Difference between revisions of "Documentation/4.4/Training"
Tag: 2017 source edit |
|||
| (7 intermediate revisions by 5 users not shown) | |||
| Line 1: | Line 1: | ||
| − | <noinclude>{{documentation/ | + | <noinclude>{{documentation/historicaltraining}}</noinclude> |
__TOC__ | __TOC__ | ||
| Line 89: | Line 89: | ||
*Audience: End-users and developers | *Audience: End-users and developers | ||
*Modules: Data, Volumes, DWI to DTI Estimation, Diffusion Tensor Scalar Measurements, Editor, Markups,Tractography Label Map Seeding, Tractography Interactive Seeding | *Modules: Data, Volumes, DWI to DTI Estimation, Diffusion Tensor Scalar Measurements, Editor, Markups,Tractography Label Map Seeding, Tractography Interactive Seeding | ||
| − | *Based on: 3D Slicer | + | *Based on: [http://www.na-mic.org/Wiki/index.php/3DSlicer_4.4_r24272 3D Slicer Version 4.4.0-2015-05-21 r24272] |
*The [[Media:DiffusionMRI_tutorialData.zip |DTI dataset]] contains an MR Diffusion Weighted Imaging scan of the brain. | *The [[Media:DiffusionMRI_tutorialData.zip |DTI dataset]] contains an MR Diffusion Weighted Imaging scan of the brain. | ||
|align="right"| | |align="right"| | ||
| Line 150: | Line 150: | ||
| | | | ||
*[[Media:NRR-CTgLiverAblation.pdf| Tutorial on non-rigid MR/CT registration for CT-guided liver ablation]] | *[[Media:NRR-CTgLiverAblation.pdf| Tutorial on non-rigid MR/CT registration for CT-guided liver ablation]] | ||
| − | *Authors: Soichiro Tani, Atsushi Yamada, Junichi Tokuda, Dominik S. Meier, and Nobuhiko Hata | + | *Authors: Soichiro Tani, M.D, Atsushi Yamada, Ph.D, Junichi Tokuda, Ph.D, Dominik S. Meier, Ph.D, and Nobuhiko Hata, Ph.D |
*Audience: End-users interested in using Slicer to fuse pre- and intra-operative images for image-guided interventions. | *Audience: End-users interested in using Slicer to fuse pre- and intra-operative images for image-guided interventions. | ||
*Modules: N4ITK MRI Bias Correction, Editor, General Registration (BRAINS) | *Modules: N4ITK MRI Bias Correction, Editor, General Registration (BRAINS) | ||
| Line 280: | Line 280: | ||
{|width="100%" | {|width="100%" | ||
| | | | ||
| − | *[http:// | + | *[http://www.slicer.org/w/img_auth.php/f/f1/OpenIGTLinkTutorial_Slicer4.1.0_JunichiTokuda_Apr2012.pdf OpenIGTLink] |
*Authors: Junichi Tokuda, BWH | *Authors: Junichi Tokuda, BWH | ||
|align="right"| | |align="right"| | ||
| Line 314: | Line 314: | ||
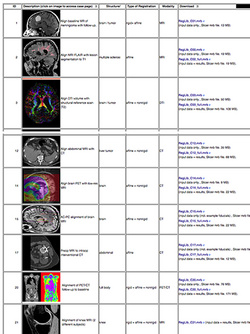
:Author: Dominik Meier, Ph.D. | :Author: Dominik Meier, Ph.D. | ||
:Audience: users interested learning/applying Slicer image registration technology | :Audience: users interested learning/applying Slicer image registration technology | ||
| − | |align="right"|[[Image:RegLib_table.png|250px|link=http://wiki.slicer.org/ | + | |align="right"|[[Image:RegLib_table.png|250px|link=http://wiki.slicer.org/wiki/Documentation/{{documentation/version}}/Registration/RegistrationLibrary]] |
|} | |} | ||
| Line 337: | Line 337: | ||
[http://www.youtube.com/results?search_query=3d+slicer&sm=3 YouTube videos about 3D Slicer] | [http://www.youtube.com/results?search_query=3d+slicer&sm=3 YouTube videos about 3D Slicer] | ||
| + | |||
| + | [https://www.youtube.com/channel/UC11x1iQ7ydSIFYw4L6wveXg/playlists 3D Slicer YouTube channel] | ||
Latest revision as of 22:12, 22 November 2022
Home < Documentation < 4.4 < Training
 |
This section is currently out-of-date and may contain errors but is retained for historical reference. Up-to-date training materials can be found at Documentation/Nightly/Training |
Contents
- 1 Introduction: Slicer 4.4 Tutorials
- 2 General Introduction
- 3 Tutorials for software developers
- 4 Specific functions
- 4.1 Slicer4 Diffusion Tensor Imaging Tutorial
- 4.2 Slicer4 Neurosurgical Planning Tutorial
- 4.3 Slicer4 3D Visualization of DICOM images for Radiology Applications
- 4.4 Slicer4 Quantitative Imaging tutorial
- 4.5 Slicer4 IGT
- 4.6 Slicer4 Non-rigid MR/CT registration for CT-guided liver ablation
- 4.7 Slicer4 3D Printing
- 4.8 Other
- 5 Summer 2014 Tutorial contest
- 6 Summer 2013 Tutorial contest
- 7 Summer 2012 Tutorial contest
- 8 Additional resources
- 9 External Resources
Introduction: Slicer 4.4 Tutorials
- This page contains "How to" tutorials with matched sample data sets. They demonstrate how to use the 3D Slicer environment (version 4.4 release) to accomplish certain tasks.
