Difference between revisions of "Documentation/4.4/gif tutorial"
From Slicer Wiki
| Line 46: | Line 46: | ||
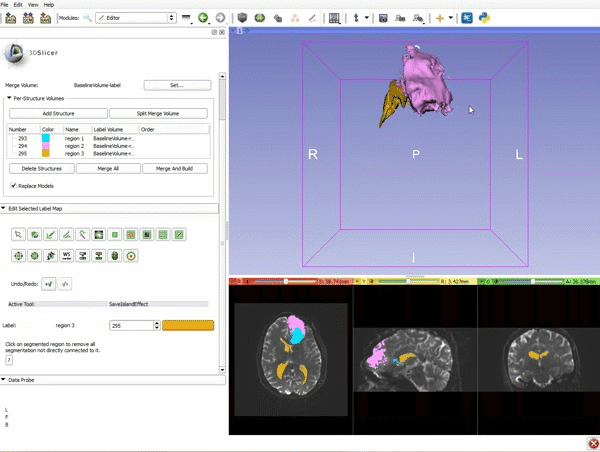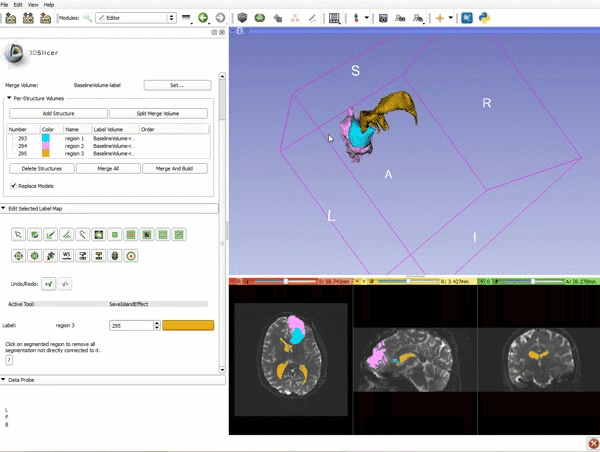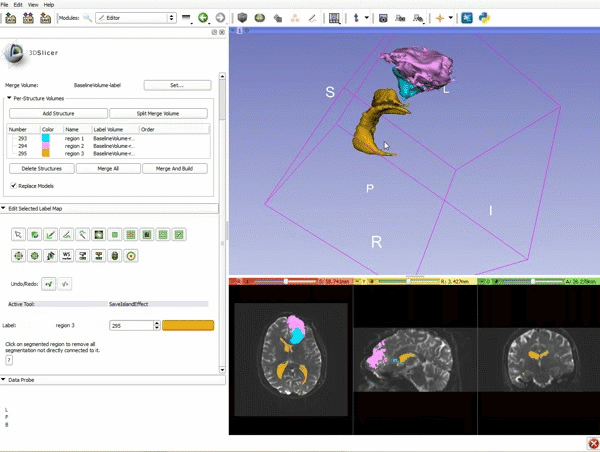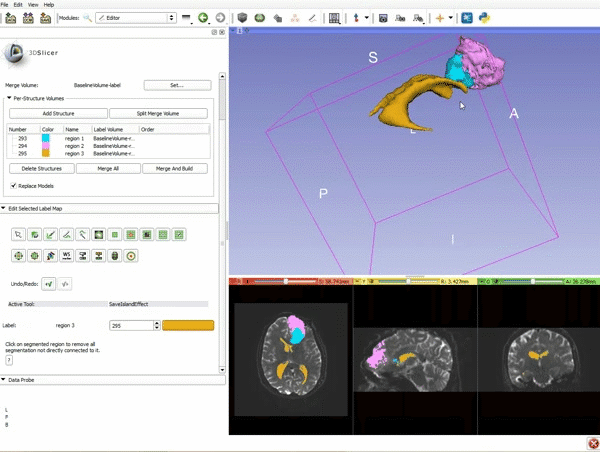
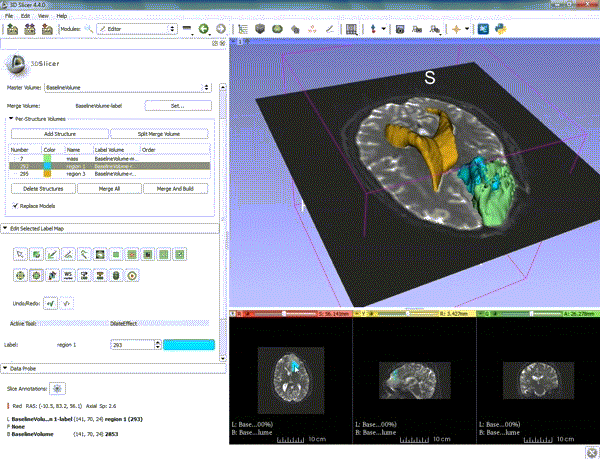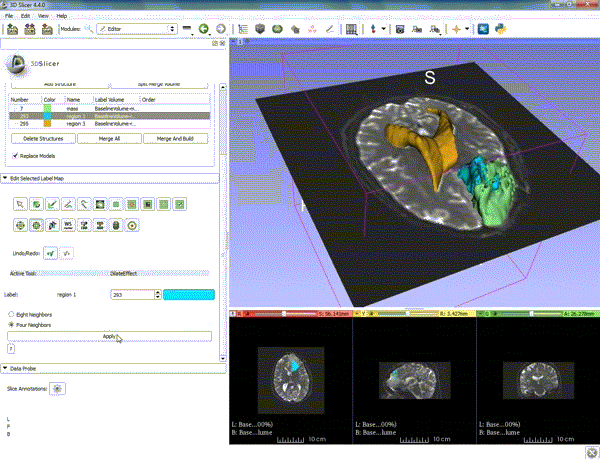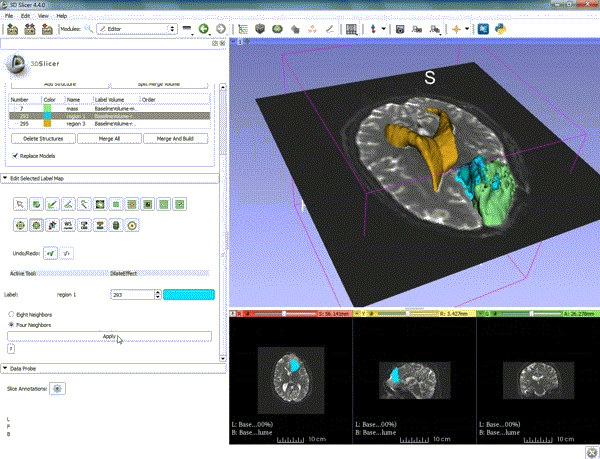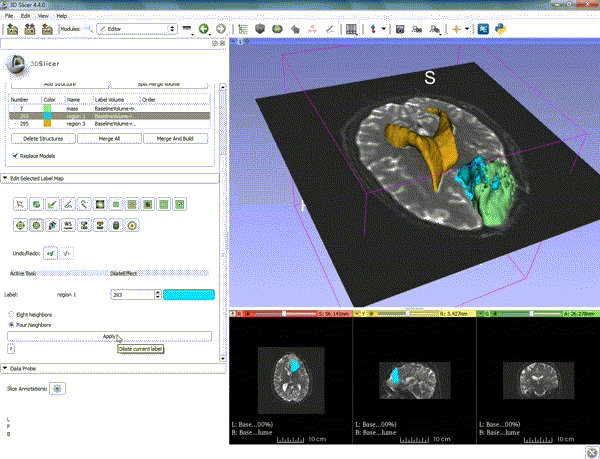
