Home < Documentation < Nightly < Extensions < SlicerPathology
Introduction and Acknowledgements
|
Extension: SlicerPathology
Acknowledgments:
This work was supported by via the NIH-National Cancer Institute Grant U24 CA180918, as well as, U24 CA180918 Quantitative Image Informatics for Cancer Research (QIICR), http://qiicr.org, PIs Ron Kikinis and Andrey Fedorov, Brigham and Women's Hospital.
Author: Erich Bremer
Contributor 1: Yi Gao
Contributor 2: Nicole Aucoin (SPL)
Contributor 3: Andrey Fedorov (SPL)
Contributor 4: Jean-Christophe Fillion-Robin (Kitware)
Contact: Erich Bremer, <email>erich.bremer@stonybrook.edu</email>
|
Module Description
This extension provides tools for automatic and semi-automatic pathology image segmentation.
Use Cases
Tutorials
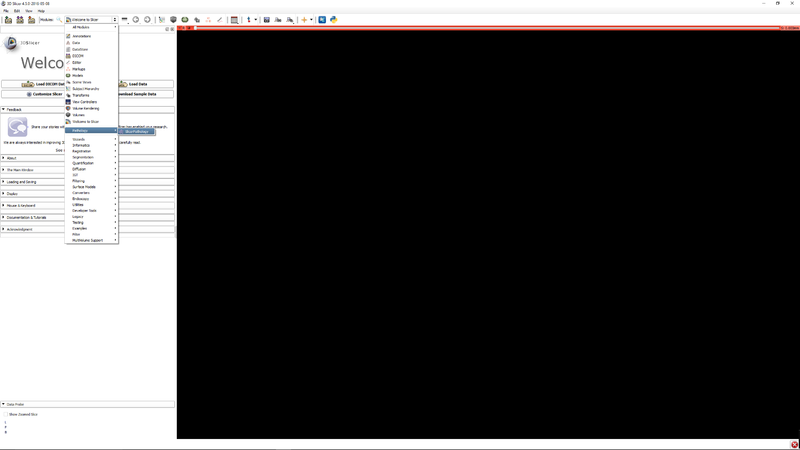 Step 1 - Go to the SlicerPathology Extension. |
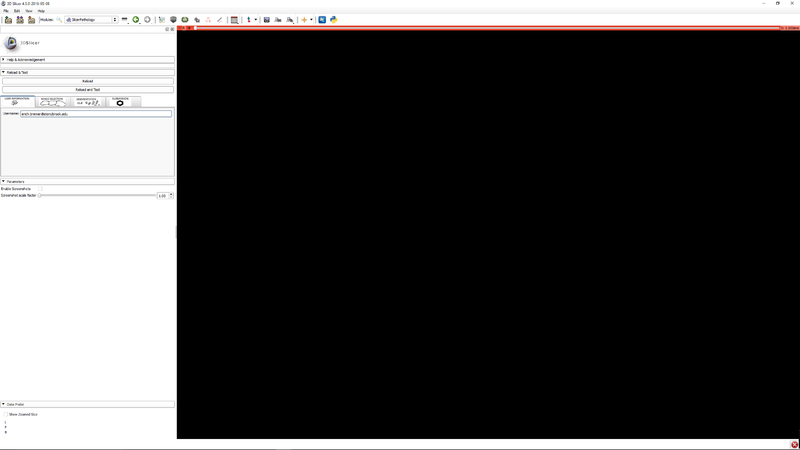 Step 2 - Click the user information tab. Enter your email address. This is used for identification of you in the file data files. |
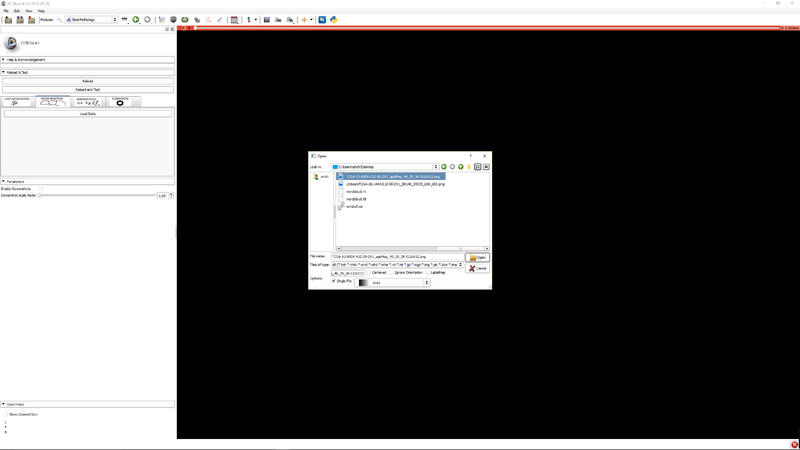 Step 3 - Click "Load data" to select an image. |
