Difference between revisions of "Documentation/Nightly/Training"
m (Text replacement - "https?:\/\/wiki.slicer.org\/slicerWiki\/index.php\/([^ ]+) " to "https://www.slicer.org/wiki/$1") |
|||
| (4 intermediate revisions by 2 users not shown) | |||
| Line 104: | Line 104: | ||
[[Image:HelloPythonTutorial.png|right|250px|]] | [[Image:HelloPythonTutorial.png|right|250px|]] | ||
|} | |} | ||
| + | |||
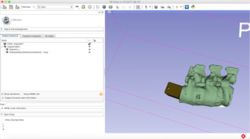
| + | == PerkLab's Slicer bootcamp training materials == | ||
| + | {|width="100%" | ||
| + | | | ||
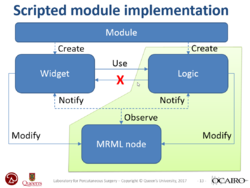
| + | *The [http://perk.cs.queensu.ca/ Laboratory for Percutaneous Surgery at Queen's University] has made available training material of its internal yearly bootcamp, covering topics, such as 3D Slicer overview, basic visualization, segmentation, registration, scripting and module development, surgical navigation, DICOM, reproducible medical image computing research methodology, version control, and research project management. | ||
| + | ** [https://github.com/PerkLab/PerkLabBootcamp/blob/master/Doc/day3_2_SlicerProgramming.pptx?raw=true Scripting and module development tutorial] | ||
| + | ** [https://github.com/PerkLab/PerkLabBootcamp/tree/master/Doc All other tutorials] | ||
| + | *Author: Andras Lasso, Csaba Pinter, Tamas Ungi, Csaba Pinter, Matthew Holden, Kyle Sunderland | ||
| + | *Audience: Developers, Users | ||
| + | *Based on: 3D Slicer version 4.10 | ||
| + | |align="right"| | ||
| + | [[Image:PerkLabSlicerProgrammingTutorial.png|right|250px|]] | ||
| + | |} | ||
| + | |||
| + | == Slicer script repository == | ||
For additional Python scripts examples, please visit the [[Documentation/{{documentation/version}}/ScriptRepository|Script Repository page]] | For additional Python scripts examples, please visit the [[Documentation/{{documentation/version}}/ScriptRepository|Script Repository page]] | ||
| Line 124: | Line 139: | ||
| | | | ||
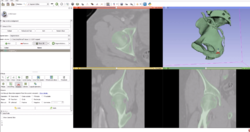
* Segmentation for 3D printing: shows how to use the Segment Editor module for combining CAD designed parts with patient-specific models. | * Segmentation for 3D printing: shows how to use the Segment Editor module for combining CAD designed parts with patient-specific models. | ||
| + | ** '''[https://discourse.slicer.org/t/new-video-tutorial-for-segment-editor-lumbar-spine-segmentation-for-3d-printing/700 Video tutorial]'''. Author: Hillary Lia. | ||
** '''[[Documentation/{{documentation/version}}/Training#Segmentation_for_3D_printing|Segmentation for 3D printing Step-by-step tutorial]]'''. Author: Csaba Pinter, MSc | ** '''[[Documentation/{{documentation/version}}/Training#Segmentation_for_3D_printing|Segmentation for 3D printing Step-by-step tutorial]]'''. Author: Csaba Pinter, MSc | ||
** Audience: Users and developers interested in segmentation and 3D printing | ** Audience: Users and developers interested in segmentation and 3D printing | ||
| + | ** Dataset: [[:File:BasePiece.zip|Phantom base STL model]] Source: [http://perk-software.cs.queensu.ca/plus/doc/nightly/modelcatalog/ PerkLab]. | ||
** Based on: 3D Slicer version 4.7 | ** Based on: 3D Slicer version 4.7 | ||
|align="right"|[[Image:20170717_3DPrintingTutorialYoutube.PNG|280px]] | |align="right"|[[Image:20170717_3DPrintingTutorialYoutube.PNG|280px]] | ||
| Line 133: | Line 150: | ||