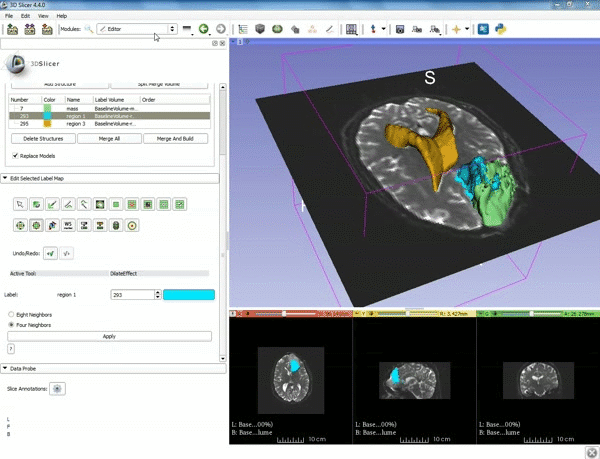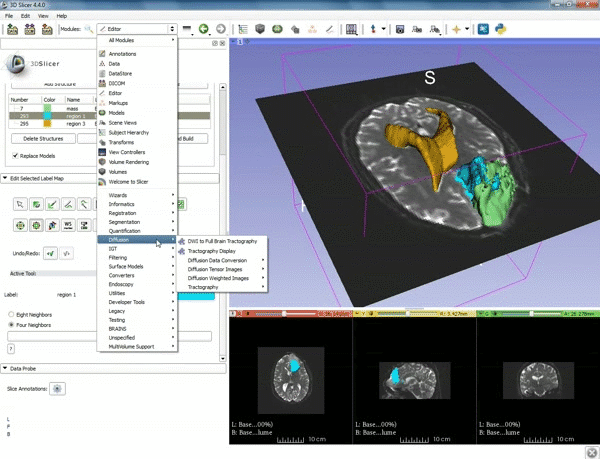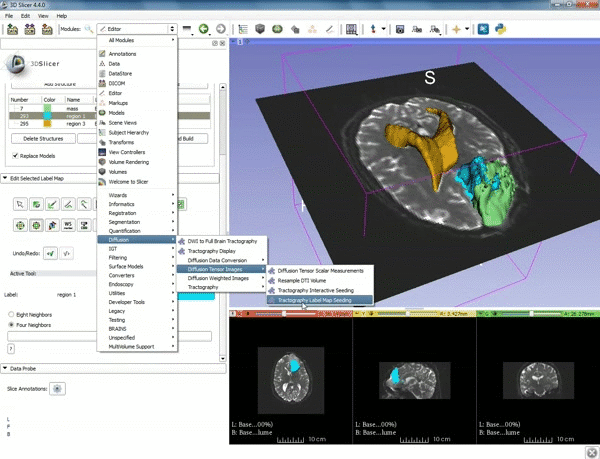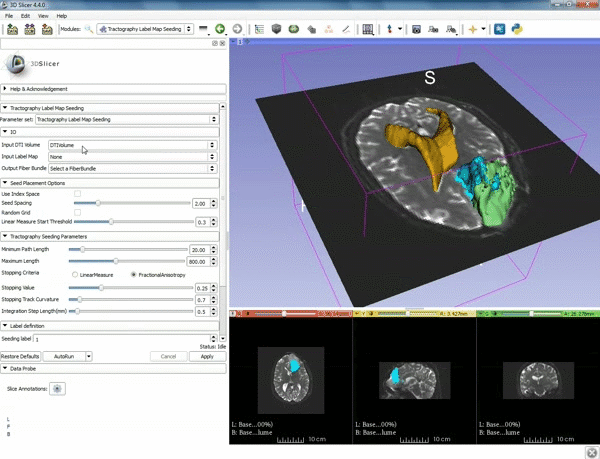
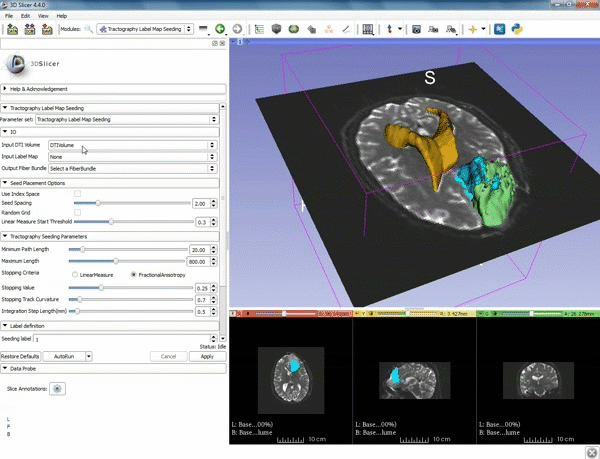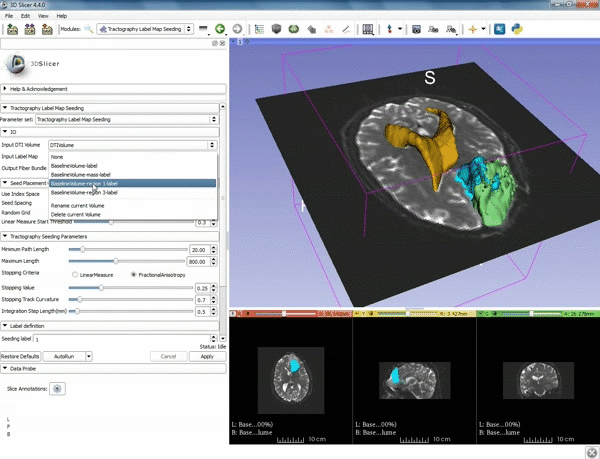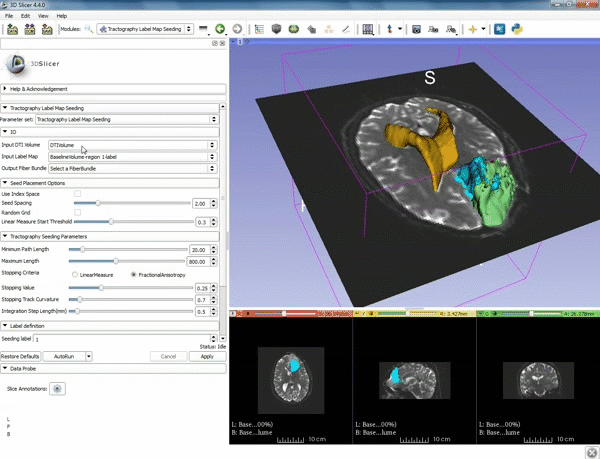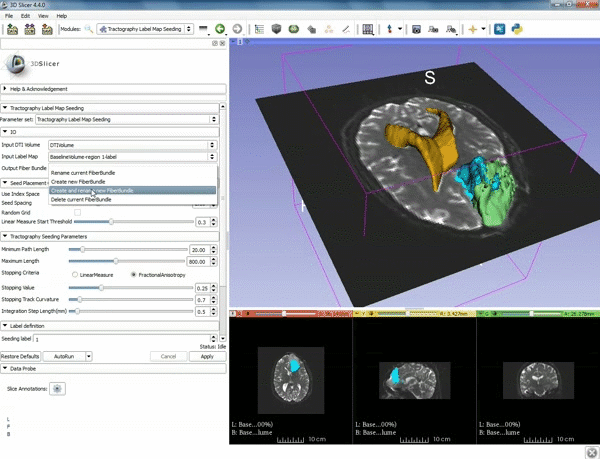
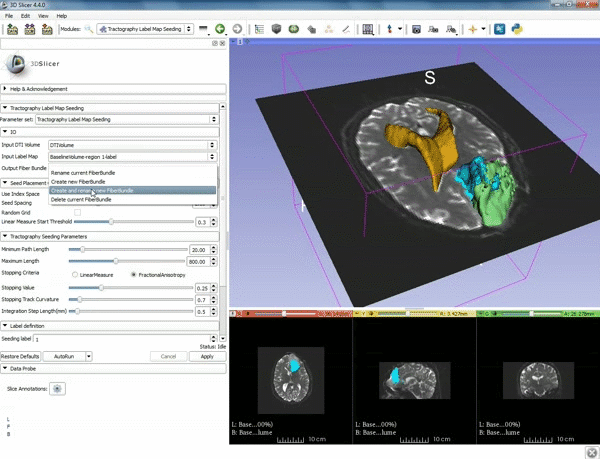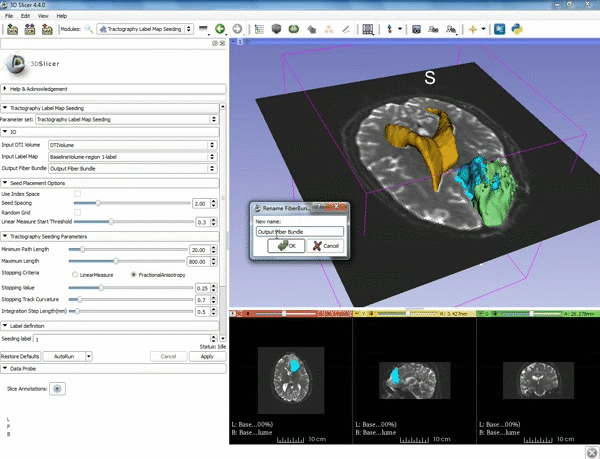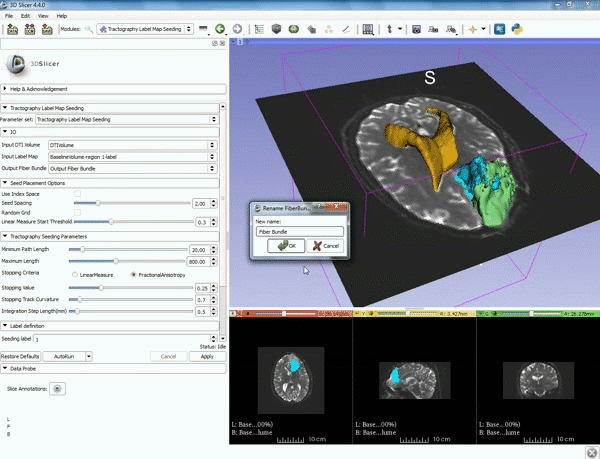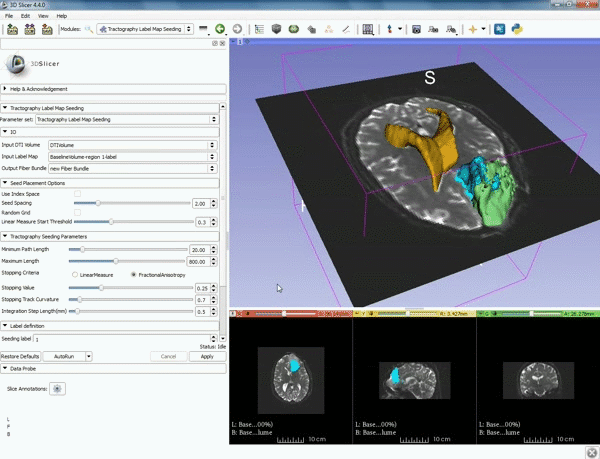
