Difference between revisions of "Modules:VMTKCenterlines"
| Line 45: | Line 45: | ||
===Examples, Use Cases=== | ===Examples, Use Cases=== | ||
| + | * '''Example centerline extraction''' | ||
| + | The centerline computation requires an existing 3D surface model. This can be obtained using the [[Modules:VMTKEasyLevelSetSegmentation|VMTKEasyLevelSetSegmentation]] module. | ||
| + | |||
| + | After selecting the existing surface model as input, the second step is to prepare this model. This includes capping holes in the surface and removing unneeded triangles. The surface model color changes to green. | ||
| + | |||
| + | {| | ||
| + | |-valign="top" | ||
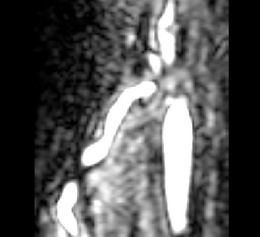
| + | |[[Image:Avf_orig_axial.png|thumb|260x260px|Axial view of a contrast-enhanced three-dimensional image acquired from a MR scanner, showing an AV fistula in an arm. (Courtesy of L. Antiga, Bioengineering Department, Mario Negri Institute, Ranica (BG), Italy and N.R. Planken, Academisch Ziekenhuis Maastricht, the Netherlands.)]] | ||
| + | |[[Image:Avf_3d_evolution_new.png|thumb|260x260px|A surface model of a segmentation of the original image obtained with the [[Modules:VMTKEasyLevelSetSegmentation|VMTKEasyLevelSetSegmentation]] module.]] | ||
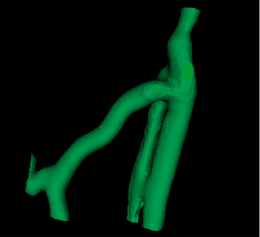
| + | |[[Image:Avf_3d_preparation.png|thumb|260x260px|This surface model after the preparation stage. Holes in the surface were closed.]] | ||
| + | |} | ||
| + | |||
| + | Now source and target fiducials have to be placed on the model in the 3D rendering window to specify the centerline routes. The fiducial positioning does not have to be exact because the module calculates the nearest position on the surface model to the input fiducials. After placing the fiducials, the centerline computation can be started. | ||
| + | |||
| + | On completion of the calculation, the inner Voronoi diagram and the corresponding centerlines are visible as polydata models in the 3D rendering window. | ||
| + | {| | ||
| + | |-valign="top" | ||
| + | |[[Image:Avf_3d_eikonal.png|thumb|260x260px|The inner Voronoi diagram colored by the solutions of the Eikonal equation obtained by Fast Marching back-tracking along the minimal path. The legend can be displayed using the [[Modules:Colors-Documentation-3.6| Colors module]].]] | ||
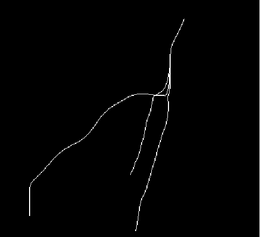
| + | |[[Image:Avf_3d_centerlines.png|thumb|260x260px|The centerline representation of the vessel's topology.]] | ||
| + | |} | ||
| + | |||
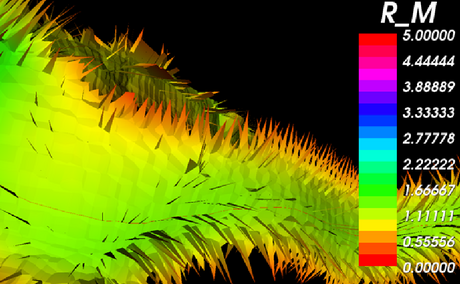
| + | A close-up of the Voronoi diagram shows the tiles separating the Voronoi regions. The modules also calculates the radius of the lumen for each point along the centerlines. | ||
| + | |||
| + | {| | ||
| + | |[[Image:Avf_3d_voronoi_big.png|thumb|460x460px|A close-up of the Voronoi diagram colored by the maximal inscribed sphere radius [vx].]] | ||
| + | |} | ||
| + | |||
| + | The extracted centerlines as well as the calculated radii along the centerlines can be exported and imported as a cloud of points to the filesystem. | ||
===Tutorials=== | ===Tutorials=== | ||
Revision as of 14:39, 20 April 2010
Home < Modules:VMTKCenterlinesReturn to Slicer 3.6 Documentation
Module Name
VMTKCenterlines, part of the  collection
collection
General Information
Module Type & Category
Type: Scripted Module
Category: Segmentation, Extension
Authors, Collaborators & Contact
- Author: Daniel Haehn, University of Heidelberg
- Acknowledgments: Luca Antiga, Mario Negri Institute; Steve Pieper, Isomics Inc.