** Author: Andras Lasso, PhD | ** Author: Andras Lasso, PhD | ||
** Audience: Users who need to segment heart structures, for example for visualization, quantification, or simulation. | ** Audience: Users who need to segment heart structures, for example for visualization, quantification, or simulation. | ||
| − | ** | + | ** Sample data set: http://slicer.kitware.com/midas3/download/bitstream/738905/CTA-cardio2.nrrd |
** Based on: 3D Slicer version 4.8 | ** Based on: 3D Slicer version 4.8 | ||
|align="right"|[[Image:WholeHeartSegYoutube.png|280px]] | |align="right"|[[Image:WholeHeartSegYoutube.png|280px]] | ||
| + | |--- | ||
| + | | | ||
| + | * '''[https://youtu.be/0at15gjk-Ns Video tutorial: Femur and pelvis segmentation from CT]''' shows how to use the Segment Editor module for segmenting pelvis and femur from CT volumes. | ||
| + | ** Author: Andras Lasso, PhD | ||
| + | ** Audience: Users who need to segment bones in CT images for visualization, quantification, or simulation. | ||
| + | ** Sample data set: https://wiki.cancerimagingarchive.net/display/Public/TCGA-PRAD (Subject TCGA-VP-A878) | ||
| + | ** Based on: 3D Slicer version 4.8 | ||
| + | |align="right"|[[Image:FemurSegmentationYoutube.png|280px]] | ||
| + | |--- | ||
| + | | | ||
| + | * [https://lassoan.github.io/SlicerSegmentationRecipes/ Slicer Segmentation Recipes] | ||
|} | |} | ||
| Line 159: | Line 187: | ||
:Author: Dominik Meier, Ph.D. | :Author: Dominik Meier, Ph.D. | ||
:Audience: users interested learning/applying Slicer image registration technology | :Audience: users interested learning/applying Slicer image registration technology | ||
| − | |align="right"|[[Image:RegLib_table.png|250px|link= | + | |align="right"|[[Image:RegLib_table.png|250px|link=https://www.slicer.org/wiki/Documentation/{{documentation/version}}/Registration/RegistrationLibrary]]|} |
| − | |} | ||
=Slicer Extensions= | =Slicer Extensions= | ||
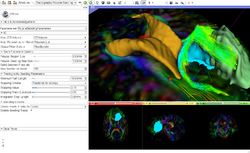
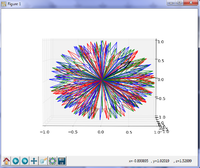
==Slicer4 Diffusion Tensor Imaging Tutorial == | ==Slicer4 Diffusion Tensor Imaging Tutorial == | ||
| Line 179: | Line 206: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
| − | + | *The [https://spujol.github.io/NeurosurgicalPlanningTutorial/ Neurosurgical Planning tutorial] course guides end-users through the generation of fiber tracts in the vicinity of a tumor. | |
| − | *The [ | ||
*Author: Sonia Pujol, Ph.D., Ron Kikinis, M.D. | *Author: Sonia Pujol, Ph.D., Ron Kikinis, M.D. | ||
| − | *Audience: | + | *Audience: Clinicians and Clinical Researchers |
| − | *Modules: | + | *Modules: Segment Editor, Tractography |
| − | *Based on | + | *Based on 3D Slicer version 4.10 |
| − | *The [[Media:WhiteMatterExplorationData.zip| White Matter Exploration dataset]] contains a Diffusion Weighted Imaging scan of | + | *The [[Media:WhiteMatterExplorationData.zip| White Matter Exploration dataset]] contains a Diffusion Weighted Imaging scan of a brain tumor patient. |
|align="right"| | |align="right"| | ||
[[Image:NeurosurgicalPlanningTutorial.png|right|250px|link=http://vimeo.com/67336069]] | [[Image:NeurosurgicalPlanningTutorial.png|right|250px|link=http://vimeo.com/67336069]] | ||
|} | |} | ||
| − | |||
| − | |||
==Slicer4 Quantitative Imaging tutorial== | ==Slicer4 Quantitative Imaging tutorial== | ||
Revision as of 14:54, 27 November 2019
Home < Documentation < Nightly < Training
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction: Slicer Nightly Tutorials
- This page contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment (version Nightly release) to accomplish certain tasks.
- For tutorials for other versions of Slicer, please visit the Slicer training portal.