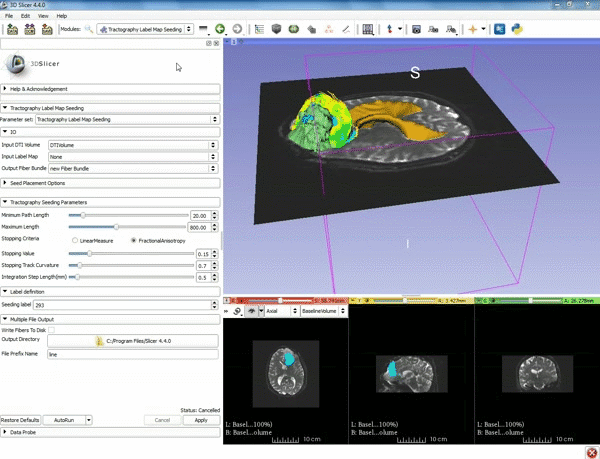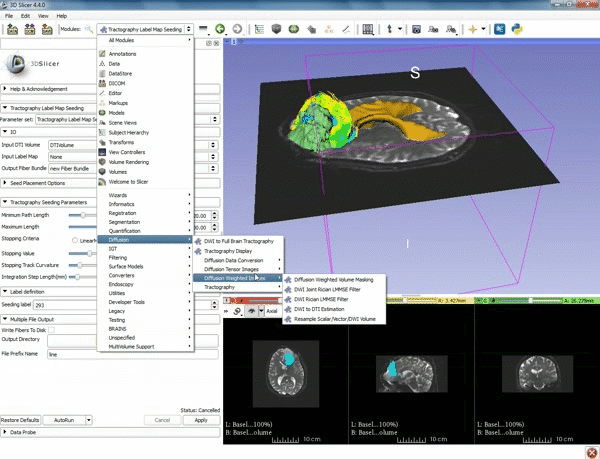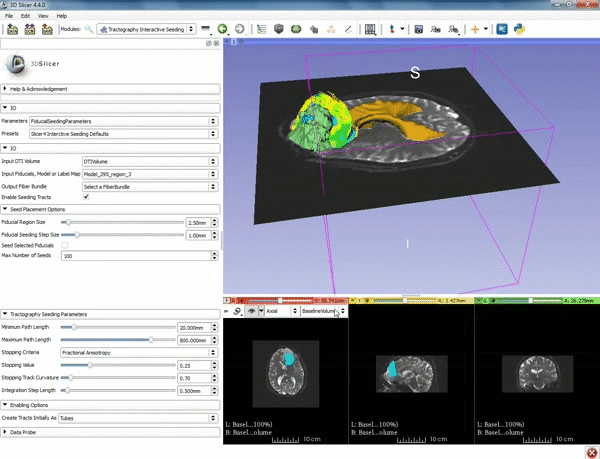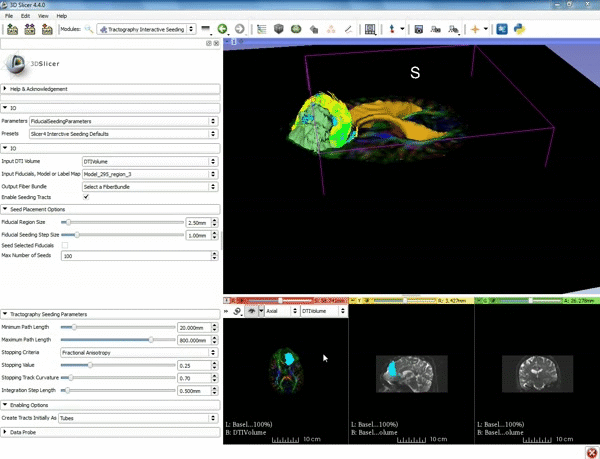
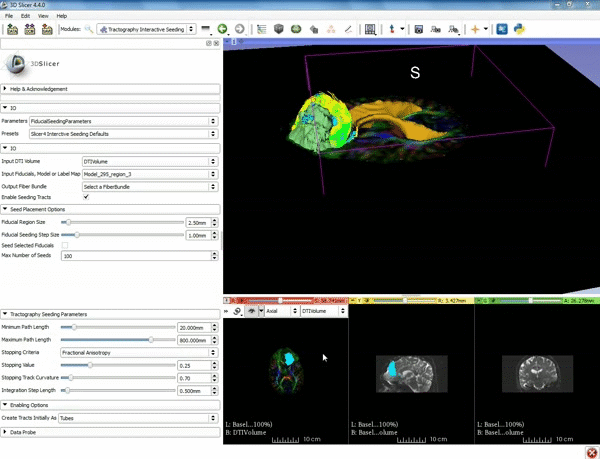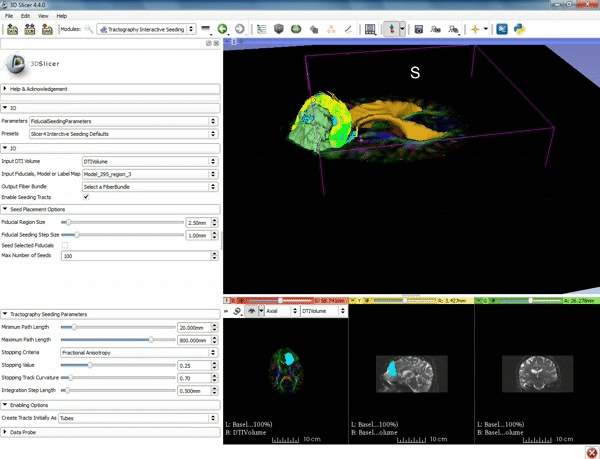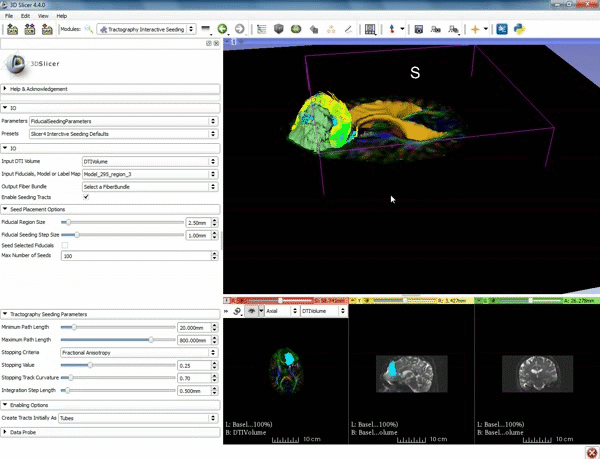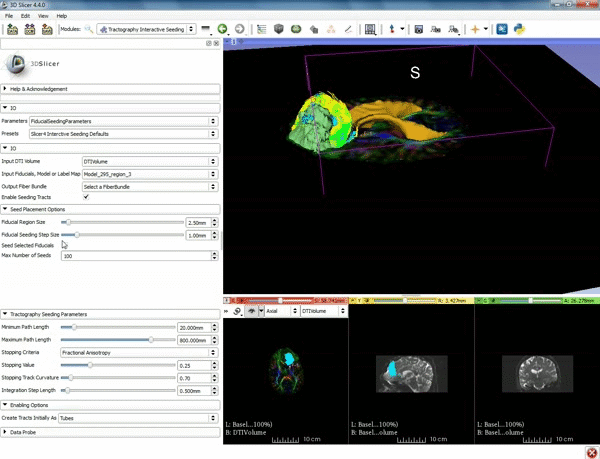
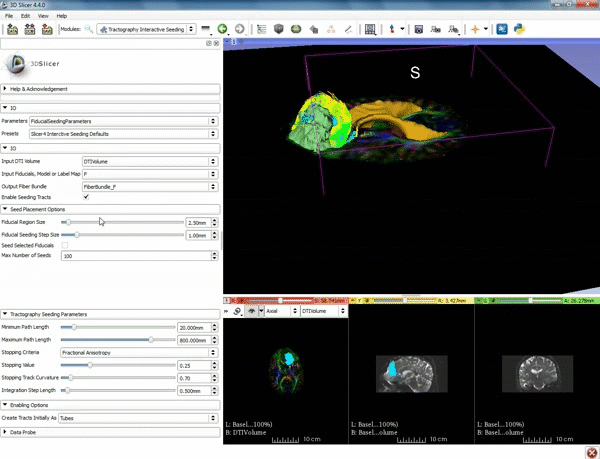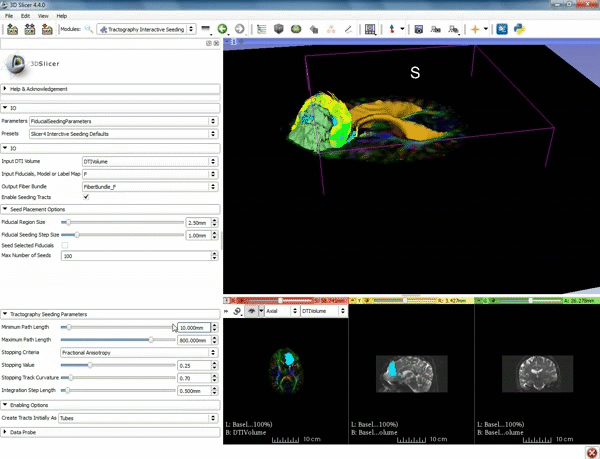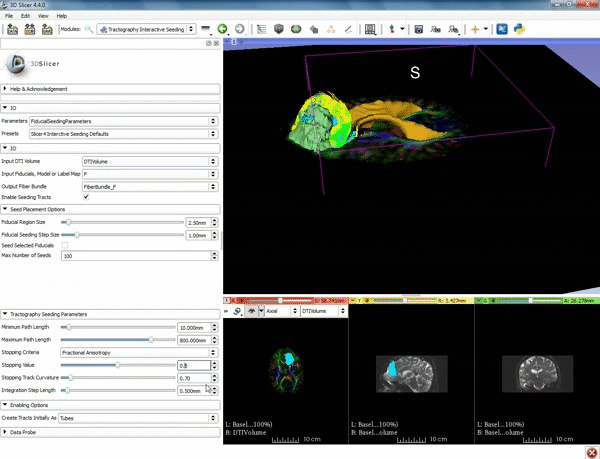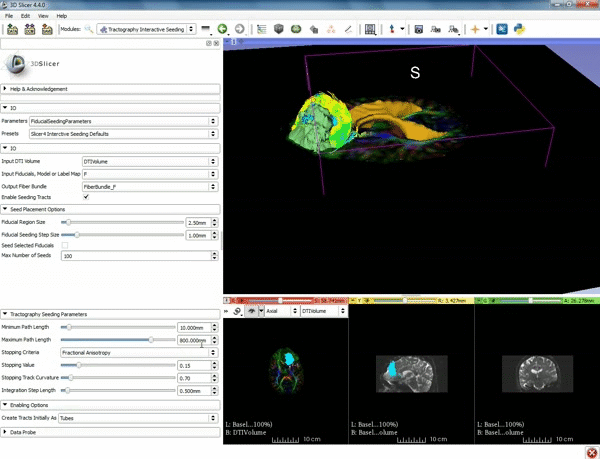
| − | ===Tractography=== | + | ===Tractography exploration of peritumoral white matter fibers=== |