- For tutorials for other versions of Slicer, please visit the Slicer training portal.
- For "reference manual" style documentation, please visit the Slicer 4.4 documentation page
- For questions related to the Slicer4 Compendium, please send an e-mail to Sonia Pujol, Ph.D
|
Some of these tutorials are based on older releases of 3D Slicer. The concepts are still useful but bear in mind that some interface elements and features will be different in updated versions. |
General Introduction
Slicer Welcome Tutorial
|
Slicer4Minute Tutorial
|
Slicer4 Data Loading and 3D Visualization
|
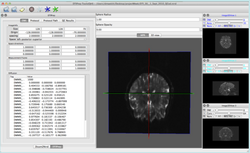
Tutorials for software developers
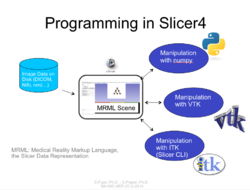
Slicer4 Programming Tutorial
|
For additional Python scripts examples, please visit the Script Repository page
Developing and contributing extensions for 3D Slicer
|
Specific functions
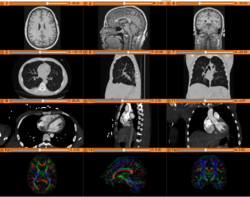
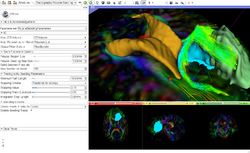
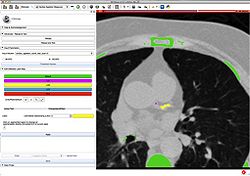
Slicer4 Diffusion Tensor Imaging Tutorial
|
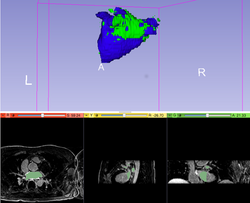
Slicer4 Neurosurgical Planning Tutorial
|
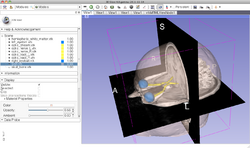
Slicer4 3D Visualization of DICOM images for Radiology Applications
|
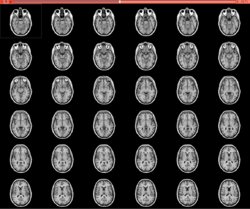
Slicer4 Quantitative Imaging tutorial
|
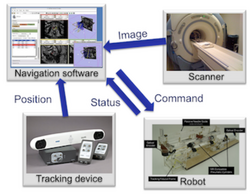
Slicer4 IGT
|
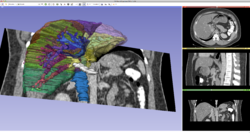
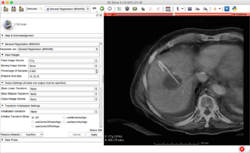
Slicer4 Non-rigid MR/CT registration for CT-guided liver ablation
|
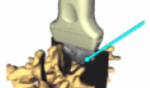
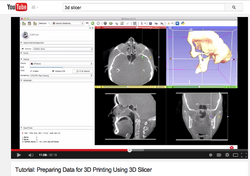
Slicer4 3D Printing
|
|
Other
Additional (non-curated) videos-based demonstrations using 3D Slicer are accessible on You Tube.
Summer 2014 Tutorial contest
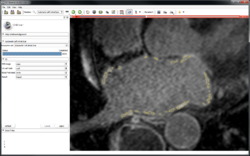
Cardiac Agatston Tutorial
|
CMR Toolkit LA workflow
|
Summer 2013 Tutorial contest
Cardiac MRI Toolkit
|
HelloCLI
|
SlicerRT
|
DTIPrep
|
Summer 2012 Tutorial contest
Automatic Left Atrial Scar Segmenter
|
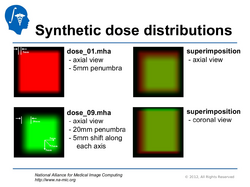
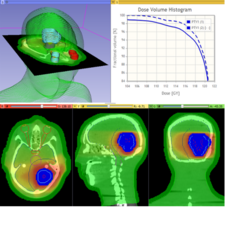
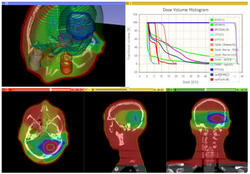
Qualitative and quantitative comparison of two RT dose distributions
|
Dose accumulation for adaptive radiation therapy
|
WebGL Export
|
OpenIGTLink
|
Additional resources
|
|
|
|
|
|
External Resources
Using the Editor
This set of tutorials about the use of slicer in paleontology is very well written and provides step-by-step instructions. Even though it covers slicer version 3.4, many of the concepts and techniques have applicability to the new version and to any 3D imaging field:
- Open Source Paleontologist: 3D Slicer: The Tutorial
- Open Source Paleontologist: 3D Slicer: The Tutorial Part II
- Open Source Paleontologist: 3D Slicer: The Tutorial Part III
- Open Source Paleontologist: 3D Slicer: The Tutorial Part IV
- Open Source Paleontologist: 3D Slicer: The Tutorial Part V
- Open Source Paleontologist: 3D Slicer: The Tutorial Part VI
Team Contributions
See the collection of videos on the Kitware vimeo album.
User Contributions
See the User Contributions Page for more content.