Difference between revisions of "Modules:ExecutionModelTour-Documentation-3.6"
| Line 6: | Line 6: | ||
{| | {| | ||
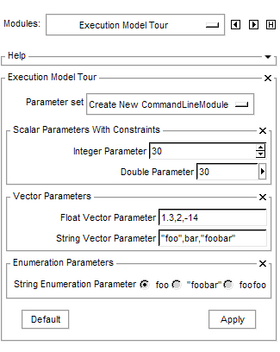
| − | |[[Image: | + | |[[Image:ExectionModelTourGUI.png|thumb|280px|Example user interface generated by a Command Line Module]] |
| − | |||
| − | |||
|} | |} | ||
Revision as of 19:34, 29 April 2010
Home < Modules:ExecutionModelTour-Documentation-3.6Return to Slicer 3.6 Documentation
Module Name
Execution Model Tour
General Information
Module Type & Category
Type: CLI
Category: Demonstration
Authors, Collaborators & Contact
- Author1: Bill Lorensen
- Author2: Dan Blezek
- Contact: bill.lorensen at gmail.com
Module Description
This module simply illustrates each type of parameter available through a Slicer Command Line Module. The parameters are described in an XML document and the user interface is constructed on the fly within Slicer.
Usage
This module is only intended to be used to demonstrate the types of parameters available and the user interface that will be generated for each parameter type.
Examples, Use Cases & Tutorials
- The Slicer Execution Model is documented here
Development
As new parameters are added to the Execution Model, we typically add a demonstration of that parameter type in this module.
Known bugs
Follow this link to the Slicer3 bug tracker.
Usability issues
Follow this link to the Slicer3 bug tracker. Please select the usability issue category when browsing or contributing.
Source code & documentation
Source Code: ExecutionModelTour.cxx
XML Description: ExecutionModelTour.xml
Usage:
./ExecutionModelTour [--processinformationaddress <std::string>]
[--xml] [--echo] [--region
<std::vector<std::vector<float> >>] ...
[--transform2 <std::string>] [--transform1
<std::string>] [--image2 <std::string>] [--image1
<std::string>] [--directory1 <std::string>]
[--file1 <std::string>] [--boolean3] [--boolean1]
[-e <Ron|Eric|Bill|Ross|Steve|Will>]
[--string_vector <std::vector<std::string>>] [-f
<std::vector<float>>] [-d <double>] [-i <int>]
[--] [--version] [-h] <std::string> <std::string>
Where:
--processinformationaddress <std::string>
Address of a structure to store process information (progress, abort,
etc.). (default: 0)
--xml
Produce xml description of command line arguments (default: 0)
--echo
Echo the command line arguments (default: 0)
--region <std::vector<std::vector<float> >> (accepted multiple times)
List of regions to process
--transform2 <std::string>
An output transform
--transform1 <std::string>
An input transform
--image2 <std::string>
An output image
--image1 <std::string>
An input image
--directory1 <std::string>
An input directory. If no default is specified, the current directory
is used,
--file1 <std::string>
An input file
--boolean3
A boolean default false (default: 0)
--boolean1
A boolean default true (default: 0)
-e <Ron|Eric|Bill|Ross|Steve|Will>, --enumeration <Ron|Eric|Bill|Ross
|Steve|Will>
An enumeration of strings (default: Bill)
--string_vector <std::vector<std::string>>
A vector of strings (default: foo,bar,foobar)
-f <std::vector<float>>, -- <std::vector<float>>
A vector of floats (default: 1.3,2,-14)
-d <double>, --double <double>
A double with constraints (default: 30)
-i <int>, --integer <int>
An integer without constraints (default: 30)
--, --ignore_rest
Ignores the rest of the labeled arguments following this flag.
--version
Displays version information and exits.
-h, --help
Displays usage information and exits.
<std::string>
(required) First index argument is an image
<std::string>
(required) Second index argument is an image
Description: Shows one of each type of parameter.
Author(s): Daniel Blezek, Bill Lorensen
Acknowledgements: This work is part of the National Alliance for Medical
Image Computing (NAMIC), funded by the National Institutes of Health
through the NIH Roadmap for Medical Research, Grant U54 EB005149.
More Information
Acknowledgment
This work is part of the National Alliance for Medical Image Computing (NAMIC), funded by the National Institutes of Health through the NIH Roadmap for Medical Research, Grant U54 EB005149. Information on the National Centers for Biomedical Computing can be obtained from National Centers for Biomedical Computing.