<div style="width: 62%; height:525px; overflow:auto; border: 2px solid #088; margin: 1em auto 1em auto;"> | <div style="width: 62%; height:525px; overflow:auto; border: 2px solid #088; margin: 1em auto 1em auto;"> | ||
Revision as of 19:22, 9 July 2015
Home < Documentation < 4.4 < gif tutorial|
Contents
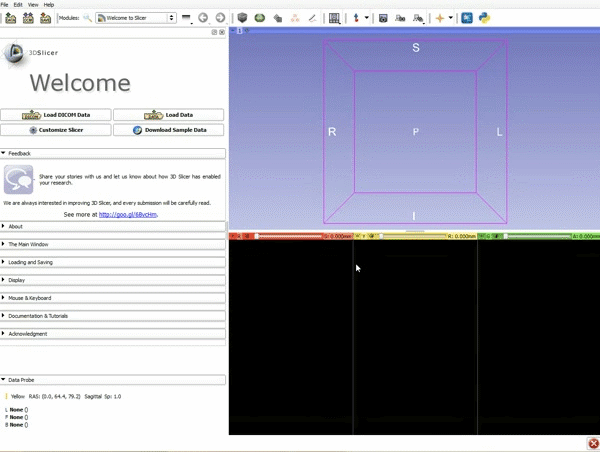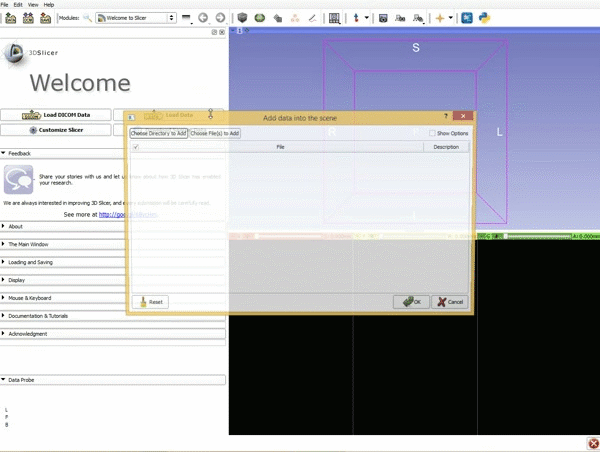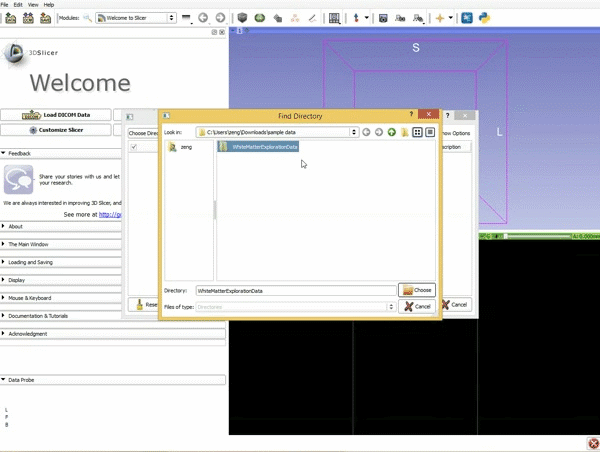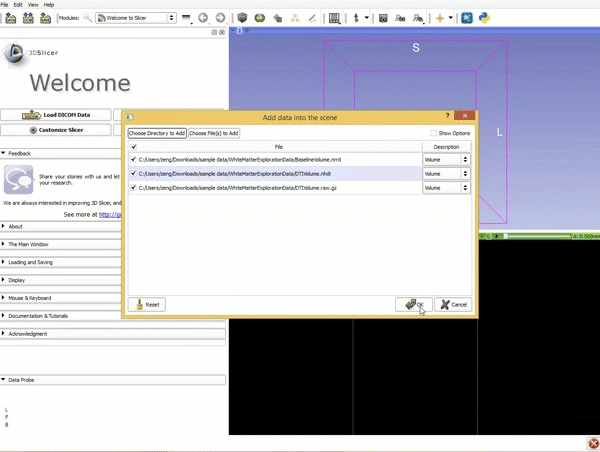
