation:Nightly:Registration:RegistrationLibrary:RegLib C06
Contents
Slicer Registration Library Case #6: Breast MRI Cancer Treatment
Input
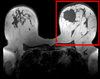
|
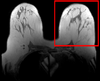
| |
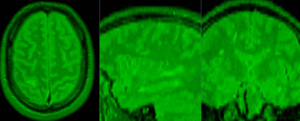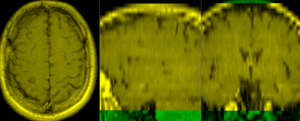
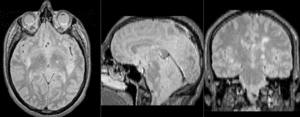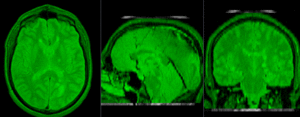
| exam 1: baseline Pre Rx | exam 1: follow-up Post Rx |
Description
This is an example of serial/logitudinal imaging for change assessment. We have a pre- and post-treatment MRI. To assess the treatment effect we seek to register the postRx scan to the baseline. Because of the strong changes in image content and contrast between the two exams, some additional work is necessary to obtain a good registration: 1) we will register only the left breast containing the tumor, i.e. we will crop the image to a much smaller FOV, while also obtaining a more isotropic resolution; 2) we have strong intensity bias in the images (originating from MR receiver coil sensitivity) that needs to be corrected first, otherwise it will negatively bias the registration; and 3) we will generate masks that include the breast but not the tumor, to ensure the nonrigid portion of the registration will not try to compensate for the tumor change.
Modules used
- Crop Volume
- N4ITKBiasFieldCorrection (filter correcting for local intensity inhomogeneity)
- Foregroud masking (BRAINS) (quick mask generation)
- Editor
- General Registration (BRAINS)
Download (from NAMIC MIDAS)
Why 2 sets of files? The "input data" mrb includes only the unregistered data to try the method yourself from start to finish. The full dataset includes intermediate files and results (transforms, resampled images etc.). If you use the full dataset we recommend to choose different names for the images/results you create yourself to distinguish the old data from the new one you generated yourself.
- RegLib_C06.mrb: input data only, use this to run the tutorial from the start (Slicer mrb file. 19 MB).
- RegLib_C06_full.mrb: includes raw data + all solutions and intermediate files, use to browse/verify (Slicer mrb file. 19 MB).
Keywords
MRI, breast, intra-subject, breast cancer, treatment effect
Video Screencasts
- Movie/screencast showing importing and cropping the image volume
- Movie/screencast showing the bias (intensity inhomogeneity) correction step
- Movie/screencast showing building the masks for registration
- Movie/screencast showing affine + BSpline registration
Procedure
- Crop:Extract left breast image of PreRx and PostRx scan
- Open the Crop Volume module (under Converters menu)
- Input volume: "PreRx"
- Input ROI: "Create & rename new annotation ROI", rename to "PreRx_CropROI" or similar
- Isotropic output voxel: yes (checkbox)
- Input spacing scaling constant:increase to 3 (the original image has a 0.4 x 0.4 x 5 mm voxel size. Isotropic resolution is highly desirable for registration, but the full resolution would yield a 784^3 image volume that would use too much memory for purposes of this demonstration).
- Interpolator: "Linear" (radio button)
- you should see a set of colored dots/beads in at least one of the slice views. If not scroll a bit to locate them. The beads represent sides and corners of an ROI box. Drag the corners of the box to completely enclose the left breast (right side of image).
- click on the Crop! button
- go to the Data module
- several new nodes were created: look for the "PreRx_subvolume-scale_3" entry, double click and rename to "PreRx_Left" or "PreRx_crop" or similar.
- Turn off the visibility of the ROI box just created: click on the eye icon next to the "PreRx_CropROI" node. (we do this to more easily distinguish the next ROI box created for the second volume below)
- repeat the same for the "PostRx" image. Since this image is smaller, choose 2 for the Input spacing scaling constant (vs. 3 above).
