Modules:AtlasCreator
Return to Slicer 3.6 Documentation
AtlasCreator
General Information
Module Type & Category
Type: Built-in Loadable Module
Category: Registration
Authors, Collaborators & Contact
- Daniel Haehn, University of Pennsylvania
- Kilian Pohl, University of Pennsylvania
- Contact: Daniel Haehn (haehn@bwh.harvard.edu)
Module Description
The Atlas Creator module aligns images paired with segmentations to generate statistical atlases for several segmented structures.
Features:
- Support for BRAINSFit/CMTK/Congeal toolkits for Registration and Resampling
- Fixed Registration against a template or Dynamic Registration against a mean image
- Normalization of output atlases to a given value
- auto-detection of Structures (labels)
- different Output Casts
- Principal Component Analysis
- Cluster Computation Mode
- using existing Transforms and skipping the Registration (f.e. to re-run a previous generation)
Usage
This module supports different usage types and interfaces:
- Simple Atlas Creation (HowTo)
- The extended graphical user interface
- The command line interface
- External invocation using the Atlas Creator MRML Node
Use Cases, Examples
Input Data Requirements
The Atlas Creator expects input data to be structured the following way:
- In general each original image is accompanied by a manual segmentation.
- The images and the segmentations have to be in two different folders but have matching filenames.
- All images and segmentations have to be in Slicer-readable format.
- For Example:
- ./originals/case1.nrrd
- ./originals/case2.nrrd
- ./originals/case3.nrrd
- ./segmentations/case1.nrrd
- ./segmentations/case2.nrrd
- ./segmentations/case3.nrrd
| HowTo: Simple Atlas Creation |
| The following steps perform a Pair Fixed Registration against an automatic chosen template for automatically detected structures. |
| 1. Select the directories containing the Original Images and the Segmentations in the Input/Output panel. |
| 2. Select an Output Directory in the Input/Output panel. It makes sense to create a new directory to use for the Output. |
| 3. Hit Start! |
| 4. After some wait (minutes or hours!, depending on the number of input cases), the generated atlases and the used template will be loaded into 3D Slicer. |
Output Directory Structure
The Output Directory contains the following content after the Atlas Creator finished.
- ./template.nrrd - The fixed or dynamic template
- ./normalizedIntensityMapOfAligned.nrrd - An intensity map of all aligned cases, divided by the number of cases.
- ./registered/ - A directory containing the registered Original Images, if Delete Aligned Images is off.
- ./resampled/ - A directory containing the resampled Segmentations used for the Atlas Creation, if Delete Aligned Segs. is off.
- ./transforms/ - A directory containing the generated transforms as a result of the registration
- ./atlasX.nrrd - Atlas for label X, if Activate PCA is off.
- ./atlasY.nrrd - Atlas for label Y, if Activate PCA is off.
- ./atlasZ.nrrd - Atlas for label Z, if Activate PCA is off.
- ./PCA/PCAX/ - A directory containing generated PCA data for label X, if Activate PCA is on.
- ./PCA/PCAY/ - A directory containing generated PCA data for label Y, if Activate PCA is on.
- ./PCA/PCAZ/ - A directory containing generated PCA data for label Z, if Activate PCA is on.
- ... more atlases depending on which structures were used
Quick Tour of Features and Use
The Extended Graphical User Interface
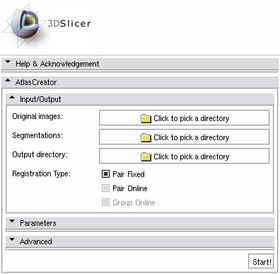
The Atlas Creator module is organized in panels from which only the Input/Output panel is expanded by default. Additional features are provided through an extended interface. This extended interface is divided into the Parameters panel and the Advanced panels which are collapsed by default.
Input/Output Panel
The Input/Output panel reflects the required settings for any atlas creation, simple or extended.
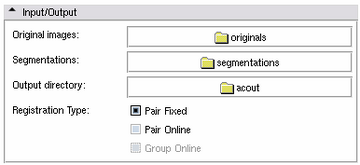
- Original Images: The folder containing the original images Required
- Segmentations: The folder containing the segmentations Required
- Output directory: The output folder - if it does not exist or is not empty, a new output folder with a similar name will be created Required
- Registration Type: The type of alignment used during registration stage
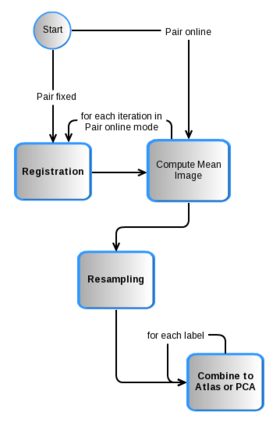
- Pair Fixed: Register all other cases against a fixed template. By default, the Atlas Creator chooses this template.
- Pair Online: Register all cases against a mean image of all cases which will be generated automatically.
