Difference between revisions of "Documentation/Nightly/Modules/Colors"
(Prepend documentation/versioncheck template. See http://na-mic.org/Mantis/view.php?id=2887) |
|||
| Line 8: | Line 8: | ||
{{documentation/{{documentation/version}}/module-introduction-start|{{documentation/modulename}}}} | {{documentation/{{documentation/version}}/module-introduction-start|{{documentation/modulename}}}} | ||
{{documentation/{{documentation/version}}/module-introduction-row}} | {{documentation/{{documentation/version}}/module-introduction-row}} | ||
| − | :'''Author(s)/Contributor(s):''' Nicole Aucoin (SPL, BWH) | + | :'''Author(s)/Contributor(s):''' Nicole Aucoin (SPL, BWH), Jim Miller (GE), Julien Finet (Kitware Inc.) |
: '''Acknowledgements:''' This work is part of the [http://www.na-mic.org/ National Alliance for Medical Image Computing] (NA-MIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 EB005149.<br> | : '''Acknowledgements:''' This work is part of the [http://www.na-mic.org/ National Alliance for Medical Image Computing] (NA-MIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 EB005149.<br> | ||
: '''Contact:''' Nicole Aucoin, <email>nicole@bwh.harvard.edu</email><br> | : '''Contact:''' Nicole Aucoin, <email>nicole@bwh.harvard.edu</email><br> | ||
| Line 28: | Line 28: | ||
The Colors Module manages color look up tables. | The Colors Module manages color look up tables. | ||
| − | + | Color look up tables are used by mappers to translate between an integer and a colour value for display of models and volumes. | |
Slicer supports three kinds of tables: | Slicer supports three kinds of tables: | ||
# Continuous scales, like the greyscale table. | # Continuous scales, like the greyscale table. | ||
| − | # Parametric tables, defined by an equation, such as the | + | # Parametric tables, defined by an equation, such as the fMRIPA table. |
# Discrete tables, such as those read in from a file. | # Discrete tables, such as those read in from a file. | ||
| Line 38: | Line 38: | ||
You can specify a directory from which to read color files using the Edit -> Application Settings window, Module Settings frame, in the User defined color file paths section. --> | You can specify a directory from which to read color files using the Edit -> Application Settings window, Module Settings frame, in the User defined color file paths section. --> | ||
| − | You can create a duplicate of a color table to allow editing the names, values and color by clicking on the | + | You can create a duplicate of a color table to allow editing the names, values and color by clicking on the [[image:Slicer43-Colors-FolderPlus.jpeg]] icon next to the drop down meny of color table nodes. You can then save the new color table via the File -> Save interface. |
You can load a color table file from the File -> Add Data dialog. | You can load a color table file from the File -> Add Data dialog. | ||
| Line 57: | Line 57: | ||
===Custom LUTs=== | ===Custom LUTs=== | ||
| − | You can create custom LUTs by [[Documentation/{{documentation/version}}/SlicerApplication/LookupTables#Custom_LUTs|creating a table with the colors on the wiki]]. | + | You can create custom LUTs by [[Documentation/{{documentation/version}}/SlicerApplication/LookupTables#Custom_LUTs|creating a table with the colors on the wiki]], saving to file and then loading them into Slicer. |
===Categories=== | ===Categories=== | ||
| Line 75: | Line 75: | ||
** [[image:DiscretefMRIPA.png]] fMRIPA: A small fMRI positive activation scale going from red to yellow from 0-19, useful for displaying functional MRI volumes when don't need the blue of the fMRI scale. | ** [[image:DiscretefMRIPA.png]] fMRIPA: A small fMRI positive activation scale going from red to yellow from 0-19, useful for displaying functional MRI volumes when don't need the blue of the fMRI scale. | ||
** [[image:DiscreteRandom.png]] Random: A random selection of 256 rgb colours, useful to distinguish between a small number of labeled regions (especially outside of the brain) | ** [[image:DiscreteRandom.png]] Random: A random selection of 256 rgb colours, useful to distinguish between a small number of labeled regions (especially outside of the brain) | ||
| − | |||