- For "reference manual" style documentation, please visit the Slicer Nightly documentation page
- For questions related to the Slicer4 Training Compendium, please send an e-mail to Sonia Pujol, Ph.D., Director of Training of 3D Slicer.
- Some of these tutorials are based on older releases of 3D Slicer and are being upgraded to Slicer4.10. The concepts are still useful but some interface elements and features may be different in updated versions.
Contents
Quick Start Guide
Downloading and Installing Slicer
|
General Introduction
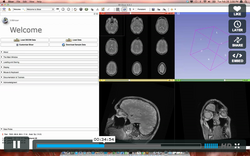
Slicer Welcome Tutorial
|
Slicer4Minute Tutorial
|
3D Visualization
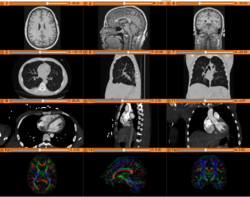
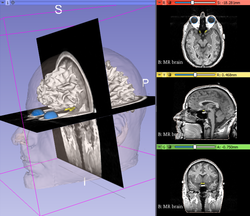
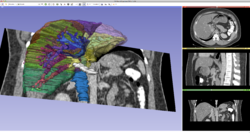
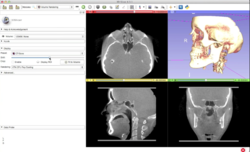
Slicer4 Data Loading and 3D Visualization
|
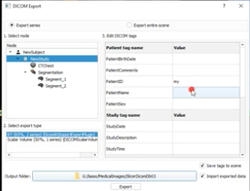
Slicer4 3D Visualization of DICOM images for Radiology Applications
|
Tutorials for software developers
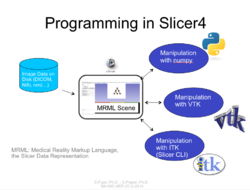
Slicer4 Programming Tutorial
|
PerkLab's Slicer bootcamp training materials
|
Slicer script repository
For additional Python scripts examples, please visit the Script Repository page
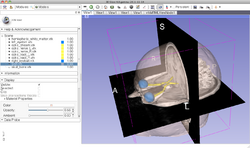
Developing and contributing extensions for 3D Slicer
|
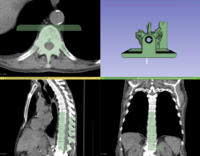
Segmentation
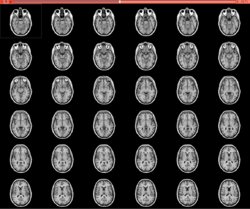
Slicer4 Image Segmentation
|
|
|
|
|
|
Registration
Slicer4 Image Registration
|

|
- Based on: 3D Slicer version 4.8
Slicer Registration Case Library
|
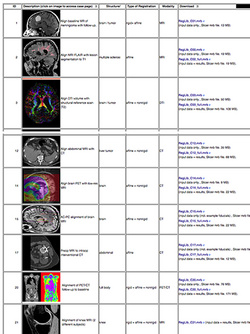 |} |}
Slicer ExtensionsSlicer4 Diffusion Tensor Imaging Tutorial
Slicer4 Neurosurgical Planning Tutorial
Slicer4 Quantitative Imaging tutorial
Slicer4 IGT
Slicer4 Radiation Therapy Tutorial
Slicer Pathology
SPHARM-PDM
Fiber Bundle Volume Measurement
3D Slicer version 4.7 Tutorial ContestFor previous editions of the contest, please visit the 3D Slicer Tutorial Contests page Segmentation for 3D printing
Slicer Pathology
Simple Python Tool for Quality Control of DWI data
SPHARM-PDM
Integration of Robot Operating System (ROS) and 3D Slicer using OpenIGTLink
Fiber Bundle Volume Measurement
YouTube videosAdditional non-curated videos-based demonstrations using 3D Slicer are accessible on YouTube. Teams Contributions
International resourcesResources in Chinese
Resources in German
Murat Maga's blog posts about using 3D Slicer for biologyUsing the (legacy) EditorFast GrowCut
Use case: Slicer in paleontologyThis set of tutorials about the use of slicer in paleontology is very well written and provides step-by-step instructions. Even though it covers slicer version 3.4, many of the concepts and techniques have applicability to the new version and to any 3D imaging field:
|