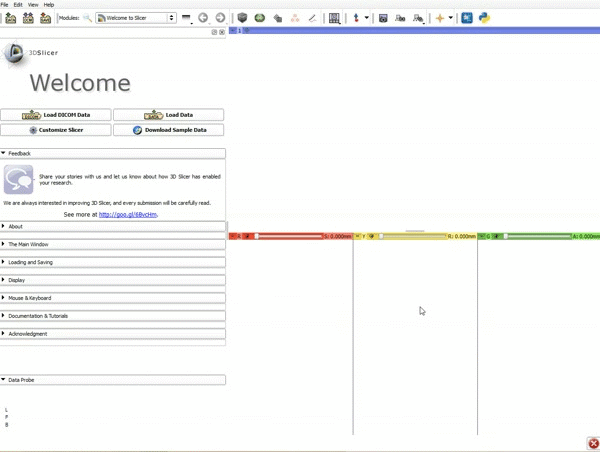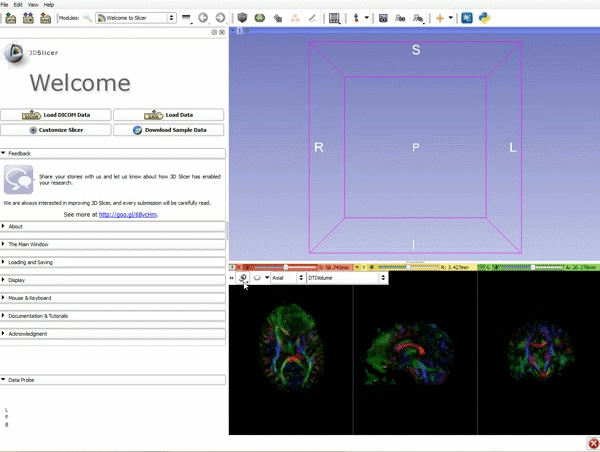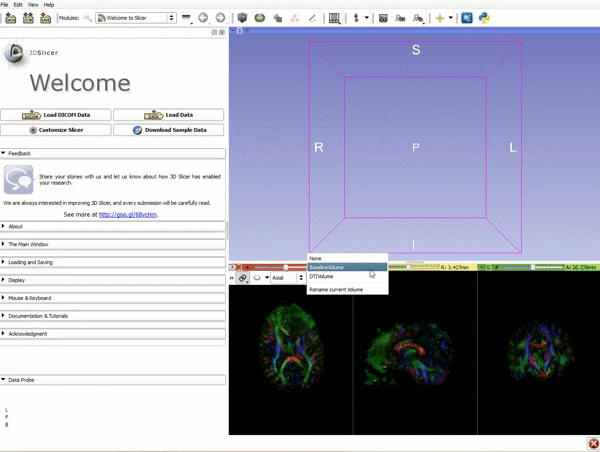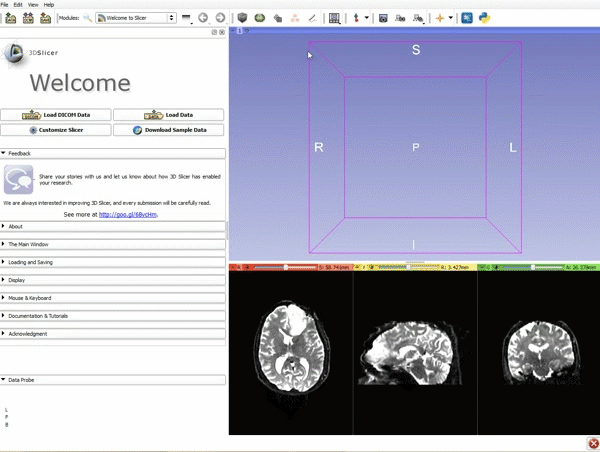
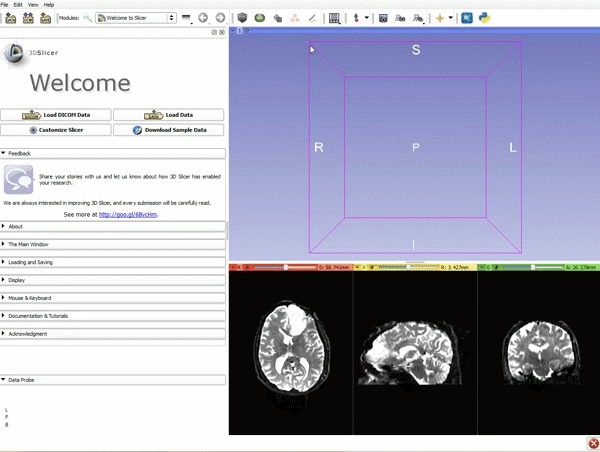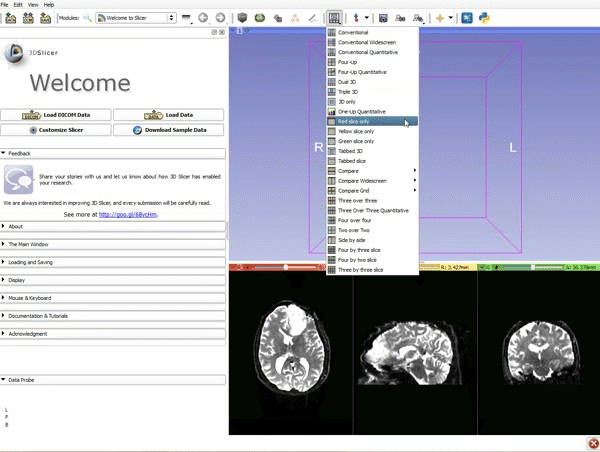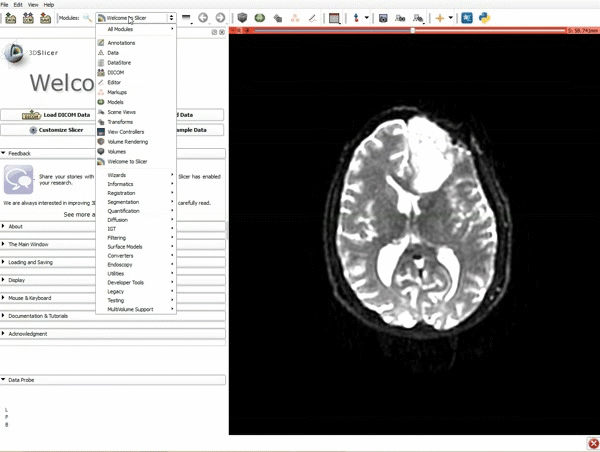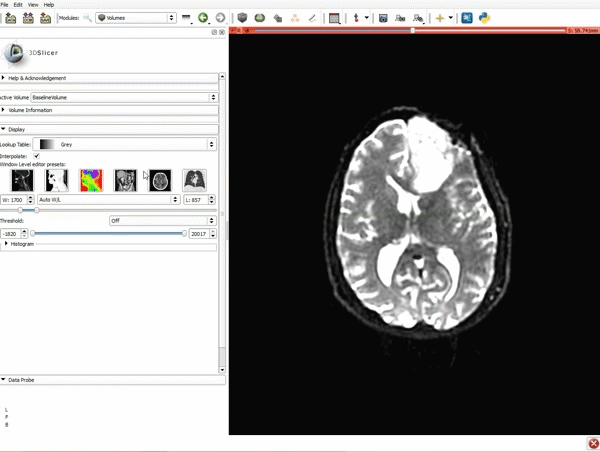
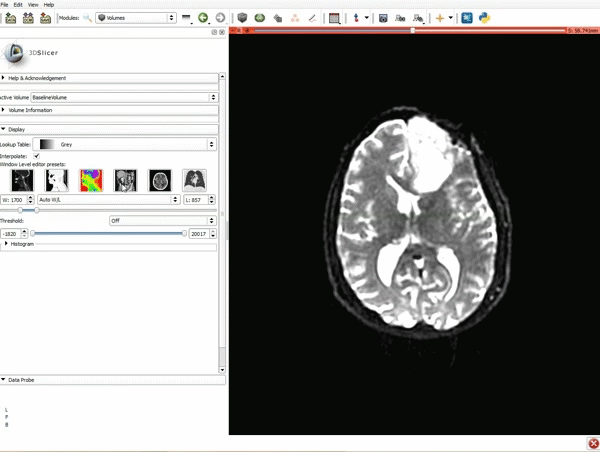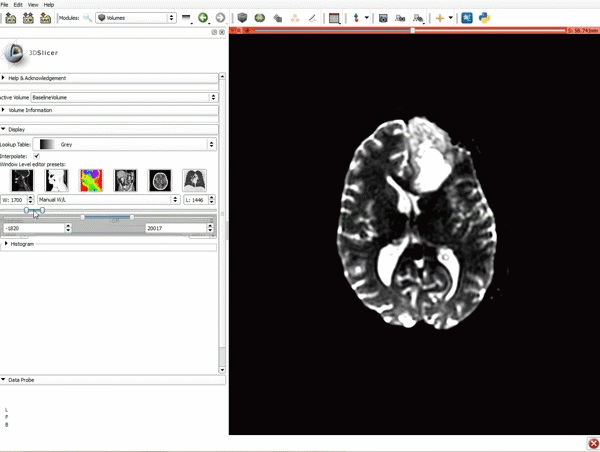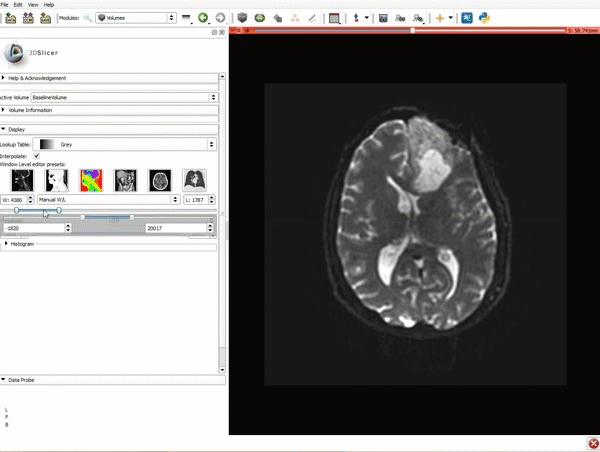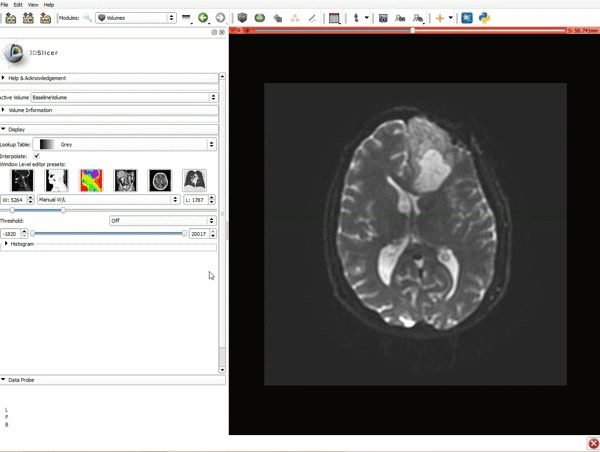