** [[image:DiscreteRed.png]] Red: A red scale of 256 values. Useful for layering with Cyan | ** [[image:DiscreteRed.png]] Red: A red scale of 256 values. Useful for layering with Cyan | ||
** [[image:DiscreteGreen.png]] Green: A green scale of 256 values, useful for layering with Magenta | ** [[image:DiscreteGreen.png]] Green: A green scale of 256 values, useful for layering with Magenta | ||
| Line 88: | Line 87: | ||
** [[image:DiscreteCool2.png]] Cool2: A scale from magenta to blue, 256 colours, ramp of cool colours that's complementary to Warm2 | ** [[image:DiscreteCool2.png]] Cool2: A scale from magenta to blue, 256 colours, ramp of cool colours that's complementary to Warm2 | ||
** [[image:DiscreteCool3.png]] Cool3: A scale from red to magenta, ramp of cool colours that's complementary to Warm3 | ** [[image:DiscreteCool3.png]] Cool3: A scale from red to magenta, ramp of cool colours that's complementary to Warm3 | ||
| − | ** [[image:DiscreteRandomIntegers.png]] RandomIntegers: A random scale | + | ** [[image:DiscreteRandomIntegers.png]] RandomIntegers: A random scale with 1000 entries. |
* Shade: | * Shade: | ||
** [[image:ShadeWarmShade1.png]] WarmShade1: A scale from black to red, of 256 colors, ramp of warm colors with variation in value that's complementary to CoolShade1 | ** [[image:ShadeWarmShade1.png]] WarmShade1: A scale from black to red, of 256 colors, ramp of warm colors with variation in value that's complementary to CoolShade1 | ||
| Line 117: | Line 116: | ||
** [[image:CartilegeMRIdGEMRIC15T.png]] dGEMRIC-1.5T: Useful for displaying 1.5 tesla delayed gadolinium-enhanced MRI of cartilage | ** [[image:CartilegeMRIdGEMRIC15T.png]] dGEMRIC-1.5T: Useful for displaying 1.5 tesla delayed gadolinium-enhanced MRI of cartilage | ||
** [[image:CartilegeMIdGEMRIC3T.png]] dGEMRIC-3T: Useful for displaying 3 Tesla delayed gadolinium-enhanced MRI of cartilage | ** [[image:CartilegeMIdGEMRIC3T.png]] dGEMRIC-3T: Useful for displaying 3 Tesla delayed gadolinium-enhanced MRI of cartilage | ||
| − | *Default Labels from File: | + | * Default Labels from File: |
** a list of color nodes loaded from files, from the default Slicer color directory | ** a list of color nodes loaded from files, from the default Slicer color directory | ||
| + | **ColdToHotRainbow: a shifted rainbow that runs from blue to red, useful when needing to display a volume for which larger values are hotter | ||
| + | **HotToColdRainbow: a shifted rainbow that runs from red to blue, useful when needing to display a volume for which larger values are colder | ||
** [[image:LabelsPelvis.png]] PelvisColor: useful for displaying segmented pelvic MRI volumes | ** [[image:LabelsPelvis.png]] PelvisColor: useful for displaying segmented pelvic MRI volumes | ||
| − | * [[image:SlicerBrainLUT2010.png]] Slicer3_2010_Brain_Labels: a brain segementation table with 16 labels defined. | + | ** [[image:SlicerBrainLUT2010.png]] Slicer3_2010_Brain_Labels: a brain segementation table with 16 labels defined. |
** [[image:LabelsNonSemantic.png]] 64Color-Nonsemantic: A color table with no semantic labels, pure color information | ** [[image:LabelsNonSemantic.png]] 64Color-Nonsemantic: A color table with no semantic labels, pure color information | ||
| + | ** MediumChartColors: A Stephen Few palette. Similar to the Dark Bright color but in the medium range of intensity. May be a better choice for bar charts. | ||