- Save intermediate results. You should have now 2 new volumes at ~ 1.3 mm isotropic resolution.
- Bias Correction: Correct for local intensity inhomogeneities
- Open the N4ITK MRI Bias Correction module (under Filters menu)
- Input Image: "PreRx_left"
- Mask Image: none
- Output Volume: create & rename new: "PreRx_left_n4"
- Number of iterations: reduce to 100,80,60
- Shrink factor: reduce to 3
- leave all parameters at defaults
- Apply.
- repeat for the Post_Rx image. Same settings
- save intermediate results
- Open the N4ITK MRI Bias Correction module (under Filters menu)
- Build Masks:
- open the Foregroud masking (BRAINS) module (under Segmentation:Specialized)
- Input Image Volume: PreRx_left_n4
- Output Mask: create & rename new: "PreRx_mask"
- Apply
- (alternatively you can also use the threshold tool in the Editor to obtain a quick mask)
- we need to exclude the tumor from the mask, to prevent the registration from trying to match that region. Much should already be excluded from the above segmentation, but we also need to dilate the mask a bit to extend beyon the edge and include some of the background, which will fill in the tumor space again. Hence we make some manual edits to delete the tumor area from the mask:
- Go to the Editor module
- Click Apply on the popup do confirm colormap selection (choice does not matter)
- Master Volume: PreRx_left
- Merge Volume: PreRx_mask (generated above)
- click on the Brush Icon ("PaintEffect")
- Label: 0 (black)
- increase radius to ~ 10-15 mm.
- navigate to an axial slice showing the tumor
- click & drag left mouse to apply, then use arrow keys to move to the next slice.
- repeat for the Post_Rx_mask
- save intermediate results
- open the Foregroud masking (BRAINS) module (under Segmentation:Specialized)
- Affine pre-Registration:
- open the * General Registration (BRAINS) module
- Fixed Image Volume: PreRx_left_n4
- Moving Image Volume: PostRx_left_n4
- Output Settings:
- Slicer BSpline Transform": none
- Slicer Linear Transform: create & rename new transform, rename to "Xf1_Affine"
- Output Image Volume: none
- Initialize Transform Mode: select Use Moments Align.
- Registration Phases: check boxes for Rigid , Rigid+Scale and Affine
- Main Parameters : Number Of Samples: 200,000
- Leave all other settings at default
- Apply
- this should generate a first alignment. note that we need not generate an output volume, because this is an intermediate result, and the BRAINS registration tool will automatically place the moving volume inside the result transform.
- Place "PreRx_left_n4" in the background and "PostRx_left_n4" in the foreground. Use fade slider to evaluate the result.
- open the * General Registration (BRAINS) module
- Nonrigid final Registration:
- open the General Registration (BRAINS) module
- Fixed Image Volume: PreRx_left_n4
- Moving Image Volume: PostRx_left_n4
- Initialization transform: "Xf1_Affine
- Initialize Transform Mode: de-select Use Moments Align, select None
- Output Settings:
- Slicer BSpline Transform": create & rename new transform, rename to "Xf2_BSpline"
- Slicer Linear Transform: none
- Output Image Volume: create & rename new transform, rename to "PostRx_left_Xf2" (this time we must request an output volume, since the nonrigid deformations cannot be visualized on the "fly.
- Registration Phases: uncheck rigid & affine boxes, check box for BSpline only
- Main Parameters:
- Number Of Samples: 300,000
- B-Spline Grid Size: 7,7,5
- Mask Option: select ROI button
- ROI Masking input fixed: select "PreRx_mask" generated in Phase 3 above
- ROI Masking input moving: select "PostRx_mask" generated in Phase 3 above
- Leave all other settings at default
- click: Apply
- open the General Registration (BRAINS) module