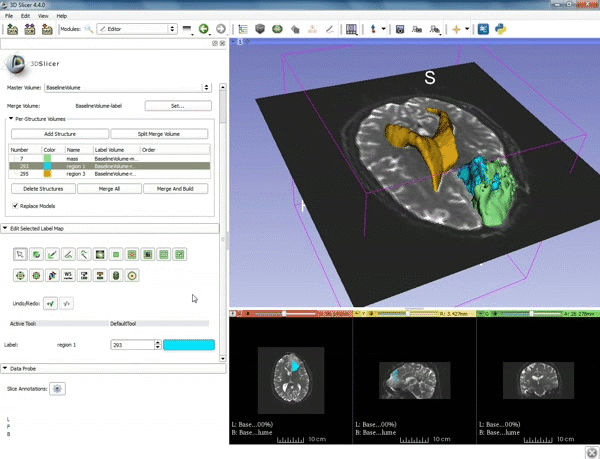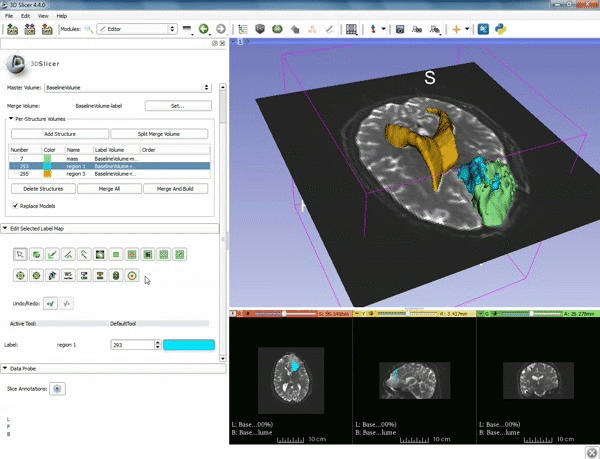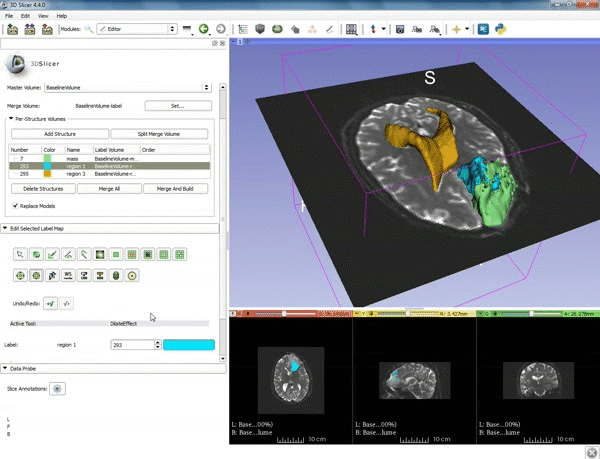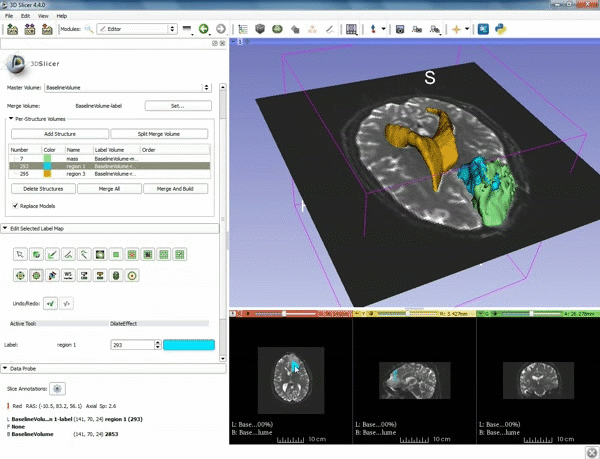
Loading DTI and Baseline Data
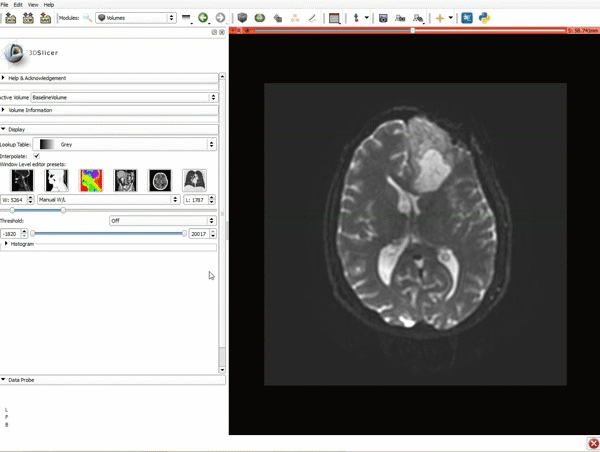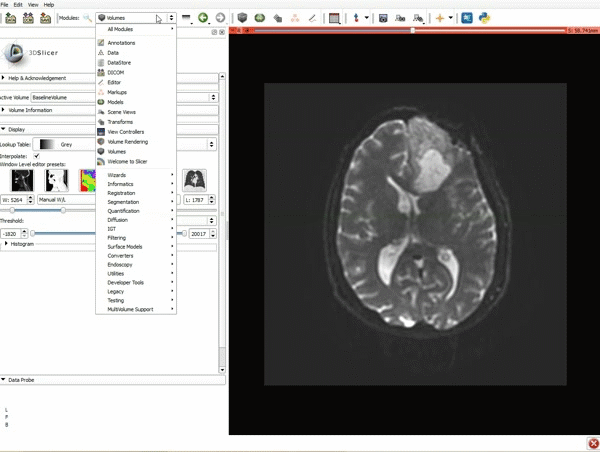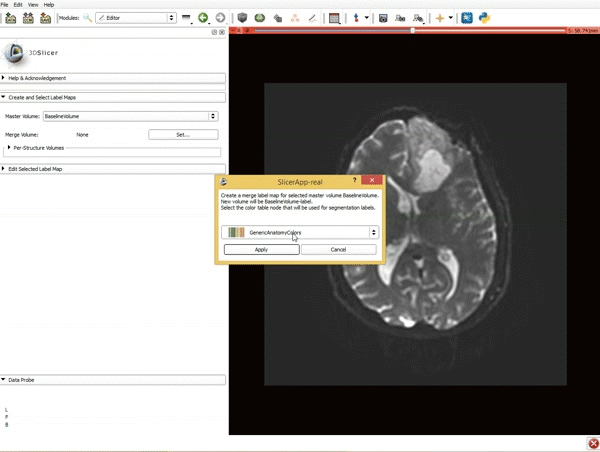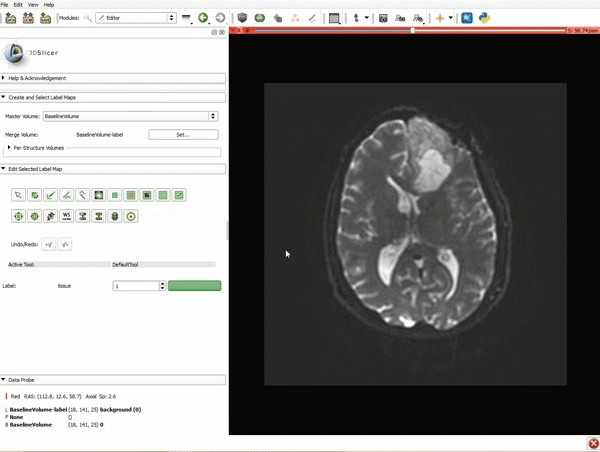
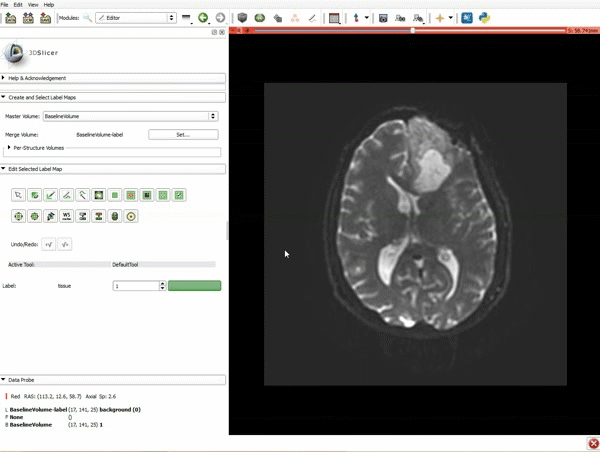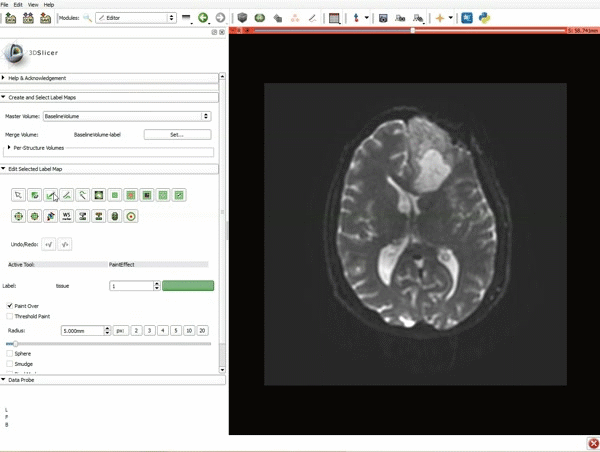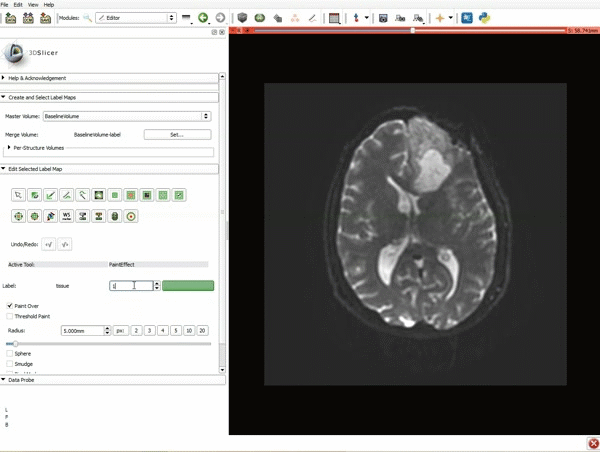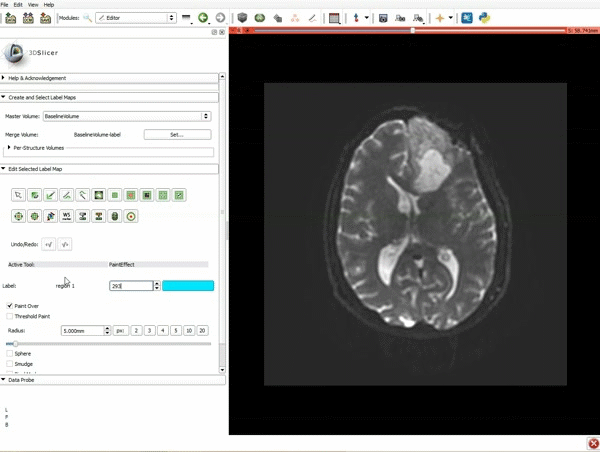