| + | ** DarkBrightChartColors: Palette designed by Stephen Few in "Practical Rules for Using Colors in Charts". Similar to the Maureen Stone palette. Stephen Few recommends this palette for highlighting data. This palette is useful when display small points or thin lines. Again, this palette is good for categorical data. | ||
** [[image:SlicerShortLUT.png]] Slicer3_2010_LabelColors: a table with 16 labels defined. | ** [[image:SlicerShortLUT.png]] Slicer3_2010_LabelColors: a table with 16 labels defined. | ||
**AbdomenColors: useful for displaying segmented abdominal MRI volumes | **AbdomenColors: useful for displaying segmented abdominal MRI volumes | ||
**SPL-BrainAtlas-ColorFile: useful for displaying segmented brain MRI volumes | **SPL-BrainAtlas-ColorFile: useful for displaying segmented brain MRI volumes | ||
| − | **SPL-BrainAtlas- | + | **SPL-BrainAtlas-2012-ColorFile: an updated brain segmentation color node |
| − | ** | + | **LightPaleChartColors: A light pale palette from Stephen Few, useful for charts. |
| − | ** | + | **SPL-BrainAtlas-2009-ColorFile: an updated brain segmentation color node |
| − | ** | + | * User Generated: A user defined color table, use the editor to specify it |
| + | ** Copies of other color tables get displayed in this category | ||
| + | ** If you create a new color node, it will appear here | ||
* File | * File | ||
** If you load a color file from File -> Add Data, it will appear here | ** If you load a color file from File -> Add Data, it will appear here | ||
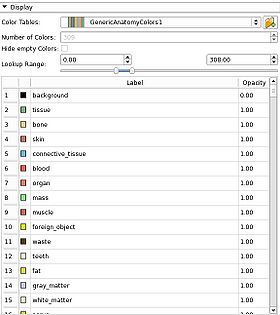
| − | * | + | *GenericAnatomyColors: a list of whole body anatomy labels and useful colors for them, the default for the Editor module creating new label map volumes |
| − | * | + | *Generic Colors: a list of colors with the names being the same as the integer value for each entry |
| + | |||
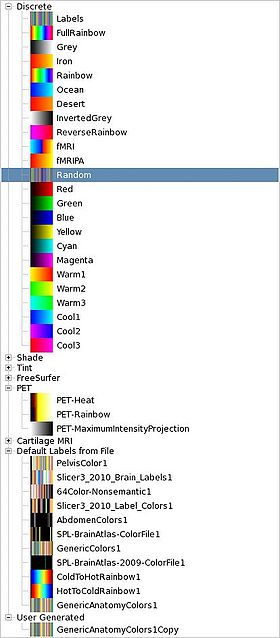
|[[Image:Slicer4-Colors-Tables.jpeg|thumb|280px|Built in color tables]] | |[[Image:Slicer4-Colors-Tables.jpeg|thumb|280px|Built in color tables]] | ||
Revision as of 19:17, 14 November 2013
Home < Documentation < Nightly < Modules < Colors
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
| |||||||
|
Module Description
|
The Colors Module manages color look up tables. Color look up tables are used by mappers to translate between an integer and a colour value for display of models and volumes. Slicer supports three kinds of tables:
You can load a color table file from the File -> Add Data dialog. File formatThe color file format is a plain text file with the .txt or .ctbl extension. Each line in the file has: label name R G B A label is an integer, name a string, and RGBA are 0-255. File example: # Comments if the line start with # 0 air 0 0 0 0 1 bone 255 255 255 255 whatever after the Alpha value is discarded 2 tumor 255 128 0 255 ... Custom LUTsYou can create custom LUTs by creating a table with the colors on the wiki, saving to file and then loading them into Slicer. CategoriesThe colors are divided up into categories:
|
Use Cases
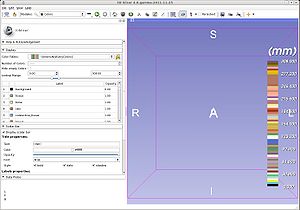
The Colors module Dispay panel can be popped up as a stand alone widget and used to select colors in other modules of Slicer4.
- The Models module uses it to select surface model colours.
- The Volumes module uses it to select color maps for label map volumes to control which colors are used to display the scalar values at each voxel
- The Editor uses it to select colors with which to paint on label map volumes.
Tutorials
N/A
Panels and their use
A list of all the panels in the interface, their features, what they mean, and how to use them. For instance:
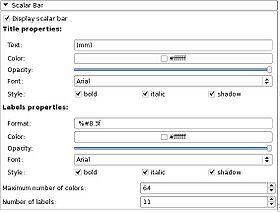
|
|
|
Similar Modules
- The Volumes, Editor and Models modules use the colours to adjust the display properties for label map volumes and surface models
References
N/A
Information for Developers
| Section under construction. |