- Contact: Daniel Haehn, haehn@bwh.harvard.edu
Module Description
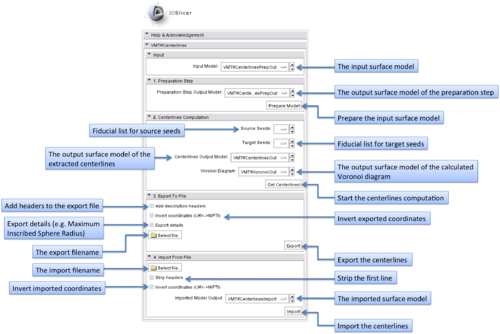
This module provides centerline computation of surface models in 3D Slicer using methods of the geometric analysis functionality of the Vascular Modeling Toolkit (http://www.vmtk.org). Centerlines are the central lumen lines in tubular structures and f.e. can be powerful descriptors of the vessel's topology. The module targets easy-to-use centerline extraction using seed and target points and also enables import and export functionality.
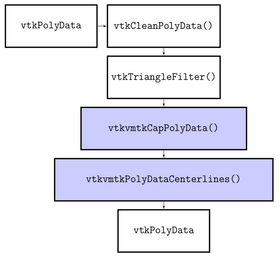
The actual centerline extraction is based on calculating an inner Voronoi diagram by computing the Delaunay tessellation.
This work is part of the NA-MIC VMTK Collaboration.
Official project page: http://www.vmtk.org/Main/VmtkIn3DSlicer
Usage
Installation
This module depends on the VmtkSlicerModule: see this page for installation notes.
The VMTKCenterlines module can be installed using the 3D Slicer extension wizard. The extension is called VMTKCenterlines.
When the module was successfully installed, it is available within 3D Slicer's module selector inside the category Vascular Modeling Toolkit.
Examples, Use Cases
- Example centerline extraction
The centerline computation requires an existing 3D surface model. This can be obtained using the VMTKEasyLevelSetSegmentation module.
After selecting the existing surface model as input, the second step is to prepare this model. This includes capping holes in the surface and removing unneeded triangles. The surface model color changes to green.
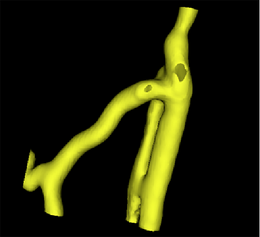 A surface model of a segmentation of the original image obtained with the VMTKEasyLevelSetSegmentation module. |
Now source and target fiducials have to be placed on the model in the 3D rendering window to specify the centerline routes. The fiducial positioning does not have to be exact because the module calculates the nearest position on the surface model to the input fiducials. After placing the fiducials, the centerline computation can be started.
On completion of the calculation, the inner Voronoi diagram and the corresponding centerlines are visible as polydata models in the 3D rendering window.
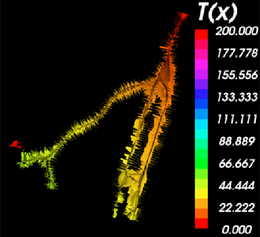 The inner Voronoi diagram colored by the solutions of the Eikonal equation obtained by Fast Marching back-tracking along the minimal path. The legend can be displayed using the Colors module. |
A close-up of the Voronoi diagram shows the tiles separating the Voronoi regions. The modules also calculates the radius of the lumen for each point along the centerlines.