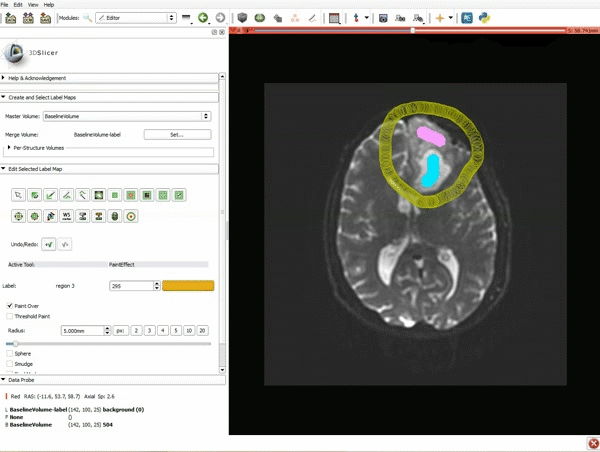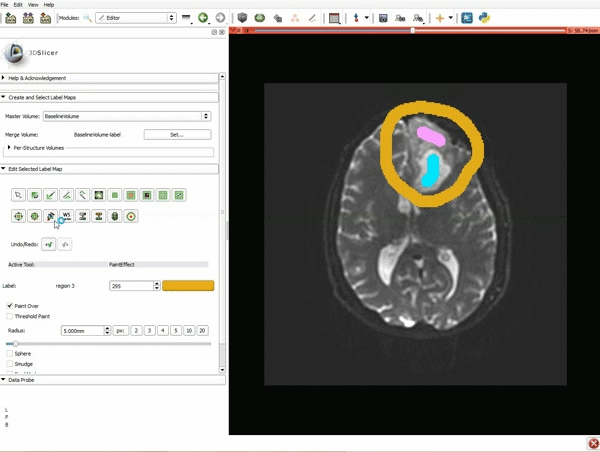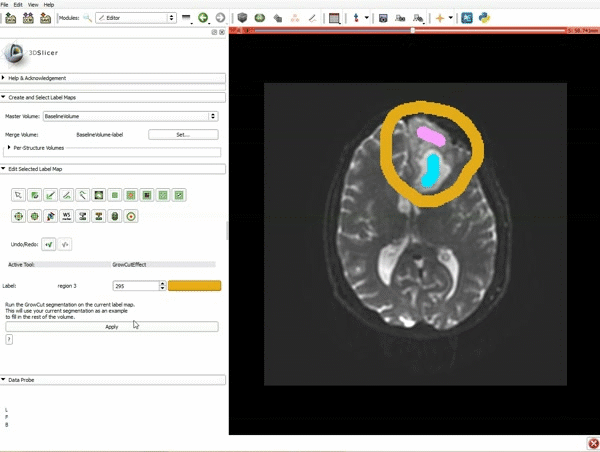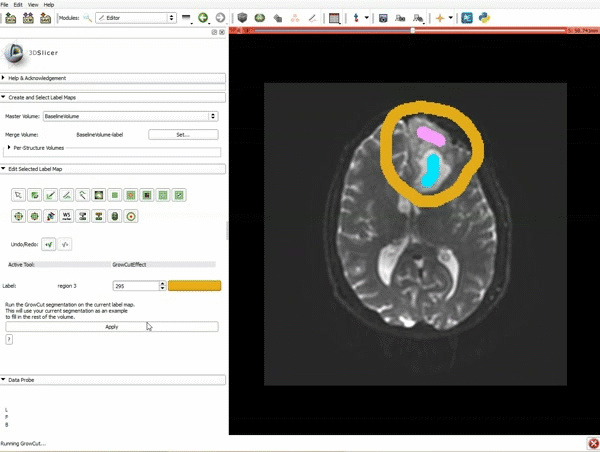
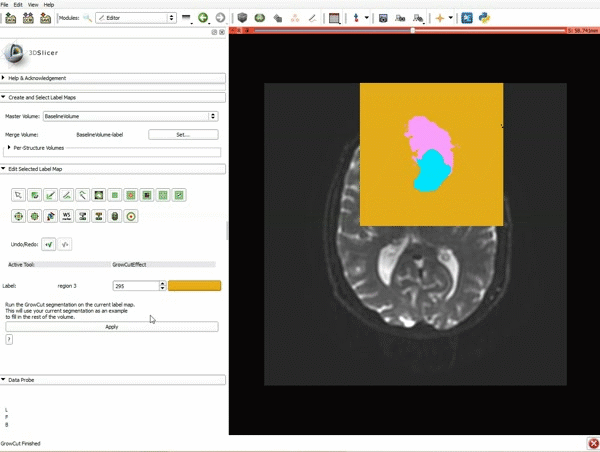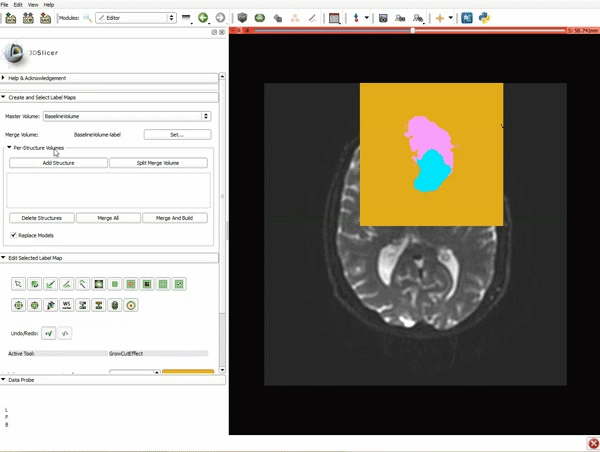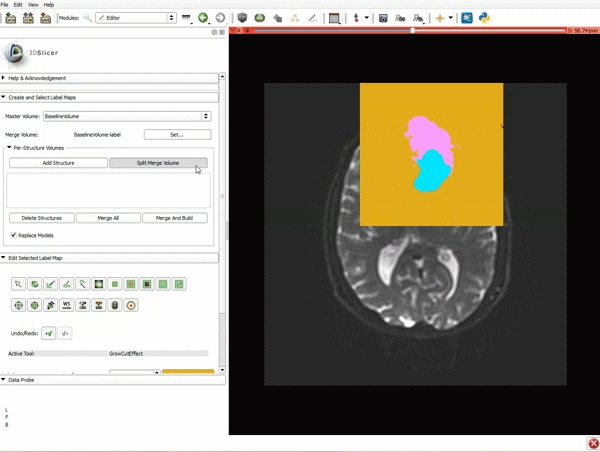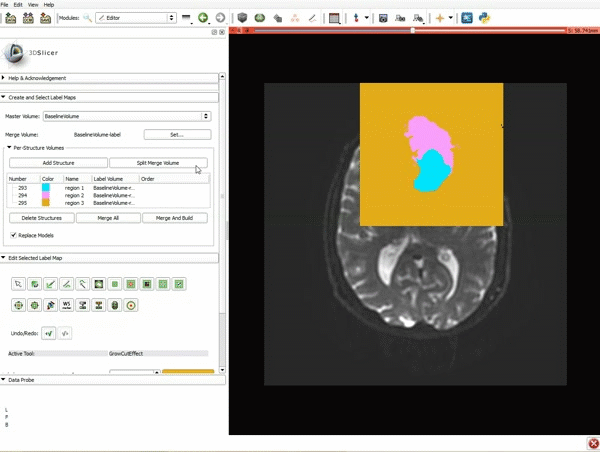
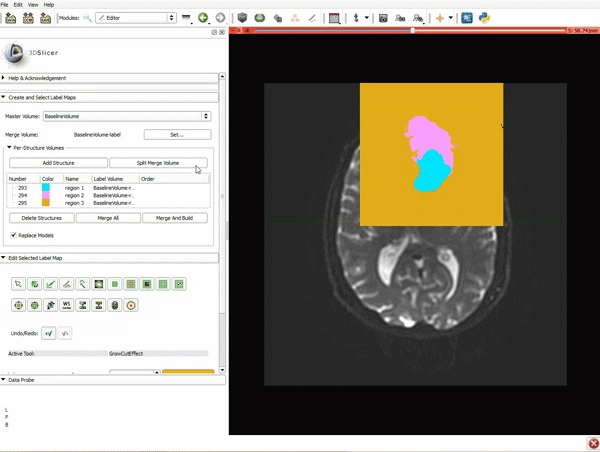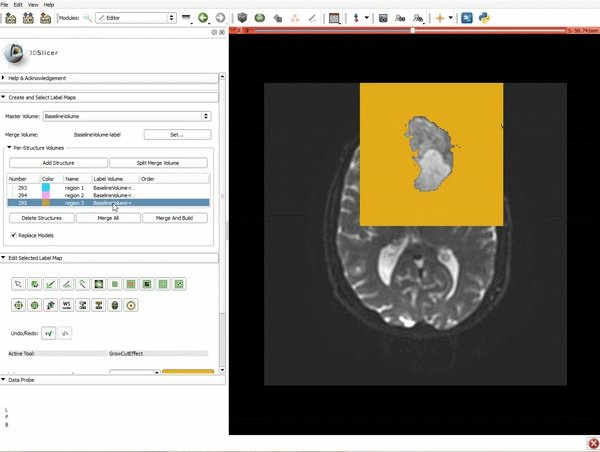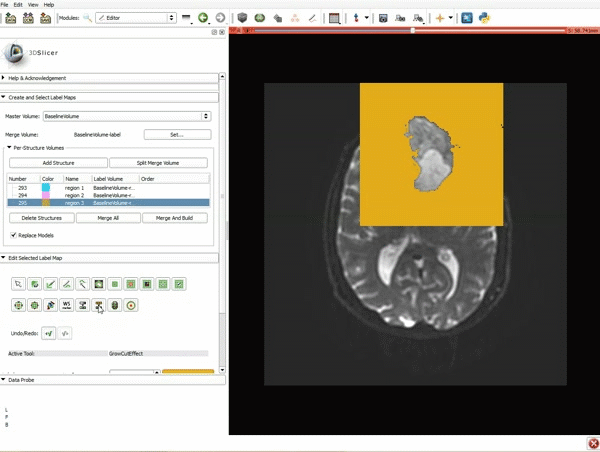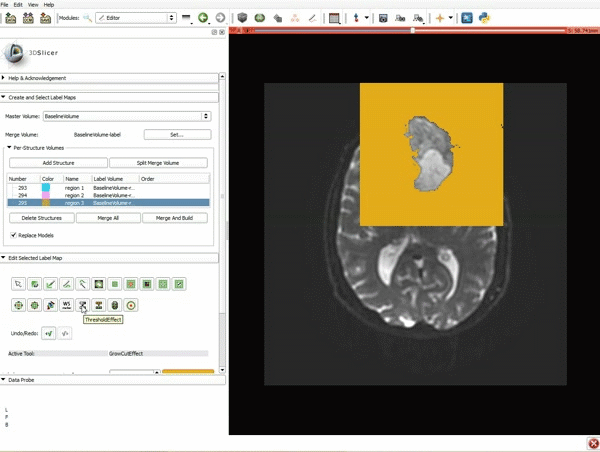