 Step 4 - Click the button Quick TCGA Effect button to activate the effect. |
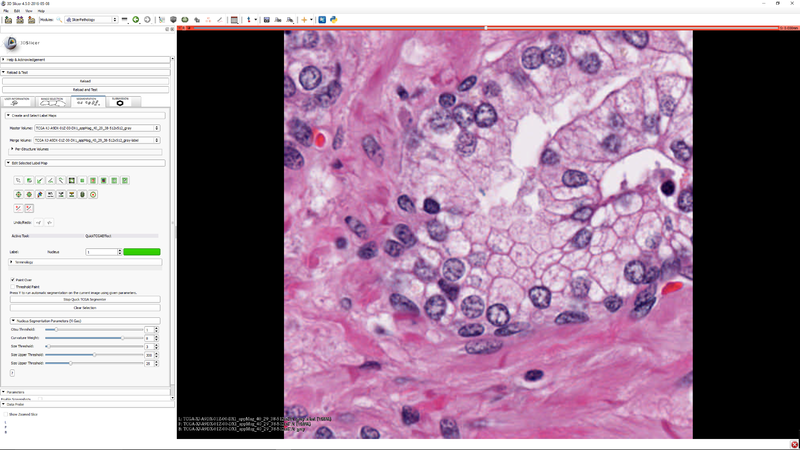 Step 5 - Click "Start TCGA Segmenter and adjust the five segmentation algorithm parameters as needed. |
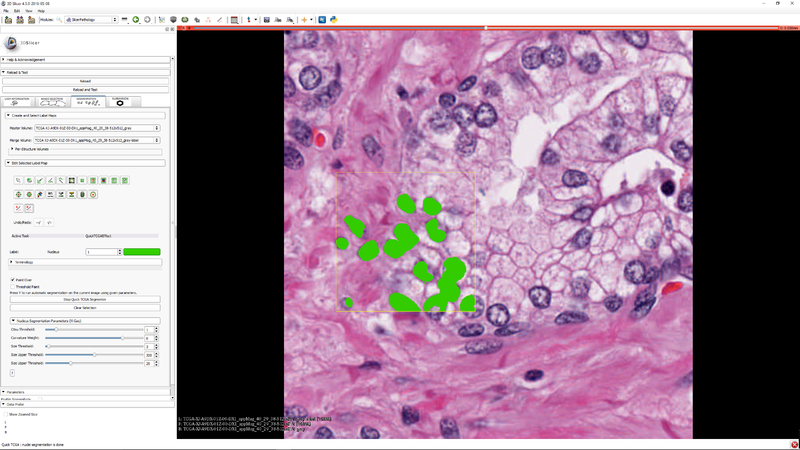 Step 6 - you can select a subregion to speed up the parameter tuning process. Press "Y" to execute the segmenter on the subregion (if it was selected) or the whole image |
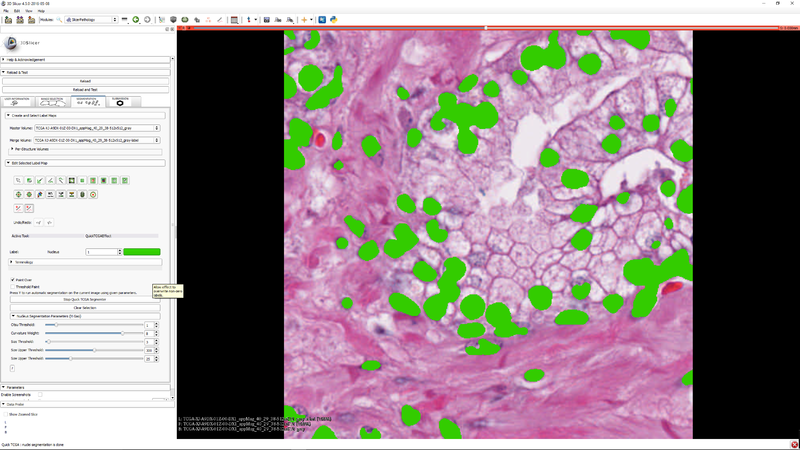 Step 7 - you can click "Clear Selection" to clear the selected sub-region so that the segmenter operates on the whole image. |
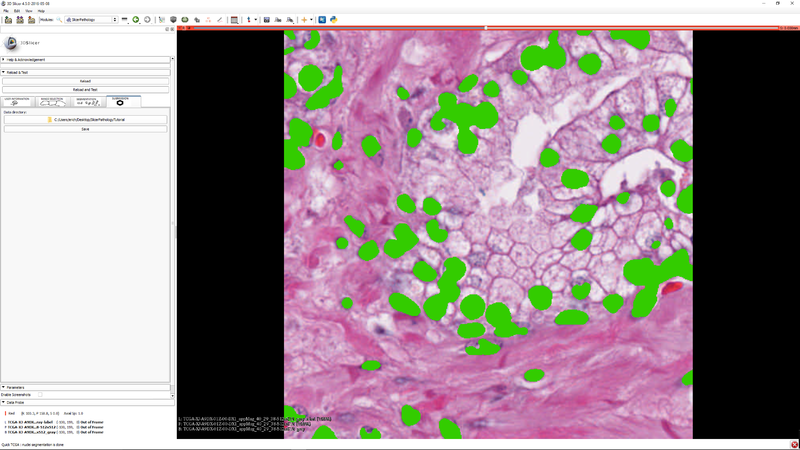 Step 8 - "Click the "Submission" tab so that you can save your label masks and related meta data. (meta data will be stored as JSON) |
 Step 9 - "This is a sample of the stored meta data. Notice the addition of your username for identification. |
References
Information for Developers
The source code for SlicerPathology is available at https://github.com/SBU-BMI/SlicerPathology