- Group Online: Register all cases using the un-biased groupwise registration provided by the Congeal tool.
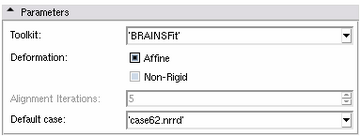
Parameters Panel
- Toolkit for Pair Fixed and Pair Online: By default BRAINSFit are used for registration and resampling. If the CMTK extensions are installed, it be used as an alternative. If CMTK was chosen but not installed, the Atlas Creator falls back to BRAINSFit.
- Deformation: Choose the deformation model for the registration stage.
- Affine: Use linear transformations (faster, less flexible)
- Non-rigid: Use non-rigid transformations on top of linear transformations (slower, more flexible)
- Alignment iterations, for Pair Online: The number of iterations for generating a mean image and registering against it in Pair Online mode.
- Default case, for Pair Fixed: The default case to use as the fixed template in Pair Fixed mode. This gets chosen automatically when selecting Original Images and Segmentations but can be modified for convenience.
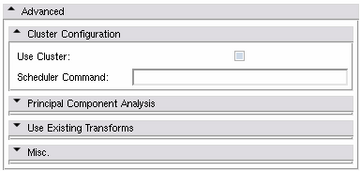
Advanced Panels
Cluster Configuration Panel
The Atlas Creator supports distributing all computations among a Grid Environment. This includes using a scheduler to start all computation jobs (registration, resampling, computing mean images, combine to atlases). All submitted jobs will be monitored for completion and will be re-started up to 4 times on failure. The atlas creation will continue if the failure is not critical but will notify of all failures.
- Use Cluster: If activated, all computation jobs will be submitted using the Scheduler Command.
- Scheduler Command: This command will be used to submit all jobs. For example, if the Scheduler Command is bsub < , the computation jobs will be run as bsub < script1.sh instead of just running script1.sh.
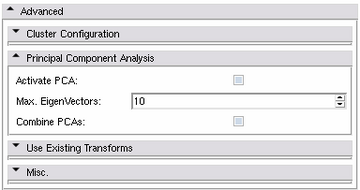
Principal Component Analysis Panel
Beside generating statistical atlases, the Atlas Creator module supports a Principal Component Analysis (PCA) to incorporate shape information.
- Activate PCA: Use Principal Component Analysis instead of statistical atlas generation.
- Max. Eigenvectors: The maximal number of eigenvectors to analyze during the PCA computation. Should be equal to the number of input cases but can be restricted for convenience. The Atlas Creator will detect if the number exceeds the number of cases.
- Combine PCAs: If activated, all structures (labels) will be defined in one PCA model instead of defining a PCA model for each structure.
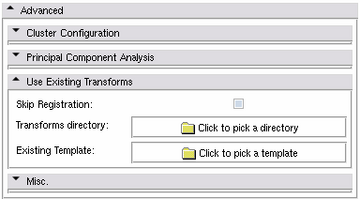
Use Existing Transforms Panel
It is possible to skip the registration stage and directly resample segmentations using existing transforms. This can be used to modify existing atlases.
- Skip Registration: If activated, the registration stage will be skipped and existing transforms are used for resampling the input segmentations.
- Transforms directory: The directory containing existing transformations. In order to be valid, these transformations must have been generated by the same toolkit (CMTK or BRAINSFit) as configured now.
- Existing Template:: The existing template which was used in the previous atlas generation (template.nrrd).
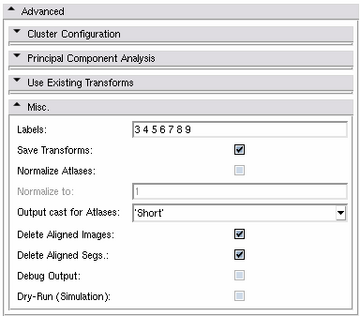
Misc. Panel
Miscellaneous settings can be specified using this panel.
- Labels: The list of structures for which to generate the different atlases. Each structure has to be represented by a label number. By default, the Atlas Creator reads this list from the default case - if specified - or from the first segmentation automatically.
- Save Transforms: If activated, the generated transforms of the registration stage will be saved. (Default setting)
- Normalize Atlases: The generated statistical atlases can be normalized to a value range from 0..X where X is the Normalize to value.
- Normalize to: If Normalize Atlases is activated, this value is the upper boundary of the range to which the atlases are normalized to. By default, this is 1.
- Output cast for Atlases: The output cast for the generated atlases can be specified. Default is short.
- Delete aligned Images: If activated, the aligned images will be deleted after atlas generation. If not activated, a sub-directory registered containing the aligned images will be available in the output directory.
- Delete aligned Segs.: If activated, the aligned images will be deleted after atlas generation. If not activated, a sub-directory resampled containing the aligned segmentations will be available in the output directory.