The extracted centerlines as well as the calculated radii along the centerlines can be exported and imported as a cloud of points to the filesystem.
Tutorials
The following tutorials demonstrate the use of VMTKCenterlines in a pipeline of VMTK modules to extract the centerlines of coronary arteries.
 Step-by-step online tutorial: Segmentation of the Right Coronary Artery using VMTK in 3D Slicer. View online here. |
 Step-by-step tutorial including example dataset: The first slide of the Centerline Extraction tutorial. Download here. |
Quick Tour of Features and Use
The graphical user interface of the VMTKEasyLevelSetSegmentation module consists of four panels.
- Main module panel:
Parameter presets exist for different use-cases. These get loaded from the MRML file presets.xml inside the module directory and can be edited. The current parameters are stored to the MRML node of the module and therefore it is possible to work with different parameter sets.
- I/O panel:
The input volume and the output volumes for the initialization and evolution stage can be selected using this panel. The output volume of the initialization stage is used as an input for the evolution stage. Fiducial lists can be configured as source and target seeds. If the selected fiducial lists are equal, target seeds will be ignored. This is useful for the segmentation of vessel trees. A switch to choose between 3D visualization using models generated by Marching Cubes or Volume Rendering is available.
- Initialization panel:
The threshold slider is used as a parameter for the Fast Marching Upwind Gradient Image Filter and updates the slice viewers during change for visualization.
- Evolution panel:
The module supports to choose between Geodesic Active Contours and CURVES evolution methods using a checkbox.
Different weights are available to configure the inflation of the initial level-set. Is is possible to adjust the importance of inflation, curvature shape and attraction to the gradient ridges. The number of iterations defines how many steps of evolution are performed.
Development
Notes from the Developer(s)
The algorithms provided by VMTK and used by this module are C++ classes implemented using VTK and ITK.
Dependencies
This module depends on the VMTK libraries which are provided in the VmtkSlicerModule. Therefore the VmtkSlicerModule has to be installed before the VMTKEasyLevelSetSegmentation module can be used.
Known bugs & Usability issues
Follow this link to the VMTK in 3D Slicer bug tracker.
Source code & documentation
VMTKCenterlines is a Python Scripted Module. It follows the conventions of the Model View Controller pattern of slicer modules. This implies the separation of logic and GUI.
The class VMTKCenterlinesGUI derives from ScriptedModuleGUI and saves the current parameters to its own MRML node. The actual calls to the VMTK libraries as well as the import and export functionality are performed in the class VMTKCenterlinesLogic. Several general supporting functions are outsourced in the class VMTKCenterlinesHelper.
The complete source code is available at a NITRC SVN repository.
More Information
Acknowledgment
This work was funded by a grant of the Thomas-Gessmann Foundation part of the Founder Federation for German Science.
References
- Antiga L, Piccinelli M, Botti L, Ene-Iordache B, Remuzzi A and Steinman DA. An image-based modeling framework for patient-specific computational hemodynamics. Medical and Biological Engineering and Computing, 46: 1097-1112, Nov 2008.
- D. Hähn. Integration of the vascular modeling toolkit in 3d slicer. SPL, 04 2009. Available online at http://www.spl.harvard.edu/publications/item/view/1728.
- D. Hähn. Centerline Extraction of Coronary Arteries in 3D Slicer using VMTK based Tools. Master's Thesis. Department of Medical Informatics, University of Heidelberg, Germany. Feb 2010.
- Piccinelli M, Veneziani A, Steinman DA, Remuzzi A, Antiga L (2009) A framework for geometric analysis of vascular structures: applications to cerebral aneurysms. IEEE Trans Med Imaging. In press.