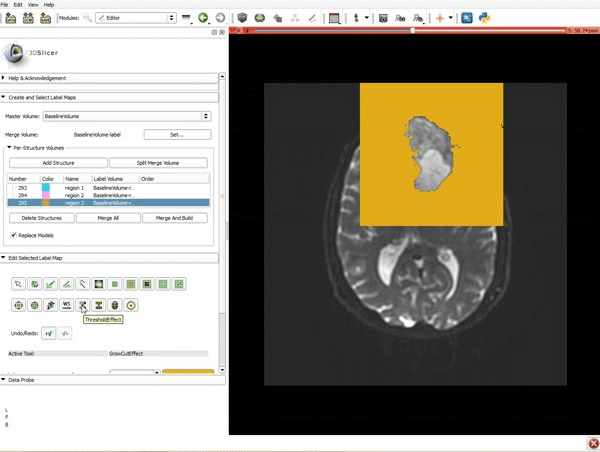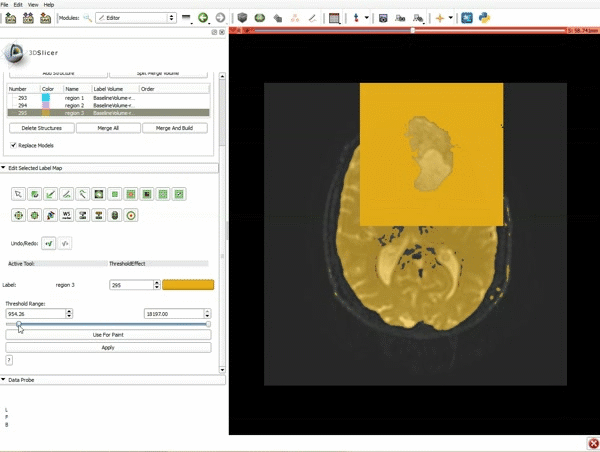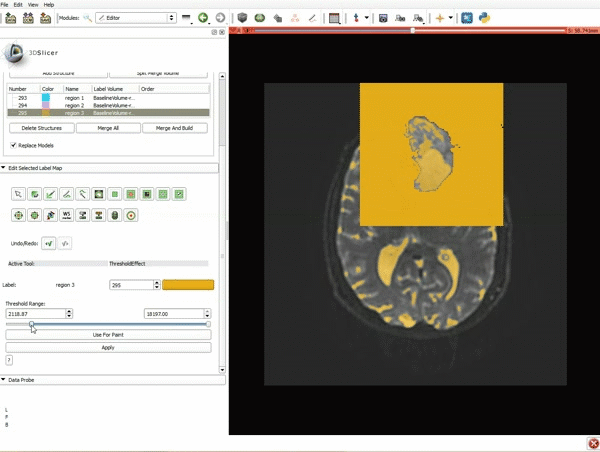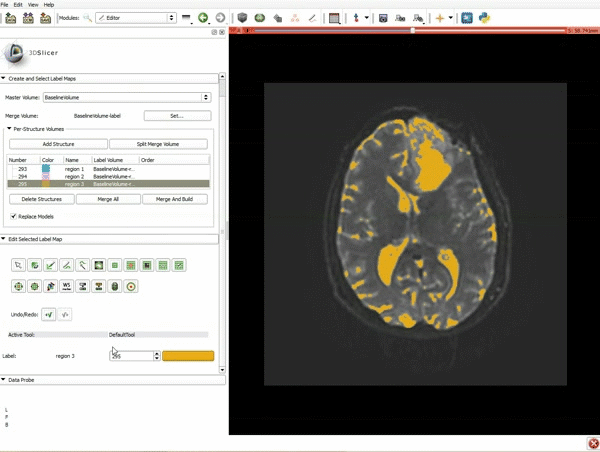
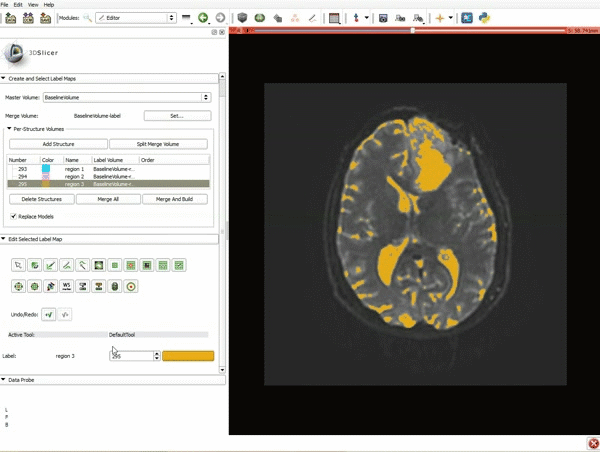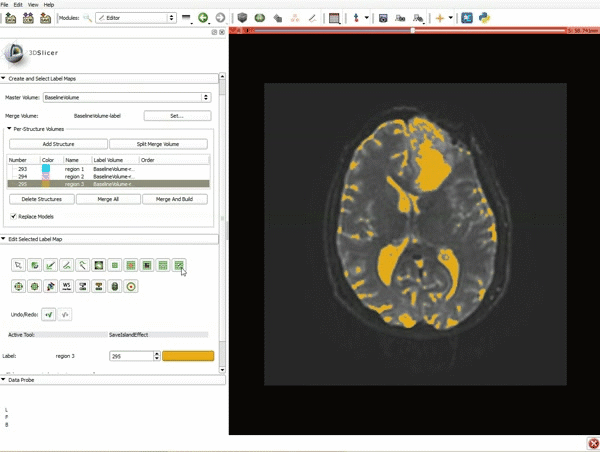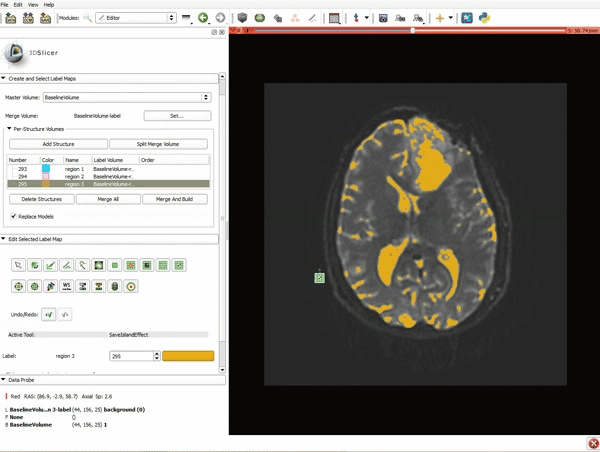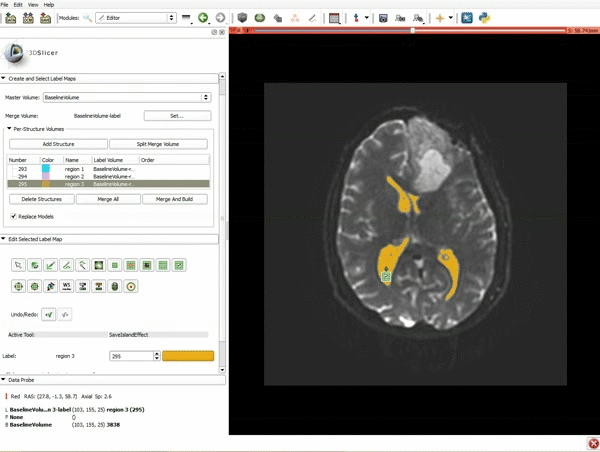
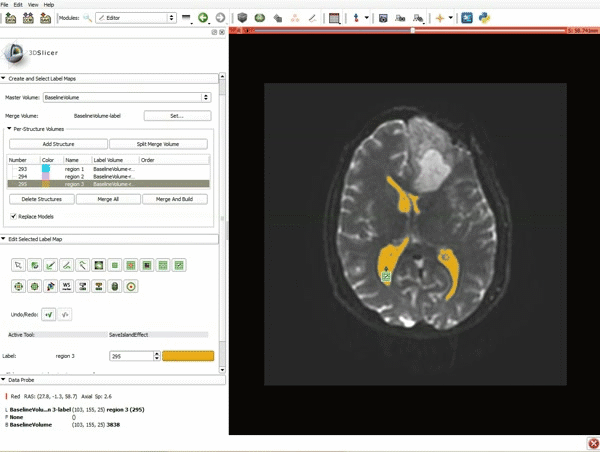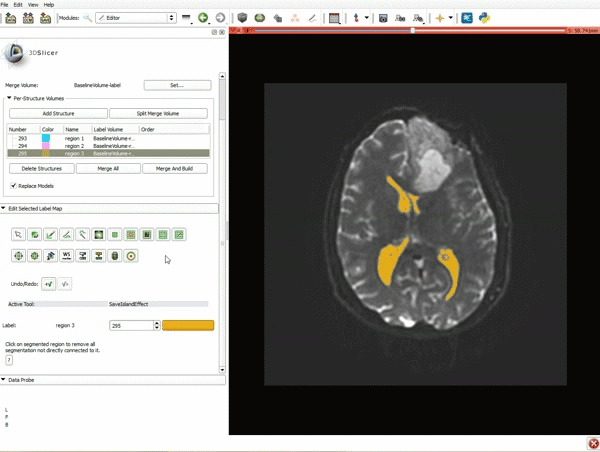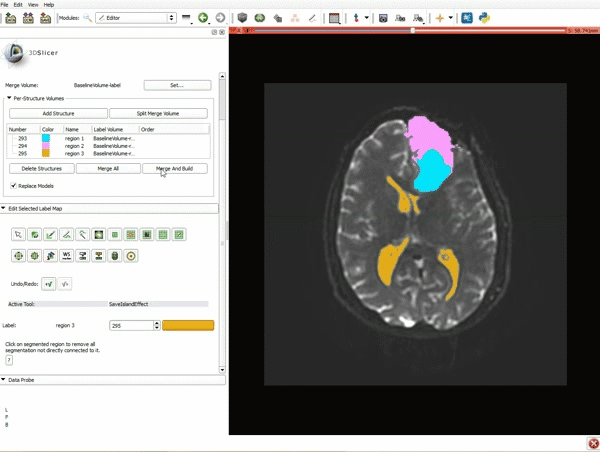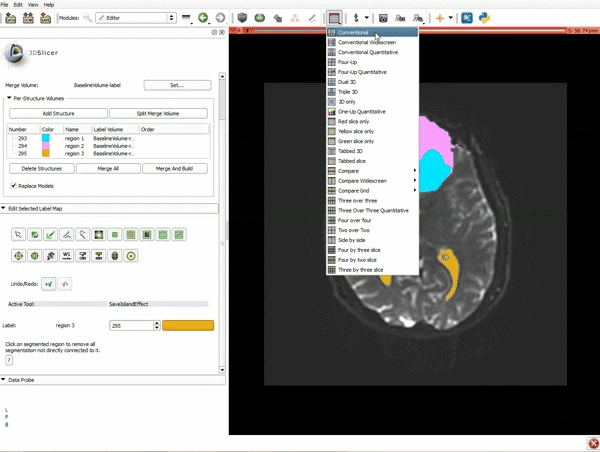
Segmentation