- Debug Output: This switch enables extra debug and verbose output during all computations.
- Dry-Run (Simulaton): If activated, no actual computation is performed and the atlas creation will be simulated. This might be helpful to test a long-run computation before actually performing it.
The Command Line Interface
Beside using the graphical user interface in 3D Slicer, the Atlas Creator can be accessed using the commandline interface.
$ cd Slicer3-release/lib/Slicer3/Modules/AtlasCreator/
$ python atlascreator.py --help
AtlasCreator for 3D Slicer
Version v0.4
Usage:
-h, --help
Show this information.
-i, --images DIR
Directory containing original images.
-s, --segmentations DIR
Directory containing segmentations.
-o, --output DIR
Output directory.
[....]
The full documentation of the commandline interface is available on a separate page.
Development
Notes from the Developer(s)
External invocation using the Atlas Creator MRML Node
It is easy to include the Atlas Creator in third-party 3D Slicer modules (C++, Tcl and Python are supported) or run it via the Tcl or Python consoles. To do so, one has to configure the vtkMRMLAtlasCreatorNode and fire a launch event.
The following example outlines the procedure by configuring a Pair fixed computation against a defaultCase (Python was chosen for this example):
from Slicer import slicer
import os
# we use the testdata which is shipped with the Atlas Creator
dataPath = os.path.normpath(slicer.Application.GetBinDir().. + '/../share/Slicer3/Modules/AtlasCreator/TestData/')
# create a new Atlas Creator MRML Node
n = slicer.vtkMRMLAtlasCreatorNode()
# Initialize with a default configuration
n.InitializeByDefault()
# set the original images by specifying the absolute filepaths for all cases divided by space
n.SetOriginalImagesFilePathList(dataPath + '/originals/case60.nrrd ' + dataPath + '/originals/case61.nrrd ' + dataPath + '/originals/case62.nrrd')
# set the segmentations by specifying the absolute filepaths for all cases divided by space
n.SetSegmentationsFilePathList(dataPath + '/segmentations/case60.nrrd ' + dataPath + '/segmentations/case61.nrrd ' + dataPath + '/segmentations/case62.nrrd')
# configure an output directory
n.SetOutputDirectory('/tmp/acout/')
# set the default case
n.SetFixedTemplateDefaultCaseFilePath(dataPath + '/originals/case62.nrrd')
# add the MRML node to the scene
slicer.MRMLScene.AddNode(n)
# start the computation by firing the launch event
n.Launch()
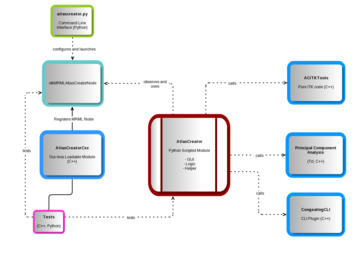
Architecture of the Atlas Creator
Dependencies
At least BRAINSFit, CMTK or Congeal have to be installed. BRAINSFit is deployed via Slicer, VMTK and Congeal are available as Slicer extensions.
Tests
On the Dashboard, these tests verify that the module is working on various platforms:
- vtkMRMLAtlasCreatorNodeTest1 Modules/AtlasCreator/Cxx/Testing/vtkMRMLAtlasCreatorNodeTest1.cxx
- AtlasCreatorLaunchFixedTest Modules/AtlasCreator/Cxx/Testing/vtkMRMLAtlasCreatorNodeLaunchTest1.py
- AtlasCreatorLaunchFixedFailProofTest Modules/AtlasCreator/Cxx/Testing/vtkMRMLAtlasCreatorNodeLaunchTest2.py
- AtlasCreatorLaunchDynamicTest Modules/AtlasCreator/Cxx/Testing/vtkMRMLAtlasCreatorNodeLaunchTest3.py
Known bugs
The AtlasCreator is currently not supported on Windows machines.
Usability issues
- Congeal support is still Under Construction.
Source code & documentation
Source code:
- Available in the 3D Slicer Trunk: Modules/AtlasCreator
Doxygen documentation:
More Information
Please cite this work as:
L. Zöllei, M. Shenton, W.M. Wells III, K.M. Pohl. The Impact of Atlas Formation Methods on Atlas-Guided Brain Segmentation, Statistical Registration. In Pair-wise and Group-wise Alignment and Atlas Formation Workshop at MICCAI 2007: Tenth International Conference on Medical Image Computing and Computer-Assisted Intervention, pp. 39 - 46, Brisbane, Australia, 2007
Acknowledgment
The research was funded by an ARRA supplement to NIH NCRR (P41 RR13218).
The Atlas Creator module benefits from several different Toolkits: Thank you for BRAINSTools (Hans Johnson et al.), for CMTK (Torsten Rolfing et al.) and for Congeal (J. De Bonet, L. Zöllei and W.M. Wells III).
Thank you to Dominique Belhachemi and Steve Pieper for their support!