Documentation/4.3/Modules/GelDosimetry
From Slicer Wiki
Home < Documentation < 4.3 < Modules < GelDosimetry
|
For the latest Slicer documentation, visit the read-the-docs. |
Introduction and Acknowledgements
Authors: Jennifer Andrea (PerkLab, Queen's University), Csaba Pinter (PerkLab, Queen's University), Mattea Welch (University of Toronto)
Contributors: Kevin Alexander (Kingston General Hospital), John Schreiner (Kingston General Hospital)
Contacts:
- Csaba Pinter, <email>csaba.pinter@queensu.ca</email>
- How to report an error
License: Slicer license
Download/install: install 3D Slicer, start 3D Slicer, open the Extension Manager, install the GelDosimetry extension
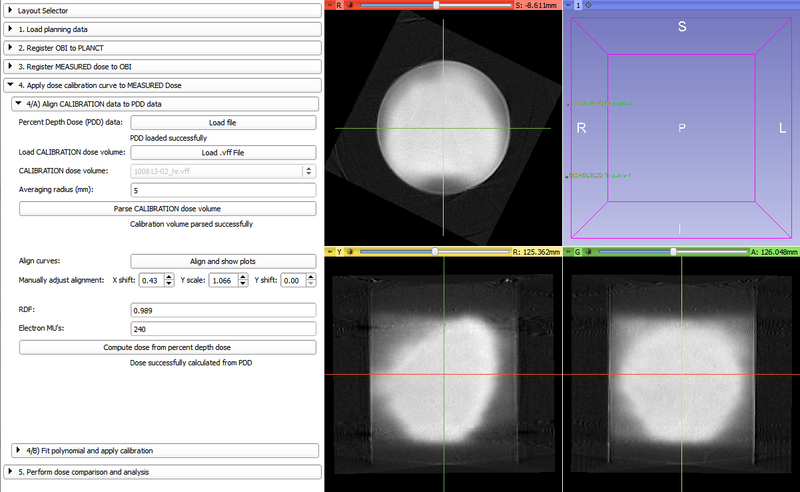
Extension Description
|
Tutorials
- N/A
Similar Extensions
- SlicerRT: GelDosimetry uses several modules from the SlicerRT extension
References
- N/A
Information for Developers
- N/A